Abstract
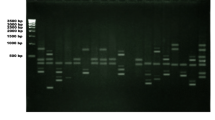
Retro-transposons are common components of plant genomes, functional at transcription, translation and integration levels. Their abundance and ability to transpose render them good potential markers. Present study was undertaken with the view of IRAP (inter-retrotransposon amplified polymorphism) and REMAP (retrotransposon-microsatellite amplified polymorphism) based DNA marker analysis for genus Citrus and related taxa. Both the IRAP and REMAP exhibited a remarkable variation among the tested genotypes. As a whole, ISSR, IRAP and REMAP analysis generated 113 score able bands ranging from 250 to 2,000 bps. There were 94 polymorphic bands, with an average of eight polymorphic bands per primer combination. The level of polymorphism was found to be 84%. No bands were shared with the ISSR pattern in REMAP analysis. Genetic similarity analysis was performed based on the Dice coefficient, and dendrogram was constructed by using the average linkage methods from combined analysis of IRAP and REMAP. A cophenetic correlation coefficient was also calculated. The clustering approach revealed a good adjustment between matrixes, with correlation coefficient of 0.77. Average similarity for all the genotype pairs was used as a cutoff value for defining the clusters. UPGMA demonstrated eight different clusters. Citrus genus showed wide range of heterogeneity, specially the mandarin group. Genera Fortunella and Poncirus were placed in relatively divergent cluster.




Similar content being viewed by others
Abbreviations
- IRAP:
-
Inter retrotransposon amplified polymorphism
- ISSR:
-
Inter simple sequence repeat
- REAMP:
-
Retro element microsatellite amplified polymorphism
References
Abkenar AA, Isshiki S, Tashiro Y (2004) Phylogenetic relationships in the “true citrus fruit trees” revealed by PCR-RFLP analysis of cpDNA. Scientia Hort 102:233–242
Amar K, Hirochika H (2001) Applications of retrotransposons as genetic tools in plant biology. Trends in Plant Sci 6:127–133
Ashalatha SN, Teo CH, Schwarzacher T, Harrison PH (2005) Genome classification of banana cultivars from south India using IRAP markers. Euphytica 144:285–290
Asins MJ, Monfort AJ, Mestre PE, Carbonell EA (1999) Citrus and Prunus copia-like retrotransposons. Thero Appl Genet 99:503–510
Barrett HC, Rhodes AM (1976) A numerical taxonomic study of affinity relationships in cultivated Citrus and its close relatives. Syst Bot 1:105–136
Bennetzen JL (2000) Transposable elements contributions to plant gene and genome evolution. Plant Mol Biol 42:351–369
Bennetzen JL (2002) Mechanisms and rates of genome expansion and contraction in flowering plants. Genetica 115:29–36
Bernet GP, Mestre PF, Pina JA, Asins MJ (2004) Molecular discrimination of lemon cultivars. Hortscience 39:165–169
Breto MP, Ruiz C, Pina JA (2001) The diversification of Citrus clementina Hort. ex Tan., a vegetatively propagated crop species. Mol Phylog Evol 21:285–293
Cameron JW, Frost HB (1968) Genetics, breeding and nucellar embryony. In: Reuther W, Batchelor LD, Webber HJ (eds) The citrus industry, vol II. Division of Agricultural Science, University of California, Berkeley, pp 325–370
Campos ET, Espinosa MAG, Warburton ML, Varela AS, Monter AV (2005) Characterization of Mandarin (Citrus spp.) using morphological and AFLP markers. Interciencia 30(11):687–693
Castelo J, Branco S, Eduardo AV, Malone G, Mauricio MK, Emilia M, Bernardes A, Claudete CM, Fernando IFC, Costa AO (2007) IRAP and REMAP assessments of genetic similarity in rice. J Appl Genet 48(2):107–113
Cheng YJ, Guo WW, Yi HL, Pang XM, Deng XX (2003) An efficient protocol for genomic DNA extraction from Citrus species. Plant Mol Biol Rep 21:177
Dalong G, Zhang H, Luo Z (2006) Genetic relationship of Diospyros kaki Thunb. and related species revealed by IRAP and REAMP analysis. Plant Sci 170:528–533
Dice LR (1945) Measure of the amount of ecological association between species. Ecology 26:297–307
Fang D, Krueger RR, Roose ML (1998) Phylogenetic relationships among selected citrus germplasm accessions revealed by inter-simple sequence repeat (ISSR) markers. J. Amer Soc Hort Sci 123(4):612–617
Federici CT, Fang DQ, Scora RW, Roose ML (1998) Phylogenetic relationships within the genus Citrus (Rutaceae) and related genera as revealed by RFLP and RAPD analysis. Theor Appl Genet 96:812–822
Gmitter FG (1995) Origin, evolution and breeding of the grapefruit. Plant Breed Rev 13:345–363
Green RM, Vardi A, Galun E (1986) The plastome of Citrus. Physical map, variation among Citrus cultivars and species and comparison with related genera. Theor Appl Genet 72:170–177
Gulsen O, Roose ML (2001) Chloroplast and nuclear genome analysis of the parentage of lemons. J Am Soc Hortic Sci 126:210–215
Handa T, Ishizawa Y, Oogaki C (1986) Phylogenetic study of fraction I protein in the genus Citrus and its close related genera. Japan J Genet 61:15–42
Herrero R, Asins MJ, Carbonell EA, Navarro L (1996) Genetic diversity in the orange subfamily Aurantioideae. I Intraspecies and intergenus variability. Theor Appl Genet 92:599–609
Higashi H, Hironaga T, Sennenbara T, Kunitake H, Komatsu H (2000) Phylogenetic classification of Fortunella species using RAPD method (in Japanese). J Japan Soc Hortic Sci 2:288
Kalendar R, Grob T, Regina MT, Suoniemi A, Schulman AH (1999) IRAP and REMAP: two new retrotransposon-based DNA fingerprinting techniques. Theor Appl Genet 98:704–711
Kalendar R, Tanskanen J, Immonen S, Nevo E, Schulman AH (2000) Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamica in response to sharp microclimatic divergence. Proc Natl Acad Sci USA 97:6603–6607
Kidwell MG, Lisch D (1997) Transposable elements as sources of variation in animals and plants. Proc Natl Acad Sci USA 94:7704–7711
Koehler-Santos P, Dornelles ALC, de Freitas LB (2003) Characterization of mandarin citrus germplasm from southern Brazil by morphological and molecular analyze. Pesq Agropec Bras 38:797–806
Kristiina AK, Kalendar R, Schulman AH (2006) TRIM retrotransposon occur in apple and are polymorphic between varieties but not sports. Theor Appl Genet. doi: 10.1007/s00122-005-0203-0
Kumar A, Bennetzen JL (1999) Plant retrotransposons. Annu Rev Genet 33:479–532
Manninan Q, Kalendar R, Robinson J, Schyknab AH (2000) Application of Bare-1 retrotransposon markers to the mapping of a major resistance gene for net blotch in barley. Mol Gen Genet 264:325–334
Moore GA (2001) Oranges and lemons: clues to the taxonomy of citrus from molecular markers. Trends Genet 17:536–540
Nicolosi E, Deng ZN, Gentile A, Malfa La S, Continella G, Tribulato E (2000) Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor Appl Genet 100:1155–1166
Pang XM, Hu CG, Deng XX (2003) Phylogenetic relationships among Citrus and its relatives as revealed by SSR markers. Acta Genet Sin 30:81–87
Pang XM, Hu CG, Deng XX (2007) Phylogenetic relationships within Citrus and its related genera as inferred from AFLP markers. Genet Resour Crop Evol 54:429–436
Queen RA, Gribbon BM, James C, Jack P, Flavell AJ (2004) Retrotransposon-based molecular markers for linkage and genetic diversity analysis in wheat. Mol Genet Geno 271(1):91–97
Raghuvanshi SS (1962) Cytogenetical studies in genus Citrus. IV. Evolution in genus Citrus. Cytologia 27:172–188
Rahman MM, Nito N (1994) Phylogenetic relationships in the kumquat (Fortunella) as revealed by isozyme analysis. Sci Hortic 57:17–28
Rohlf FJ (2000) NTSYS-pc: numerical taxonomy and multivariate analysis system, version 2.1. Exeter Software, New York
Sankar AA, Moore GA (2001) Evolution of inter-simple sequence repeat analysis from mapping in Citrus and extension of the genetic linkage map. Thero Appl Genet 102:206–214
Sokal RR, Rohlf FJ (1962) The comparison of dendrograms by objective methods. Taxon 11:30–40
Swengle WT (1946) The botany of Citrus and its wild relatives in the orange sub family. In: Webber HJ, Batcheler LD (eds) The citrus industry, vol 1. University of California Press, Berkeley, pp 128–474
Swingle WT, Reece PC (1967) The botany of Citrus and its wild relatives. In: Reuther W, Webber HJ, Batchelor LD (eds) The citrus industry, vol 1. University of California, Berkeley, pp 90–430
Tanaka T (1977) Fundamental discussion of Citrus classification. Stud Citrol 14:1–6
Torres MA, Soost RK, Diedenhofen U (1978) Leaf isozymes as genetic markers in citrus. Amer J Bot 65:869–881
Wei J (2007) Characterization of retrotransposon elements and development of related molecular markers in citrus. PhD Thesis, Huazhong Agricultural University, Wuhan, China
Yamamoto M, Kobayashi S, Nakamura Y, Yamada Y (1993) Phylogenic relationships of Citrus revealed by diversity of cytoplasmic genomes. In: Hayashi T, Omura M, Scott NS (eds) Techniques on gene diagnosis and breeding in fruit trees. Fruit Trees Research Station, Okitsu, pp 39–46
Yannic G, Baumel A, Ainouche M (2004) Uniformity of the nuclear and chloroplast genomes of Spartina maritima (Poaceae), a salt-marsh species in decline along the Western European Coast. Heredity 93:182–188
Acknowledgments
This research was supported by the Natural Science Foundation of China (NSFC No. 30921002).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Biswas, M.K., Baig, M.N.R., Cheng, YJ. et al. Retro-transposon based genetic similarity within the genus Citrus and its relatives. Genet Resour Crop Evol 57, 963–972 (2010). https://doi.org/10.1007/s10722-010-9533-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-010-9533-0