Abstract
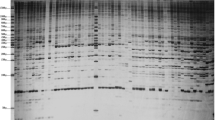
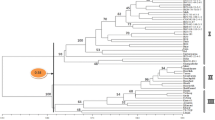
To evaluate the genetic diversity and to clarify the genetic relationships of Japanese peach cultivars, we analyzed the amplified fragment length polymorphism (AFLP) and traced the pedigree of 17 Japanese commercial peach cultivars and six traditional accessions. Sixteen AFLP primer combinations produced a total of 837 fragments and 146 polymorphic bands with a polymorphism percentage of 17.5%. All of the peach accessions could be identified from differences in at least 10 polymorphic bands. A cluster analysis showed that all the Japanese commercial peach cultivars, except ‘Kiyomi’ and ‘Jichigetsuto’, formed a major group consisting of three sub-groups. Of the six traditional accessions, four were genetically distant from the Japanese commercial peach cultivars while two accessions from China were classified into the Japanese commercial peach cultivars group. Both the AFLP analysis and pedigree tracing suggested that Japanese commercial peach cultivars are mainly derived from ‘Shanhai Suimitsuto’, one of the traditional accessions from China. Although the genetic relationships revealed by AFLP were generally in agreement with those shown by the pedigree information, some contradictions were found. Combining the AFLP results and pedigree information can provide a better understanding of the genetic relationships of Japanese peach cultivars.
Similar content being viewed by others
References
W.V. Baird A.S. Estager J.K. Wells (1994) ArticleTitleEstimating nuclear DNA content in peach and related diploid species using laser flow cytometry and DNA hybridization J. Am. Soc. Hort. Sci. 119 1312–1316
B.A. Barrett K.K. Kidwell (1998) ArticleTitleAFLP-based genetic diversity assessment among wheat cultivars from the Pacific Northwest Crop Sci. 38 1261–1271 Occurrence Handle1:CAS:528:DyaK1cXlvFyhsbg%3D Occurrence Handle10.2135/cropsci1998.0011183X003800050025x
J. Felsenstein (1995) PHYLIP (Phylogeny inference package) version 3.57c Univ. Washington Press Seattle
T. Helms J. Orf G. Valllad P. McClean (1997) ArticleTitleGenetic variancecoefficient of parentageand genetic distance of six soybean populations Theor. Appl. Genet. 94 20–26 Occurrence Handle10.1007/s001220050376 Occurrence Handle1:STN:280:DC%2BD1M3msVOmug%3D%3D Occurrence Handle19352740
D.J. Mackill Z. Zhang E.D. Redoña P.M. Colowit (1996) ArticleTitleLevel of polymorphism and genetic mapping of AFLP markers in rice Genome 39 969–977 Occurrence Handle8890522 Occurrence Handle1:CAS:528:DyaK28XntVSnurs%3D
P.J. Maughan M.A. Saghai Maroof G.R. Buss G.M. Huestis (1996) ArticleTitleAmplified fragment length polymorphism (AFLP) in soybean species diversity, inheritancenear-isogenic line analysis Theor. Appl. Genet. 93 392–401 Occurrence Handle1:CAS:528:DyaK28XmtF2ns7w%3D Occurrence Handle10.1007/s001220050293
M. Nei (1978) ArticleTitleEstimation of average heterozygosity and genetic distance from a small number of individuals Genetics 89 583–590 Occurrence Handle17248844 Occurrence Handle1:STN:280:DC%2BC3crpt1Kqtg%3D%3D
M. Nei W.H. Li (1979) ArticleTitleMathematical model for studying genetic variation in terms of restriction endonuclease Proc. Natl. Acad. Sci. USA 76 5256–5273 Occurrence Handle10.1073/pnas.76.10.5269
T. Nicola V. Negri (2002) ArticleTitleEfficiency of three PCR-based markers in assessing genetic variation among cowpea (Vigna unguiculata subsp. unguiculata) landraces Genome 45 268–275 Occurrence Handle10.1139/g01-146
W. Powell M. Morgante C. Andre M. Hanafey J. Vogel S. Tingey A. Rafalki (1996) ArticleTitleThe comparison of RFLP, RAPD, AFLP, and SSR (microsatellite) markers for germplasm analysis Molecular breeding 2 225–238 Occurrence Handle1:CAS:528:DyaK28XntlGrtLk%3D Occurrence Handle10.1007/BF00564200
R. Scorza S.A. Mehlenbacher G.W. Lightner (1985) ArticleTitleInbreeding and coancestry of freestone peach cultivars of the eastern United States and implications for peach germplasm improvement J. Am. Soc. Hort. Sci. 110 547–552
T. Shimada T. Yamamoto H. Yaegaki M. Yamaguchi M. Yoshida T. Hayashi (1999) ArticleTitleApplication of AFLP to molecular genetic analysis in peach J. Jpn. Soc. Hort. Sci. 68 67–69 Occurrence Handle1:CAS:528:DyaK1MXnt1artQ%3D%3D
A. Tenkouano J.H. Crouch H.K. Crouch D. Vuylsteke R. Ortiz (1999) ArticleTitleComparison of DNA markers and pedigree-based methods of genetic analysis of plantain and banana (Musa spp.) clones I. Estimation of genetic relationships Theor. Appl. Genet. 98 62–68 Occurrence Handle1:CAS:528:DyaK1MXhslOmtbk%3D Occurrence Handle10.1007/s001220051040
P. Vos R. Hogers M. Bleeker M. Reijans T. Lee ParticleVan de M. Hornes A. Frijters J. Pot J. Pelemen M. Kuiper M. Zabeau (1995) ArticleTitleAFLP: a new technique for DNA fingerprinting Nucleic Acids Res. 23 4407–4414 Occurrence Handle7501463 Occurrence Handle1:CAS:528:DyaK2MXpslensbo%3D
T. Yamamoto T. Shimada K. Kotobuki Y. Morimoto M. Yoshida (1998) ArticleTitleGenetic characterization of Asian chestnut varieties assessed by AFLP Breeding Sci. 48 359–363 Occurrence Handle1:CAS:528:DyaK1MXhtFGnsr4%3D
T. Yamamoto K. Mochida T. Hayashi (2003) ArticleTitleShanhai Suimitsutoone of the origins of Japanese peach cultivars J. Japan Soc. Hort. Sci. 72 116–121 Occurrence Handle1:CAS:528:DC%2BD3sXitlygu74%3D Occurrence Handle10.2503/jjshs.72.116
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, D.H., Wahyuni, S., Sato, Y. et al. Genetic Diversity and Relationships of Japanese Peach (Prunus persica L.) Cultivars Revealed by AFLP and Pedigree Tracing. Genet Resour Crop Evol 53, 883–889 (2006). https://doi.org/10.1007/s10722-004-0575-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-004-0575-z