Abstract
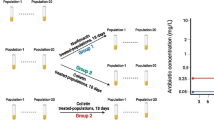
Since antibiotic resistance is a growing public health problem worldwide, it is important to understand how antibiotics and spontaneous mutations cooperate and shape the genome-wide mutation rate and spectrum. Here, we quantitatively evaluate genome-wide mutational profiles of Escherichia coli after long-term subinhibitory exposure to a broad-spectrum (streptomycin) and a narrow-spectrum antibiotic (nalidixic acid), using a mutation accumulation design combined with whole-genome resequencing of replicate lines as a mutagenicity test. We determined that, while the genome-wide mutation rate is slightly higher in the streptomycin-treated lines compared to the control lines, there is a significant increase in the nalidixic acid-treated lines. Our findings suggest that both broad and narrow-spectrum antibiotics may elevate the mutation rates in E. coli, but mechanisms of action may affect the consequence, thus contribute to accelerating the rate of adaptation and conferring antibiotic resistance.


Similar content being viewed by others
Data availability
Raw data were deposited in the NCBI SRA under Bio Project ID of PRJNA630872.
References
Boe L, Tolker-Nielsen T, Eeegholm KM, Spliid H, Vrang A (1994) Fluctuation analysis of mutations to nalidixic acid resistance in Escherichia coli. J Bacteriol 176:2781–2787. https://doi.org/10.1128/jb.176.10.2781-2787.1994
Chu X-L, Zhang B-W, Zhang Q-G, Zhu B-R, Lin K, Zhang D-Y (2018) Temperature responses of mutation rate and mutational spectrum in an Escherichia coli strain and the correlation with metabolic rate. BMC Evol Biol 18:126. https://doi.org/10.1186/s12862-018-1252-8
Davies J, Davies D (2010) Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev 74:417–433. https://doi.org/10.1128/MMBR.00016-10
DePristo MBE et al (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43:491–498. https://doi.org/10.1038/ng.806
Didier J-P, Villet R, Huggler E, Lew DP, Hooper DC, Kelley WL, Vaudaux P (2011) Impact of ciprofloxacin exposure on Staphylococcus aureus genomic alterations linked with emergence of rifampin resistance. Antimicrob Agents Chemother 55:1946–1952. https://doi.org/10.1128/AAC.01407-10
Fair RJ, Tor Y (2014) Antibiotics and bacterial resistance in the 21st century. Perspect Medicin chem 6:25–64. https://doi.org/10.4137/PMC.S14459
Fijalkowska IJ, Dunn RL, Schaaper RM (1997) Genetic requirement and mutational specificity of the Escherichia coli SOS mutator activity. J Bacteriol 179:7435–7445
Frana TS et al (2013) Isolation and characterization of methicillin-resistant Staphylococcus aureus from pork farms and visiting veterinary students. PLoS ONE 8:e53738. https://doi.org/10.1371/journal.pone.0053738
Hershberg R, Petrov DA (2010) Evidence that mutation is universally biased towards AT in bacteria. PLoS Genet. https://doi.org/10.1371/journal.pgen.1001115
Hooper DC, Jacoby GA (2016) Topoisomerase inhibitors: fluoroquinolone mechanisms of action and resistance. Cold Spring Harb Perspect Med 6:a025320. https://doi.org/10.1101/cshperspect.a025320
Hughes D (2014) Selection and evolution of resistance to antimicrobial drugs. IUBMB Life 66:521–529. https://doi.org/10.1002/iub.1278
Huitric E, Verhasselt P, Koul A, Andries K, Hoffner S, Andersson DI (2010) Rates and mechanisms of resistance development in Mycobacterium tuberculosis to a novel diarylquinoline ATP synthase inhibitor. Antimicrob Agents Chemother 54:1022–1028. https://doi.org/10.1128/AAC.01611-09
Isler B et al (2019) Antibiotic overconsumption and resistance in Turkey. Clin Microbiol Infect 25:651–653. https://doi.org/10.1016/j.cmi.2019.02.024
Ittisoponpisan S, Islam SA, Khanna T, Alhuzimi E, David A, Strenberg MJE (2019) Can predicted protein 3D structures provide reliable insights into whether missense variants are disease associated? J Mol Biol 431:2197–2212. https://doi.org/10.1016/j.jmb.2019.04.009
Jørgensen KM, Wassermann T, Jensen PØ, Hengzuang W, Molin S, Høiby N, Ciofu O (2013) Sublethal ciprofloxacin treatment leads to rapid development of high-level ciprofloxacin resistance during long-term experimental evolution of Pseudomonas aeruginosa. Antimicrob Agents Chemother 57:4215–4221. https://doi.org/10.1128/AAC.00493-13
Keightley PD, Trivedi U, Thomson M, Oliver F, Kumar S, Blaxter ML (2009) Analysis of the genome sequences of three Drosophila melanogaster spontaneous mutation accumulation lines. Genome Res 19:1195–1201. https://doi.org/10.1101/gr.091231.109
Kibota TT, Lynch M (1996) Estimate of the genomic mutation rate deleterious to overall fitness in E. coli. Nature 381:694–696. https://doi.org/10.1038/381694a0
Kohanski MA, DePristo MA, Collins JJ (2010) Sublethal antibiotic treatment leads to multidrug resistance via radical-induced mutagenesis. Mol Cell 37:311–320. https://doi.org/10.1016/j.molcel.2010.01.003
Komp Lindgreen P, Karlsson A, Hughes D (2003) Mutation rate and evolution of fluoroquinolone resistance in Escherichia coli isolates from patients with urinary tract infection. Antimicrob Agents Chemother 47:3222–3232. https://doi.org/10.1128/aac.47.10.3222-3232.2003
Krasovec M, Eyre-Walker A, Sanchez-Ferandin S, Piganeau G (2017) Spontaneous mutation rate in the smallest photosynthetic eukaryotes. Mol Biol Evol 34:1770–1779
Krause KM, Serio AW, Kane TR, Connolly LE (2016) Aminoglycosides: an overview. Cold Spring Harb Perspect Med 6:a027029. https://doi.org/10.1101/cshperspect.a027029
Kucukyildirim S (2020) In vitro evaluation of the effect of streptomycin on the rate and molecular spectrum of spontaneous mutations in Klebsiella aerogenes. J Turk Soc Microbiol 50:156–163
Lazarus B, Paterson DL, Mollinger JL, Rogers BA (2015) Do human extraintestinal Escherichia coli infections resistant to expanded-spectrum cephalosporins originate from food-producing animals? A systematic review. Clin Infect Dis 60:439–452. https://doi.org/10.1093/cid/ciu785
Lee H, Popodi E, Tang H, Foster PL (2012) Rate and molecular spectrum of spontaneous mutations in the bacterium Escherichia coli as determined by whole-genome sequencing. Proc Natl Acad Sci USA 109:E2774-2783. https://doi.org/10.1073/pnas.1210309109
Leeds JA, Sachdeva M, Mullin S, Barnes SW, Ruzin A (2013) In vitro selection, via serial passage, of Clostridium difficile mutants with reduced susceptibility to fidaxomicin or vancomycin. J Antimicrob Chemother 69:41–44. https://doi.org/10.1093/jac/dkt302
Levin BR, Rozen DE (2006) Non-inherited antibiotic resistance. Nat Rev Microbiol 4:556–562. https://doi.org/10.1038/nrmicro1445
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Lind PA, Andersson DI (2008) Whole-genome mutational biases in bacteria. Proc Nat Acad Sci USA 105:17878–17883. https://doi.org/10.1073/pnas.0804445105
Lobanovska M, Pilla G (2017) Penicillin`s discovery and antibiotic resistance: lessons for the future? Yale J Biol Med 90:135–145
Long H et al (2016) Antibiotics treatment enhances the genome-wide mutation of target cells. Proc Nat Acad Sci USA 113:2498–2505. https://doi.org/10.1073/pnas.160120811
Long H et al (2018) Evolutionary determinants of genome-wide nucleotide composition. Nat Ecol Evol 2:237–240. https://doi.org/10.1038/s41559-017-0425-y
Luzzatto L, Apirion D, Schlessinger D (1968) Mechanism of action of streptomycin in E. coli: interruption of the ribosome cycle at the initiation of protein synthesis. Proc Nat Acad Sci USA 60:873–880. https://doi.org/10.1073/pnas.60.3.873
McKenna AHM et al (2010) The genome analysis toolkit: a Map reduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Michaels ML, Cruz C, Grollman AP, Miller JH (1992) Evidence that MutY and MutM combine to prevent mutations by an oxidatively damaged form of guanine in DNA. Proc Nat Acad Sci USA 89:7022–7025
Ness RW, Morgan AD, Vasanthakrishnan RB, Colegrave N, Keightley PD (2015) Extensive de novo mutation rate variation between individuals and across the genome of Chlamydomonas reinhardtii. Genome Res 25:1739–1749. https://doi.org/10.1101/gr.191494.115
Oz T et al (2014) Strength of selection pressure is an important parameter contributing to the complexity of antibiotic resistance evolution. Mol Biol Evol 31:2387–2401. https://doi.org/10.1093/molbev/msu191
Pham TDM, Ziora ZM, Blaskovich MAT (2019) Quinolone antibiotics. Medchemcomm 10:1719–1739. https://doi.org/10.1039/c9md00120d
R Development Core Team (2015) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rodrigues CHM, Pires DEV, Ascher DB (2018) DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res 46:W350-355. https://doi.org/10.1093/nar/gky300
Schaaper RM, Bond BI, Fowler RG (1989) A.T — C.G transversions and their prevention by the Escherichia coli mutT and mutHLS pathways. Mol Gen Genet 219:256–262. https://doi.org/10.1007/BF00261185
Shewaramani S, Finn TJ, Leahy SC, Kassen R, Rainey PB, Moon CD (2017) Anaerobically grown Escherichia coli has an enhanced mutation rate and distinct mutational spectra. PLOS Genet 13:e1006570. https://doi.org/10.1371/journal.pgen.1006570
Song LY et al (2016) Mutational consequences of ciprofloxacin in Escherichia coli. Antimicrob Agents Chemother 60:6165–6172. https://doi.org/10.1128/AAC.01415-16
Springer B, Kidan YG, Prammananan T, Ellrott K, Böttger EC, Sander P (2001) Mechanisms of streptomycin resistance: selection of mutations in the 16S rRNA gene conferring resistance. Antimicrob Agents Chemother 45:2877–2884. https://doi.org/10.1128/AAC.45.10.2877-2884.2001
Stewart FM (1994) Fluctuation tests: how reliable are the estimates of mutation rates? Genetics 137:1139–1146
Sung W, Ackerman MS, Dillon MM, Platt TG, Fuqua C, Cooper VS, Lynch M (2016) Evolution of the insertion-deletion mutation rate across the tree of life. G3 (Bethesda) 6:2583–2591. https://doi.org/10.1534/g3.116.030890
Thorvaldsdottir H, Robinson JT, Mesirov JP (2013) Integrated genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14:178–192. https://doi.org/10.1093/bib/bbs017
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G et al (2013) From fastQ data to high confidence variant calls: the genome analysis toolkit best practices pipeline. Curr Protoc Bioinformatics 43:11.10.11-11.10.33. https://doi.org/10.1002/0471250953.bi1110s43
van Hoek AHAM, Mevius D, Guerra B, Mullany P, Roberts AP, Aarts HJM (2011) Acquired antibiotic resistance genes: an overview. Front Microbiol 2:203. https://doi.org/10.3389/fmicb.2011.00203
Wijker CA, Lafleur VM (1998) Influence of the UV-activated SOS response on the γ-radiation-induced mutation spectrum in the lacI gene. Mutat Res/DNA Repair 408:195–201
Windels EM, Michiels JE, Fauvart M, Wenseleers T, Van den Bergh B, Michiels J (2019) Bacterial persistence promotes the evolution of antibiotic resistance by increasing survival and mutation rates. ISME J 13:1239–1251. https://doi.org/10.1038/s41396-019-0344-9
Woodford N, Ellington MJ (2007) The emergence of antibiotic resistance by mutation. Clin Microbiol Infect 13:5–18. https://doi.org/10.1111/j.1469-0691.2006.01492.x
Zurek L, Ghosh A (2014) Insects represent a link between food animal farms and the urban environment for antibiotic resistance traits. Appl Environ Microbiol 80:3562–3567. https://doi.org/10.1128/AEM.00600-14
Acknowledgements
We thank Arif Duyar and Merve Dokez for technical support. This work supported by Hacettepe University Research Fund (Grant Number FBB-2017-15738 to S.K.).
Funding
Hacettepe University Research Fund (Grant Number FBB-2017-15738 to S.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ozdemirel, H.O., Ulusal, D. & Kucukyildirim Celik, S. Streptomycin and nalidixic acid elevate the spontaneous genome-wide mutation rate in Escherichia coli. Genetica 149, 73–80 (2021). https://doi.org/10.1007/s10709-021-00114-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-021-00114-w