Abstract
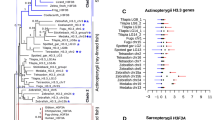
HIF-α transcription factors, as key master regulators of oxygen homeostasis, constitute a subgroup of the large bHLH-PAS transcription factor family and have been identified in many vertebrates. Although the amino acid sequences of bHLH-PAS domain are conserved, the physiological and pathological roles of this family are variable. They also have different patterns of expression. It is possible that the HIF-α copies have been retained as a consequence of adaptive amino acid replacements or relaxed selective constraint which have conferred subtle changes in function after duplications. Phylogenetic analysis indicated that at least two major duplications had occurred early in the vertebrate lineages. Analyses of the ratios of nonsynonymous/synonymous substitution rates revealed that relaxation of selective constraints might play important roles over evolutionary time and shape variation in some members of the family. The coefficients of functional divergence (θ) estimated between pairwise comparisons of gene groups from HIF-1α, HIF-2α, and HIF-3α indicated statistically significant site-specific shift of evolutionary rates between them, suggesting that altered functional constraints may have taken place at some amino acid residues after their duplications. Moreover, we also mapped sites identified to have been relaxed from purifying selection onto the three-dimensional structure of human HIF-2α. Overall, our study demonstrated that the functional diversity of HIF-αs members may be caused by relaxed negative selection on the N-terminal transactivation domains after HIF-αs duplications, which recruited new partners leading to functional specificity.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Altschul SF, Madden TL, Schaffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Brahimi-Horn MC, Pouyssegur J (2007) Harnessing the hypoxia-inducible factor in cancer and ischemic disease. Biochem Pharmacol 73:450–457
Carmeliet P, Dor Y, Herbert JM et al (1998) Role of HIF-1alpha in hypoxia-mediated apoptosis, cell proliferation and tumour angiogenesis. Nature 394:485–490
Compernolle V, Brusselmans K, Acker T et al (2002) Loss of HIF-2alpha and inhibition of VEGF impair fetal lung maturation, whereas treatment with VEGF prevents fatal respiratory distress in premature mice. Nat Med 8:702–710
Cowden Dahl KD, Fryer BH, Mack FA et al (2005) Hypoxia-inducible factors 1alpha and 2alpha regulate trophoblast differentiation. Mol Cell Biol 25:10479–10491
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform 5:113
Force A, Lynch M, Pickett FB et al (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Goodman M, Moore GW, Matsuda G (1975) Darwinian evolution in the genealogy of haemoglobin. Nature 253:603–608
Gu X (1999) Statistical methods for testing functional divergence after gene duplication. Mol Biol Evol 16:1664–1674
Gu X (2001) Maximum-likelihood approach for gene family evolution under functional divergence. Mol Biol Evol 18:453–464
Gu X, Vander Velden K (2002) DIVERGE: phylogeny-based analysis for functional-structural divergence of a protein family. Bioinformatics 18:500–501
Gu Y-Z, Moran SM, Hogenesch JB et al (1998) Molecular characterization and chromosomal localization of a third α-class hypoxia inducible factor subunit, HIF3α. Gene Expr 7:205–213
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Gustafsson MV, Zheng X, Pereira T et al (2005) Hypoxia requires notch signaling to maintain the undifferentiated cell state. Dev Cell 9:617–628
Heidbreder M, Frohlich F, Johren O et al (2003) Hypoxia rapidly activates HIF-3α mRNA expression. FASEB J 17:1541–1543
Hu CJ, Wang LY, Chodosh LA et al (2003) Differential roles of hypoxia-inducible factor 1alpha (HIF-1alpha) and HIF-2alpha in hypoxic gene regulation. Mol Cell Biol 23:9361–9374
Huang LE, Gu J, Schau M et al (1998) Regulation of hypoxia-inducible factor 1α is mediated by an O2-dependent degradation domain via the ubiquitin-proteasome pathway. Proc Natl Acad Sci U S A 95:7987–7992
Hughes J, Criscuolo F (2008) Evolutionary history of the UCP gene family: gene duplication and selection. BMC Evol Biol 8:306
Hughes DA, Jastroch M, Stoneking M et al (2009) Molecular evolution of UCP1 and the evolutionary history of mammalian non-shivering thermogenesis. BMC Evol Biol 9:4
Iyer NV, Kotch LE, Agani F et al (1998a) Cellular and developmental control of O2 homeostasis by hypoxia-inducible factor 1 alpha. Genes Dev 12:149–162
Iyer NV, Leung SW, Semenza GL (1998b) The human hypoxia-inducible factor 1alpha gene: HIF1A structure and evolutionary conservation. Genomics 52:159–165
Jaakkola P, Mole DR, Tian YM et al (2001) Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 292:468–472
Kimura M (1979) The neutral theory of molecular evolution. Sci Am 241:98–100, 102, 108 passim
Lynch VJ, Roth JJ, Wagner GP (2006) Adaptive evolution of Hox-gene homeodomains after cluster duplications. BMC Evol Biol 6:86
Makino Y, Cao R, Svensson K et al (2001) Inhibitory PAS domain protein is a negative regulator of hypoxia-inducible gene expression. Nature 414:550–554
Martinez-Castilla LP, Alvarez-Buylla ER (2003) Adaptive evolution in the Arabidopsis MADS-box gene family inferred from its complete resolved phylogeny. Proc Natl Acad Sci U S A 100:13407–13412
Maxwell PH, Wiesener MS, Chang GW et al (1999) The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399:271–275
Nielsen R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148:929–936
Nylander JAA (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University, Uppsala
Park JI, Semyonov J, Chang CL et al (2008) Origin of INSL3-mediated testicular descent in therian mammals. Genome Res 18:974–985
Patel SA, Simon MC (2008) Biology of hypoxia-inducible factor-2alpha in development and disease. Cell Death Differ 15:628–634
Peng J, Zhang L, Drysdale L et al (2000) The transcription factor EPAS-1/hypoxia-inducible factor 2alpha plays an important role in vascular remodeling. Proc Natl Acad Sci U S A 97:8386–8391
Raval RR, Lau KW, Tran MG et al (2005) Contrasting properties of hypoxia-inducible factor 1 (HIF-1) and HIF-2 in von Hippel-Lindau-associated renal cell carcinoma. Mol Cell Biol 25:5675–5686
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Ryan HE, Lo J, Johnson RS (1998) HIF-1 alpha is required for solid tumor formation and embryonic vascularization. EMBO J 17:3005–3015
Rytkonen KT, Ryynanen HJ, Nikinmaa M et al (2008) Variable patterns in the molecular evolution of the hypoxia-inducible factor-1 alpha (HIF-1alpha) gene in teleost fishes and mammals. Gene 420:1–10
Sanchez-Puig N, Veprintsev DB, Fersht AR (2005) Binding of natively unfolded HIF-1alpha ODD domain to p53. Mol Cell 17:11–21
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612
Taylor JS, Van de Peer Y, Braasch I et al (2001) Comparative genomics provides evidence for an ancient genome duplication event in fish. Philos Trans R Soc Lond B Biol Sci 356:1661–1679
Taylor JS, Braasch I, Frickey T et al (2003) Genome duplication, a trait shared by 22000 species of ray-finned fish. Genome Res 13:382–390
Thompson CE, Salzano FM, de Souza ON et al (2007) Sequence and structural aspects of the functional diversification of plant alcohol dehydrogenases. Gene 396:108–115
Tian H, Hammer RE, Matsumoto AM et al (1998) The hypoxia-responsive transcription factor EPAS1 is essential for catecholamine homeostasis and protection against heart failure during embryonic development. Genes Dev 12:3320–3324
Wagner A (2008) Neutralism and selectionism: a network-based reconciliation. Nat Rev Genet 9:965–974
Wang GL, Semenza GL (1995) Purification and characterization of hypoxia-inducible factor 1. J Biol Chem 270:1230–1237
Wong WS, Yang Z, Goldman N et al (2004) Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics 168:1041–1051
Yang Z (2002) Likelihood and Bayes estimation of ancestral population sizes in hominoids using data from multiple loci. Genetics 162:1811–1823
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Nielsen R (2002) Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol 19:908–917
Zhang J, Nielsen R, Yang Z (2005) Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol Biol Evol 22:2472–2479
Acknowledgments
This work is supported by the National High Technology Research and Development Program of China (863 project) (grant no. 2006AA10Z1E3), the National Natural Science Foundation of China (grant no. 30671492), and the National 973 Key Basic Research Program (grant no. 2006CB102102, 2004CB117502).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, X., Wang, M., Tan, G. et al. Molecular selection and functional divergence of HIF-α proteins in vertebrates. Genetica 138, 1241–1250 (2010). https://doi.org/10.1007/s10709-010-9523-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-010-9523-3