Abstract
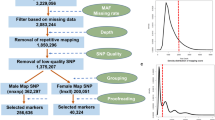
Hop is one of the few dioecious plants with dimorphic sex chromosomes. Because the entire Cannabaceae family is dioecious, hop and other members of this family are thought to have a relatively older sex chromosomal system than other plant species. Hop cones are only produced in female hops with or without fertilization. This has lead to most genomic research being directed toward female plants. The work we present provides genomic resources surrounding male plants. We have produced a draft genome for the male hop line USDA 21422M using a novel genome assembly method. In addition, we identified a 1.3 Mb set of scaffolds, which appear to be the male specific region based upon specificity with male hop accessions. This set includes a smaller high confidence total length 18 Kb set of scaffolds, which are supported by over 500 individuals, including the USDA world collection of hop varieties and two mapping populations, with genotyping by sequencing. We also have identified a portion of the Teamaker × 21422M linkage map to be associated with the pseudo-autosomal region (PAR). Within the genomic scaffolds, we identified a set of genes that are sex-linked and likely located in the PAR.




Similar content being viewed by others

References
Charlesworth B (2015) Causes of natural variation in fitness: evidence from studies of Drosophila populations. Proc Natl Acad Sci 112:1662–1669. doi:10.1073/pnas.1423275112
Danilova TV, Karlov GI (2006) Application of inter simple sequence repeat (ISSR) polymorphism for detection of sex-specific molecular markers in hop (Humulus lupulus L.). Euphytica 151:15–21
Divashuk MG, Alexandrov OS, Kroupin PY, Karlov GI (2011) Molecular cytogenetic mapping of humulus lupulus sex chromosomes. Cytogenet Genome Res 134:213–219
Divashuk MG, Alexandrov OS, Razumova OV, Kirov IV, Karlov GI (2014) Molecular cytogenetic characterization of the dioecious Cannabis sativa with an xy chromosome sex determination system. PLoS ONE 9:e85118. doi:10.1371/journal.pone.0085118
Dunn OJ (1961) Multiple comparisons among means. J Am Stat Assoc 56:52–64
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6:e19379
Eri S, Khoo BK, Lech J, Hartman TG (2000) Direct thermal desorption-gas chromatography and gas chromatography-mass spectrometry profiling of hop (Humulus lupulus L.) essential oils in support of varietal characterization. J Agric Food Chem 48:1140–1149
Filatov DA (2005) Substitution rates in a new Silene latifolia sex-linked gene. SlssX/Y Mol Biol Evol 22:402–408
Glaubitz JC, Casstevens TM, Lu F, Harriman J, Elshire RJ, Sun Q, Buckler ES (2014) TASSEL-GBS: a high capacity genotyping by sequencing analysis pipeline. PLoS One 9:E90346
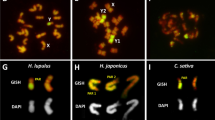
Grabowska-Joachimiak A, Mosiolek M, Lech A, Góralski G (2011) C-banding/DAPI and in situ hybridization reflect karyotype structure and sex chromosome differentiation in Humulus japonicus Siebold & Zucc. Cytogenet Genome Res 132:203–211
Henning JA, Gent DH, Twomey MC, Townsend MS, Pitra NJ, Matthews PD (2015) Precision QTL mapping of downy mildew resistance in hop (Humulus lupulus L.). Euphytica 202:487–498. doi:10.1007/s10681-015-1356-9
Hough J, Hollister JD, Wang W, Barrett SCH, Wright SI (2014) Genetic degeneration of old and young Y chromosomes in the flowering plant Rumex hastatulus. Proc Natl Acad Sci 111:7713–7718. doi:10.1073/pnas.1319227111
Jakse J, Stajner N, Kozjak P, Cerenak A, Javornik B (2008) Trinucleotide microsatellite repeat is tightly linked to male sex in hop (Humulus lupulus L.). Mol Breed 21:139–148. doi:10.1007/s11032-007-9114-x
Jiang H, Lei R, Ding S-W, Zhu S (2014) Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinform 15:182
Johnson M, Zaretskaya I, Raytselis Y, Merezhuk Y, McGinnis S, Madden TL (2008) NCBI BLAST: a better web interface. Nucleic Acids Res 36:W5–W9. doi:10.1093/nar/gkn201
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. doi:10.1093/bioinformatics/btp324
Liu Z et al (2004) A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427:348–352
Majer A, Javornik B, Cerenak A, Jakse J (2014) Development of novel EST-derived resistance gene markers in hop (Humulus lupulus L.). Mol Breed 33:61–74
McAdam EL et al (2013) Quantitative trait loci in hop (Humulus lupulus L.) reveal complex genetic architecture underlying variation in sex, yield and cone chemistry. BMC Genom 14:360
Ming R, Bendahmane A, Renner SS (2011) Sex chromosomes in land plants. Annu Rev Plant Biol 62:485–514. doi:10.1146/annurev-arplant-042110-103914
Natsume S et al (2015) The Draft Genome of Hop (Humulus lupulus), an Essence for Brewing. Plant Cell Physiol 56:428–441. doi:10.1093/pcp/pcu169
Ono T (1961) The hop native to Japan. Sasaki Printing and Publishing Company Ltd, Sendai
Otto SP et al (2011) About PAR: the distinct evolutionary dynamics of the pseudoautosomal region. Trends Genet 27:358–367
Oyama RK, Silber MV, Renner SS (2010) A specific insertion of a solo-LTR characterizes the Y-chromosome of Bryonia dioica (Cucurbitaceae). BMC research notes 3:166
Polley A, Ganal MW, Seigner E (1997) Identification of sex in hop (Humulus lupulus) using molecular markers. Genome 40:357–361
Russell GC, Guest JR (1991) Sequence similarities within the family of dihydrolipoamide acyltransferases and discovery of a previously unidentified fungal enzyme. Biochimica et Biophysica Acta (BBA) 1076:225–232
Seefelder S, Ehrmaier H, Schweizer G, Seigner E (2000) Male and female genetic linkage map of hops, Humulus lupulus. Plant Breeding 119:249–255
Shephard HL, Parker JS, Darby P, Ainsworth CC (2000) Sexual development and sex chromosomes in hop. New Phytol 148:397–411. doi:10.1046/j.1469-8137.2000.00771.x
Skaletsky H et al (2003) The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423:825–837
Xie Y et al (2014) SOAPdenovo-Trans: De novo transcriptome assembly with short RNA-Seq reads. Bioinformatics. doi:10.1093/bioinformatics/btu077
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using deBruijn graphs. Genome Res 18:821–829
Zhang W, Wang X, Yu Q, Ming R, Jiang J (2008) DNA methylation and heterochromatinization in the male-specific region of the primitive Y chromosome of papaya. Genome Res 18:1938–1943
Author information
Authors and Affiliations
Corresponding author
Additional information
D. Hendrix and J. A. Henning are co-investigators.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hill, S.T., Coggins, J., Liston, A. et al. Genomics of the hop pseudo-autosomal regions. Euphytica 209, 171–179 (2016). https://doi.org/10.1007/s10681-016-1655-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-016-1655-9