Abstract
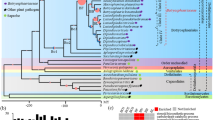
A survey of bacterial wilt in China collected 286 strains of Ralstonia solanacearum from 17 plant species in 13 Chinese provinces to investigate genetic diversity using the biovar (bv.) and phylotype classification schemes. A phylotype-specific multiplex-PCR showed that 198 isolates belonged to phylotype I (bv. 3, 4 and 5) and 68 to phylotype II (bv. 2 and bv. 1). A phylogenetic analysis examined the partial sequence of the egl and hrpB gene of all strains and the genetic diversity of 95 representatives was reported, demonstrating that Chinese strains are partitioned into phylotype I (Asia) and II (Americas). Phylotype I strains (historically typed bv. 3, 4 and 5), had considerable phylogenetic diversity, including 10 different sequevars: seven previously described sequevars 12 to 18 and three new sequevars: 34, 44 and 48. Chinese strains Z1, Z2, Z3, Z7, Pe74 and Tm82 were not genetically distinguishable from the edible ginger reference strain ACH92 (r4-bv. 4) for sequevar 16. This is believed to be the first report of this ginger group in China. All Chinese bv. 2 strains falling into the genetically and phenotypically diverse phylotype II were placed into phylotype IIB sequevar 1 (historically the Andean race3-bv. 2 potato brown rot agent). In both the egl and hrpB sequence-based trees, strains isolated from mulberry were present in two distinct branches found in sequevars 12 and 48 (reference strains R292 and M2, respectively).



Similar content being viewed by others
References
Buddenhagen, I., Sequeira, L., & Kelman, A. (1962). Designation of races in Pseudomonas solanacearum. Phytopathology, 52, 726.
Cook, D., & Sequeira, L. (1994). Strain differentiation of Pseudomonas solanacearum by molecular genetic methods. In A. C. Hayward & G. L. Hartman (Eds.), Bacterial wilt: The disease and its causative agent, Pseudomonas solanacearum (pp. 77–93). United Kingdom: CAB International.
Cook, D., Barlow, E., & Sequeira, L. (1989). Genetic diversity of Pseudomonas solanacearum: detection of restriction fragment polymorphisms with DNA probes that specify virulence and hypersensitive response. Molecular Plant-Microbe Interactions, 2, 113–121.
Elphinstone, J. G. (1996). Survival and possibilities for extinction of Pseudomonas solanacearum (Smith) Smith in cool climates. Potato Research, 39, 403–410.
Fegan, M., & Prior, P. (2005). How complex is the “Ralstonia solanacearum species complex”. In C. Allen, P. Prior & A. C. Hayward (Eds.), Bacterial wilt disease and the Ralstonia solanacearum species complex (pp. 449–462). St. Paul: APS.
Fegan, M., Taghavi, M., Sly, L. I., & Hayward, A. C. (1998). Phylogeny, diversity and molecular diagnostics of Ralstonia solanacearum. In P. Prior, C. Allen & J. Elphinstone (Eds.), Bacterial wilt disease: Molecular and ecological aspects (pp. 19–33). Paris: INRA Editions.
Gillings, M., Fahy, P., & Davies, C. (1993). Restriction analysis of an amplified polygalacturonase gene fragment differentiates strains of the phytopathogenic bacterium Pseudomonas solanacearum. Letters in Applied Microbiology, 17, 44–48.
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic acids symposium series, 41, 95–98.
Hayward, A. C. (1964). Characteristics of Pseudomonas solanacearum. Journal of Applied Bacteriology, 27, 265–277.
Hayward, A. C. (1991). Biology and epidemiology of bacterial wilt caused by Pseudomonas solanacearum. Annual Review of Phytopathology, 29, 67–87.
Hayward, A. C. (1994). Systematics and phylogeny of Pseudomonas solanacearum and related bacteria. In A. C. Hayward & G. L. Hartman (Eds.), Bacterial wilt: The disease and its causative agent, Pseudomonas solanacearum (pp. 123–135). United Kingdom: CAB International.
He, L.Y. (1985). Bacterial wilt in the People’s Republic of China. In: G.J. Perley (Eds.), Bacterial wilt disease in Asia and the south pacific (pp. 40–48). Proceedings of an international workshop held at PCARRD, Los Banos, Philippine, 8–10 October 1985.
He, L. Y., Sequeira, L., & Kelman, A. (1983). Characteristics of strains of Pseudomonas solanacearum. Plant Disease, 67, 1357–1361.
Jukes, T. H., & Cantor, C. R. (1969). Evolution of protein molecules. In H. N. Munro (Ed.), Mammalian protein metabolism (pp. 121–132). New York: Academic.
Kelman, A. (1954). The relationship of pathogenicity in Pseudomonas solanacearum to colony appearance on a tetrazolium chloride medium. Phytopathology, 44, 693–695.
Kumar, S., Tamura, K., & Nei, M. (2004). MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform, 5, 150–163.
Opina, N., Tavner, F., Hollway, G., Wang, J. F., Li, T. H., Maghirang, R., et al. (1997). A novel method for development of species and strain-specific DNA probes and PCR primers for identifying Burkholderia solanacearum. Asia and Pacific Journal of Molecular Biology and Biotechnology, 5, 19–30.
Page, R. D. M. (1996). TREEVIEW: an application to display phylogenetic trees on personal computers. Computer applications in the biosciences, 12, 357–358.
Pegg, K. G., & Moffett, M. (1971). Host range of the ginger strain of Pseudomonas solanacearum in Queensland. Australian Journal of Experimental Agriculture and Animal Husbandry, 11, 696–698.
Poussier, S., Vandewalle, P., & Luisetti, J. (1999). Genetic diversity of African and worldwide strains of Ralstonia solanacearum as determined by PCR-restriction fragment length polymorphism analysis of the hrp gene region. Applied and Environmental Microbiology, 65, 2184–2194.
Poussier, S., Prior, P., Luisetti, J., Hayward, C., & Fegan, M. (2000a). Partial sequencing of the hrpB and endoglucanase genes confirms and expands the known diversity within the Ralstonia solanacearum species complex. Systematic and Applied Microbiology, 23, 479–486.
Poussier, S., Trigalet-Demery, D., Vandewalle, P., Goffinet, B., Luisetti, J., & Trigalet, A. (2000b). Genetic diversity of Ralstonia solanacearum as assessed by PCR-RFLP of the hrp gene region, AFLP and 16S rRNA sequence analysis, and identification of an African subdivision. Microbiology, 146, 1679–1692.
Prior, P., & Fegan, M. (2005a). Recent development in the phylogeny and classification of Ralstonia solanacearum. Acta Horticulturae, 695, 127–136.
Prior, P., & Fegan, M. (2005b). Diversity and molecular detection of Ralstonia solanacearum race 2 strains by multiplex PCR. In C. Allen, P. Prior & A. C. Hayward (Eds.), Bacterial wilt disease and the Ralstonia solanacearum species complex (pp. 405–414). Madison: APS.
Rozas, J., Sanchez-DelBarrio, J. C., Messeguer, X., & Rozas, R. (2003). DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics, 19, 2496–2497.
Seal, S. E., Jackson, L. A., & Daniels, M. J. (1992). Use of tRNA consensus primers to indicate subgroups of Pseudomonas solanacearum by polymerase chain reaction amplification. Applied and Environmental Microbiology, 58, 3759–3761.
Smith, J. J., Offord, L. C., Holderness, M., & Saddler, G. S. (1995). Genetic diversity of Burkholderia solanacearum (synonym Pseudomonas solanacearum) race 3 in Kenya. Applied and Environmental Microbiology, 61, 4263–4268.
Taghavi, M., Hayward, C., Sly, L. I., & Fegan, M. (1996). Analysis of the phylogenetic relationships of strains of Burkholderia solanacearum, Pseudomonas syzygii, and the blood disease bacterium of banana based on 16S rRNA gene sequences. International Journal of Systematic Bacteriology, 46, 10–15.
Thompson, J. D., Higgins, D. G., & Gibson, T. J. (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Research, 22, 4673–4680.
Van der Wolf, J. M., Bonants, P. J. M., Smith, J. J., Hagenaar, M., Nijhuis, E., Van Beckhoven, J. R. C. M., et al. (1998). Genetic diversity of Ralstonia solanacearum race 3 in Western Europe determined by AFLP, RCPFGE and Rep-PCR. In P. Prior, C. Allen & J. Elphinstone (Eds.), Bacterial wilt disease: Molecular and ecological aspects (pp. 44–49). Paris: INRA Editions.
Villa, J. E., Tsuchiya, K., Horita, M., Natural, M., Opina, N., & Hyakumachi, M. (2005). Phylogenetic relationships of Ralstonia solanacearum species complex strains from Asia and other continents based on 16S rDNA, endoglucanase, and hrpB gene sequences. Journal of General Plant Pathology, 71, 39–46.
Wicker, E., Grassart, L., Coranson-Beaudu, R., Mian, D., Guilbaud, C., Fegan, M., et al. (2007). Ralstonia solanacearum Strains from Martinique (French West Indies) Exhibiting a New Pathogenic Potential. Applied and Environmental Microbiology, 71, 6790–6801.
Williamson, L., Nakaho, K., Hudelson, B., & Allen, C. (2002). Ralstonia solanacearum race 3, biovar 2 strains isolated from geranium are pathogenic on potato. Plant Disease, 86, 987–991.
Acknowledgements
This research was supported by the ‘National Basic Research and Development Programme (973 Programme) (No.2009CB119200)’, the ‘National Key Technologies Research and Development Programme (No. 2006 BAD08A14 & No. 2006 BAD08A15)’, the ‘National High Technology Research and Development Programme (863 Programme) (No.2006AA10Z432)’ and the ‘National Natural Science Foundation (No.30671418)’ of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
J. Xu and Z. C. Pan contributed equally towards this paper.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplemental Table 1
(DOC 122 kb)
Rights and permissions
About this article
Cite this article
Xu, J., Pan, Z.C., Prior, P. et al. Genetic diversity of Ralstonia solanacearum strains from China. Eur J Plant Pathol 125, 641–653 (2009). https://doi.org/10.1007/s10658-009-9512-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-009-9512-5