Abstract
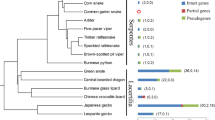
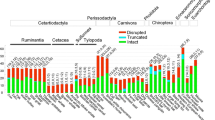
Bitter taste perception is important for vertebrates to select food and avoid toxic substances. A large number of Tas2r genes have been identified from vertebrate species previously; however, few studies have been conducted on the Tas2r genes of Ovalentaria species that have various dietary niches and are widely distributed, ranging from the sea to freshwater environments. Several genomes of Ovalentaria species have been released recently, allowing us to study Tas2r genes in these fishes. Thus, we explored the genomes of these fishes and identified 34 Tas2r genes in 21 species, including 27 intact Tas2r genes and seven pseudogenes. The results suggest that Ovalentaria species generally carry a small repertoire of Tas2r genes. To determine the phylogenetic relationship of Tas2r genes among 21 fishes, we constructed neighbor-joining (NJ) trees. The results showed that gene duplication may not occur in these fishes. Phylogenetic independent contrast (PIC) analysis showed that the fish Tas2r gene repertoire size was not positively correlated with diet, indicating that the food swallowing behavior might reduce the importance of bitter taste sense.




Similar content being viewed by others
Abbreviations
- Tas2r:
-
Bitter taste receptor
- NJ:
-
Neighbor-joining
- PIC:
-
Phylogenetically independent contrasts
- Calhm1:
-
calcium homeostasis modulator 1
References
Adler E, Hoon MA, Mueller KL, Chandrashekar J, Ryba NJP, Zuker CS (2000) A novel family of mammalian taste receptors. Cell 100(6):693–702. doi:10.1016/s0092-8674(00)80705-9
Albertson RC, Markert JA, Danley PD, Kocher TD (1999) Phylogeny of a rapidly evolving clade: the cichlid fishes of Lake Malawi, East Africa. Proc Natl Acad Sci U S A 96(9):5107. doi:10.1073/pnas.96.9.5107
Aoki M, Takao T, Takao K, Koike F, Suganuma N (2014) Lower expressions of the human bitter taste receptor TAS2R in smokers: reverse transcriptase-polymerase chain reaction analysis. Tob Induc Dis 12(1):12. doi:10.1186/1617-9625-12-12
Arthington AH (1989) Diet of Gambusia Affinis Holbrooki, Xiphophorus Helleri, X. Maculatus and Poecilia Reticulata (Pisces: Poeciliidae) in streams of southeastern Queensland, Australia. doi:10.1520/STP29045S
Bachmanov AA, Bosak NP, Lin C, Matsumoto I, Ohmoto M, Reed DR, Nelson TM (2013) Genetics of taste receptors. Curr Pharm Des 20(16):2669–2683. doi:10.1146/annurev.nutr.26.061505.111329
Barletta M, Blaber SJM, Craig JF (2016) Fish and aquatic habitat conservation in South America. J Fish Biol 89(1):1. doi:10.1111/jfb.13032
Behrens M, Korsching S, Meyerhof W (2014) Tuning properties of avian and frog bitter taste receptors dynamically fit gene repertoire sizes. Mol biol Evol:msu254. doi:10.1093/molbev/msu254
Booth DJ, Hixon MA (1999) Food ration and condition affect early survival of the coral reef damselfish, Stegastes Partitus. Oecologia 121(3):364–368. doi:10.1007/s004420050940
Bouton N, Os NV, Witte F (1998) Feeding performance of Lake Victoria rock cichlids: testing predictions from morphology. J Fish Biol 53(sA):118–127. doi:10.1111/j.1095-8649.1998.tb01022.x
Brawand D, Wagner CE, Li YI, Malinsky M, Keller I, Fan S, Simakov O, Ng AY, Lim ZW, Bezault E (2014) The genomic substrate for adaptive radiation in African cichlid fish. Nature 513(7518):375–381. doi:10.1038/nature13726
Breitholtz M, Hill C, Bengtsson BE (2016) Toxic substances and reproductive disorders in Baltic fish and crustaceans. Ambio 30(4–5):210. doi:10.1579/0044-7447-30.4.210
Chandrashekar J, Mueller KL, Hoon MA, Adler E, Feng L, Guo W, Zuker CS, Ryba NJP (2000) T2Rs function as bitter taste receptors. Cell 100(6):703–711. doi:10.1016/s0092-8674(00)80706-0
Cripe GM, Hansen DJ, Hansen DJ, Macauley SF, Macauley SF, Forester J (1986) Effects of diet quantity on sheepshead minnows (Cyprinodon Variegatus) during early life-stage exposures to chlorpyrifos. doi:10.1520/STP29045S
Darriba D, Taboada GL, Doallo R, Posada D (2011) ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27(8):1164–1165. doi:10.1093/bioinformatics/btr088
Dekoven DL, Núñez JM, Lester SM, Conklin DE, Marty GD, Parker LM, Hinton DE (1992) A purified diet for medaka (Oryzias Latipes): refining a fish model for toxicological research. Lab Anim Sci 42(2):180–189
Dussault GV, Kramer DL (1981) Food and feeding behavior of the guppy, Poecilia Reticulata (Pisces: Poeciliidae). Can J Zool 59(4):684–701. doi:10.1139/z81-098
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. Bmc Bioinformatics 5(4):1–19. doi:10.1093/nar/gkh340
Felsenstein J (1985) Phylogenies and the comparative method. Am Nat 125(1):1–15. doi:10.2307/2408678
Feng P, Zheng J, Rossiter SJ, Wang D, Zhao H (2014) Massive losses of taste receptor genes in toothed and baleen whales. Genome Biol Evol 6(6):1254–1265. doi:10.1093/gbe/evu095
Fink SV, Fink WL (1996) Interrelationships of the ostariophysan fishes (Teleostei). Interrelationships of Fishes 72(4):209–249. doi:10.1111/j.1096-3642.1981.tb01575.x
Hong W, Zhao H (2014) Vampire bats exhibit evolutionary reduction of bitter taste receptor genes common to other bats. Proc Biol Sci /Roy Soc 281(1788):20141079. doi:10.1098/rspb.2014.1079
Justi KC, Hayashi C, Visentainer JV, Souza NED, Matsushita M (2003) The influence of feed supply time on the fatty acid profile of Nile tilapia ( Oreochromis Niloticus ) fed on a diet enriched with n-3 fatty acids. Food Chem 80(4):489–493. doi:10.1016/S0308-8146(02)00317-5
Klingenberg CP, Barluenga M, Meyer A (2003) Body shape variation in cichlid fishes of the Amphilophus Citrinellus species complex. Biol J Linn Soc 80(3):397–408. doi:10.1046/j.1095-8312.2003.00246.x
Krogh A, Larsson B, Heijne GV, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden markov model: application to complete genomes 1. J Mol Biol 305(3):567–580. doi:10.1006/jmbi.2000.4315
Lampert KP, Lamatsch DK, Fischer P, Schartl M (2008) A tetraploid Amazon molly, Poecilia Formosa. J Hered 99(2):223–226. doi:10.1093/jhered/esm102
Li D, Zhang J (2014) Diet shapes the evolution of the vertebrate bitter taste receptor gene repertoire. Mol Biol Evol 31(2):303–309. doi:10.1093/molbev/mst219
Lindemann B (1996) Taste reception. Physiol Rev 76(3):718–766
Ling-Ling HU, Shi P (2013) Smallest bitter taste receptor (T2Rs) gene repertoire in carnivores. Zool Res 34(3):75–81. doi:10.1360/972009-1142
Liu Z, Liu G, Hailer F, Orozco-Terwengel P, Tan X, Tian J, Yan Z, Zhang B, Ming L (2016) Dietary specialization drives multiple independent losses and gains in the bitter taste gene repertoire of Laurasiatherian mammals. Front Zool 13(1):1–10. doi:10.1186/s12983-016-0161-1
Logan J, Haas H, Deegan L, Gaines E (2006) Turnover rates of nitrogen stable isotopes in the salt marsh mummichog, Fundulus Heteroclitus, following a laboratory diet switch. Oecologia 147(3):391–395. doi:10.1007/s00442-005-0277-z
Matsunami H, Montmayeur JP, Buck LB (2000) A family of candidate taste receptors in human and mouse. Nature 404(6778):601–604. doi:10.1038/35007072
Morais S (2016) The physiology of taste in fish: potential implications for feeding stimulation and gut chemical sensing. Rev Fish Sci Aquacult:1–17. doi:10.1080/2330849.2016.1249279
Naiman RJ (1979) Preliminary food studies of Cyprinodon Macularius and Cyprinodon Nevadensis (Cyprinodontidae). Southwest Nat 24(3):538. doi:10.2307/3671312
Nair GA, Balakrishnan N, Suryanarayanan H (1985) On the growth, food and conversion efficiency of a cichlid fish, Sarotherodon Mossambicus (Peters) exposed to sublethal concentrations of effluents from a titanium dioxide factory. J Anim Morphol Physiol 32:43–54
Nature GIUfCo, Resources N (1979) IUCN [International Union for Conservation of Nature and Natural Resources] annual report 1978–79
Oike H, Nagai T, Furuyama A, Okada S, Aihara Y, Ishimaru Y, Marui T, Matsumoto I, Misaka T, Abe K (2007) Characterization of ligands for fish taste receptors. J Neurosci Off J Soc Neurosci 27(21):5584–5592. doi:10.1523/JNEUROSCI.0651-07.2007
Paradis E, Claude J, Strimmer K (2004) APE: analyses of Phylogenetics and evolution in R language. Bioinformatics 20(2):289–290. doi:10.1093/bioinformatics/btg412
Parzefall J (2001) A review of morphological and Behavioural changes in the cave Molly, Poecilia Mexicana, from tabasco, Mexico. Environ Biol Fish 62(1):263–275. doi:10.1023/A:1011899817764
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Ther 4(6):406–425. doi:10.1093/oxfordjournals.molbev.a040454
Sanciangco MD, Carpenter KE, Betancur-R R (2015) Phylogenetic placement of enigmatic percomorph families (Teleostei: Percomorphaceae). Mol Phylogenet Evol 94(Pt B):565–576. doi:10.1016/j.ympev.2015.10.006
Shang S, Wu X, Chen J, Zhang H, Zhong H, Wei Q, Yan J, Li H, Liu G, Sha W, Zhang H (2017) The repertoire of bitter taste receptor genes in canids. Amino Acids:1–9. doi:10.1007/s00726-017-2422-5
Shi P, Zhang J (2006) Contrasting modes of evolution between vertebrate sweet/umami receptor genes and bitter receptor genes. Mol Biol Evol 23(2):292–300. doi:10.1093/molbev/msj028
Syed AS, Korsching SI (2014) Positive Darwinian selection in the singularly large taste receptor gene family of an ‘ancient’fish, Latimeria Chalumnae. BMC Genomics 15(1):1. doi:10.1186/1471-2164-15-650
Taruno A, Vingtdeux V, Ohmoto M, Ma Z, Dvoryanchikov G, Li A, Adrien L, Zhao H, Leung S, Abernethy M (2013) CALHM1 ion channel mediates purinergic neurotransmission of sweet, bitter and umami tastes. Nature 495(7440):223–226. doi:10.1038/nature11906
Terzibasi E, Lefrançois C, Domenici P, Hartmann N, Graf M, Cellerino A (2009) Effects of dietary restriction on mortality and age-related phenotypes in the short-lived fish Nothobranchius Furzeri. Aging Cell 8(2):88–99. doi:10.1111/j.1474-9726.2009.00455.x
Wang K, Zhao H (2015) Birds generally carry a small repertoire of bitter taste receptor genes. Genome Biol Evol 7(9):2705–2715. doi:10.1093/gbe/evv180
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24(8):1586–1591. doi:10.1093/molbev/msm088
Acknowledgements
This work was supported by the Special Fund for Forest Scientific Research in the Public Welfare (NO. 201404420), the National Natural Science Fund of China (NO. 31372220, 31672313). The data is obtained from the NCBI database, thus no Ethics Committees are needed in this research.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Shang, S., Zhang, H., Wu, X. et al. The repertoire of bitter taste receptor genes in Ovalentaria fish. Environ Biol Fish 100, 1489–1496 (2017). https://doi.org/10.1007/s10641-017-0659-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10641-017-0659-1