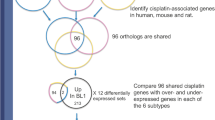
Summary
Introduction Given the high level of uncertainty surrounding the outcomes of early phase clinical trials, whole genome and transcriptome analysis (WGTA) can be used to optimize patient selection and study assignment. In this retrospective analysis, we reviewed the impact of this approach on one such program. Methods Patients with advanced malignancies underwent fresh tumor biopsies as part of our personalized medicine program (NCT02155621). Tumour molecular data were reviewed for potentially clinically actionable findings and patients were referred to the developmental therapeutics program. Outcomes were reviewed in all patients, including those where trial selection was driven by molecular data (matched) and those where there was no clear molecular rationale (unmatched). Results From January 2014 to January 2018, 28 patients underwent WGTA and enrolled in clinical trials, including 2 patients enrolled in two trials. Fifteen patients were matched to a treatment based on a molecular target. Five patients were matched to a trial based upon single-gene DNA changes, all supported by RNA data. Ten cases were matched on the basis of genome-wide data (n = 4) or RNA gene expression only (n = 6). With a median follow-up of 6.7 months, the median time on treatment was 8.2 weeks. Discussion When compared to single-gene DNA-based data alone, WGTA led to a 3-fold increase in treatment matching. In a setting where there is a high level of uncertainty around both the investigational agents and the biomarkers, more data are needed to fully evaluate the impact of routine use of WGTA.


Similar content being viewed by others
References
Thompson MA, Godden JJ, Weissman SM, Wham D, Wilson A, Ruggeri A, et al. Implementing an oncology precision medicine clinic in a large community health system. Am J Manag Care. 2017;23(10 Spec No.):SP425–7
Tannock IF, Hickman JA (2016) Limits to personalized Cancer medicine. N Engl J Med 375(13):1289–1294
Laskin J, Jones S, Aparicio S, Chia S, Ch'ng C, Deyell R et al (2015) Lessons learned from the application of whole-genome analysis to the treatment of patients with advanced cancers. Cold Spring Harb Mol Case Stud 1(1):a000570
Mateo J, Chakravarty D, Dienstmann R, Jezdic S, Gonzalez-Perez A, Lopez-Bigas N et al (2018) A framework to rank genomic alterations as targets for cancer precision medicine: the ESMO scale for clinical Actionability of molecular targets (ESCAT). Ann Oncol 29(9):1895–1902
Tuxen IV, Rohrberg KS, Oestrup O, Ahlborn LB, Schmidt AY, Spanggaard I et al (2019) Copenhagen prospective personalized oncology (CoPPO)-clinical utility of using molecular profiling to select patients to phase I trials. Clin Cancer Res 25(4):1239–1247
Rothwell DG, Ayub M, Cook N, Thistlethwaite F, Carter L, Dean E et al (2019) Utility of ctDNA to support patient selection for early phase clinical trials: the TARGET study. Nat Med 22
Le Tourneau C, Delord J, Gonçalves A, Gavoille C, Dubot C, Isambert N et al (2015) Molecularly targeted therapy based on tumour molecular profiling versus conventional therapy for advanced cancer (SHIVA): a multicentre, open-label, proof-of-concept, randomised, controlled phase 2 trial. Lancet Oncol 16(13):1324–1334
Tsimberidou AM, Kurzrock R (2015) Precision medicine: lessons learned from the SHIVA trial. Lancet Oncol. 16(16):e579–e580
van der Velden DL, Hoes LR, van der Wijngaart H, van Berge Henegouwen JM, van Werkhoven E, Roepman P et al (2019) The drug rediscovery protocol facilitates the expanded use of existing anticancer drugs. Nature 574(7776):127–131
Funding
The BC Personalized OncoGenomics project is supported by the BC Cancer Foundation. JML is supported by the University of British Columbia Clinician Investigator Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
JML received honoraria from Pfizer and Astellas. TM declares that she has no conflict of interest. SEL declares that she has no conflict of interest. BD declares that she has no conflict of interest. SKC received honoraria from Novartis, Hoffmann-La Roche, Pfizer, and AstraZeneca. KAG received honoraria from Pfizer, AstraZeneca, Novartis, Lilly, Genomic Health, NanoString, Merck, Mylan, Roche, BMS and Genentech. CKK received honoraria from Pfizer, Astellas, BMS, Ipsen, Eisai, Roche, Merck and Janssen. AVT received honoraria from AstraZeneca. SJMJ declares that he has no conflict of interest. MM declares that he has no conflict of interest. JL received honoraria from Roche, BI and Astra Zeneca. DJR received research funding and honoraria from Bayer, and honoraria from Servier, Celgene, Taiho and Ipsen.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study as part of Personalized OncoGenomics.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lavoie, JM., Mitchell, T., Lee, SE. et al. Patient selection for a developmental therapeutics program using whole genome and Transcriptome analysis. Invest New Drugs 38, 1601–1604 (2020). https://doi.org/10.1007/s10637-020-00892-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-020-00892-8