Abstract
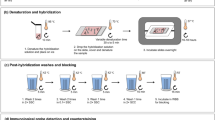
Tandemly repetitive DNA sequences, also named satellite repeats, are major DNA components of heterochromatin and are often organized as long arrays in the pericentromeric, centromeric, and subtelomeric regions of eukaryotic chromosomes. An increasing amount of evidence indicates that transcripts derived from some satellite repeats play important roles in various biological functions. We used a RNA-fluorescence in situ hybridization (RNA-FISH) technique to investigate the transcription of the four well-characterized satellite repeats of maize (Zea mays), including the 180-bp knob repeat, the telomeric (TTTAGGG)n repeat, the 156-bp centromeric repeat CentC, and a 350-bp subtelomeric repeat. Although few transcripts derived from these four repeats were found in the expressed sequence tag and RNA-seq databases, RNA-FISH consistently detected the transcripts from three of the four repeats on interphase nuclei, suggesting that the transcripts from the three repeats are largely integrated into chromatin. The transcripts from the knob and telomeric repeats were mapped to the related DNA loci. In contrast, the transcripts from the CentC repeats were mainly localized to the nucleolus, although nucleoplasmic CentC transcripts were also detectable. The nucleolus and nuclear RNAs appeared to be important for the nuclear localization for at least one centromeric protein, Mis12. We demonstrate that RNA-FISH is a powerful tool to assess the level of transcription as well as to physically map the nuclear locations of the transcripts derived from satellite repeats.







Similar content being viewed by others
Abbreviations
- CENP:
-
Centromere protein
- EST:
-
Expressed sequence tags
- FISH:
-
Fluorescence in situ hybridization
- TERRA:
-
Telomeric repeat-containing RNA
References
Ananiev EV, Phillips RL, Rines HW (1998) Chromosome-specific molecular organization of maize (Zea mays L.) centromeric regions. Proc Natl Acad Sci 95:13073–13078
Azzalin CM, Reichenbach P, Khoriauli L, Giulotto E, Lingner J (2007) Telomeric repeat-containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318:798–801
Biscotti MA, Canapa A, Forconi M, Olmo E, Barucca M (2015) Transcription of tandemly repetitive DNA: functional roles. Chromosom Res 23:463–477
Bouzinba-Segard H, Guais A, Francastel C (2006) Accumulation of small murine minor satellite transcripts leads to impaired centromeric architecture and function. Proc Nat Acad Sci USA 103:8709–8714
Burr B, Burr FA, Matz EC, Romeroseverson J (1992) Pinning down loose ends: mapping telomeres and factors affecting their length. Plant Cell 4:953–960
Chan FL, Marshall OJ, Saffery R, Kim BW, Earle E, et al. (2012) Active transcription and essential role of RNA polymerase II at the centromere during mitosis. Proc Nat Acad Sci USA 109:1979–1984
Cheeseman IM, Desai A (2008) Molecular architecture of the kinetochore-microtubule interface. Nat Rev Mol Cell Biol 9:33–46
Chen ES, Zhang K, Nicolas E, Cam HP, Zofall M, et al. (2008) Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451:734–737
Cheng ZK, Stupar RM, Gu MH, Jiang JM (2001) A tandemly repeated DNA sequence is associated with both knob-like heterochromatin and a highly decondensed structure in the meiotic pachytene chromosomes of rice. Chromosoma 110:24–31
Chun Y, Park B, Koh W, Lee S, Cheon Y, et al. (2011) New centromeric component CENP-W is an RNA-associated nuclear matrix protein that interacts with nucleophosmin/B23 protein. J Biol Chemi 286:42758–42769
Cox KH, Deleon DV, Angerer LM, Angerer RC (1984) Detection of mRNAs in sea urchin embryos by in situ hybridization using asymmetric RNA probes. Dev Biol 101:485–502
Du YQ, Topp CN, Dawe RK (2010) DNA binding of centromere protein C (CENPC) is stabilized by single stranded RNA. PLoS Genet 6:e1000835
Eymery A, Horard B, El Atifi-Borel M, Fourel G, Berger F, et al. (2009) A transcriptomic analysis of human centromeric and pericentric sequences in normal and tumor cells. Nucleic Acids Res 37:6340–6354
Fajkus J, Sykorova E, Leitch AR (2005) Telomeres in evolution and evolution of telomeres. Chromosom Res 13:469–479
Ferri F, Bouzinba-Segard H, Velasco G, Hube F, Francastel C (2009) Non-coding murine centromeric transcripts associate with and potentiate Aurora B kinase. Nucleic Acids Res 37:5071–5080
Galbraith DW, Harkins KR, Maddox JM, Ayres NM, Sharma DP, et al. (1983) Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 220:1049–1051
Gall JG, Atherton DD (1974) Satellite DNA sequences in Drosophila virilis. J Mol Biol 85:633–664
Garrido-Ramos MA (2015) Satellite DNA in plants: more than just rubbish. Cytogenetic Genome Res 146:153–170
Gaubatz JW, Cutler RG (1990) Mouse satellite DNA is transcribed in senescent cardiac muscle. J Biol Chemi 265:17753–17758
Grewal SIS, Elgin SCR (2007) Transcription and RNA interference in the formation of heterochromatin. Nature 447:399–406
He L, Liu J, Torres GA, Zhang HQ, Jiang JM, et al. (2013) Interstitial telomeric repeats are enriched in the centromeres of chromosomes in Solanum species. Chromosom Res 21:5–13
Henikoff S, Ahmad K, Malik HS (2001) The centromere paradox: stable inheritance with rapidly evolving DNA. Science 293:1098–1102
Heslop-Harrison JS, Schwarzacher T (2011) Organisation of the plant genome in chromosomes. Plant J 66:18–33
Highett MI, Beven AF, Shaw PJ (1993) Localization of 5S genes and transcripts in Pisum sativum nuclei. J Cell Sci 105:1151–1158
Jiang JM, Gill BS (2006) Current status and the future of fluorescence in situ hybridization (FISH) in plant genome research. Genome 49:1057–1068
Jiang JM, Birchler JA, Parrott WA, Dawe RK (2003) A molecular view of plant centromeres. Trends Plant Sci 8:570–575
Jin WW, Melo JR, Nagaki K, Talbert PB, Henikoff S, et al. (2004) Maize centromeres: organization and functional adaptation in the genetic background of oat. Plant Cell 16:571–581
Jolly C, Mongelard F, RobertNicoud M, Vourch C (1997) Optimization of nuclear transcript detection by FISH and combination with fluorescence immunocytochemical detection of transcription factors. J Histochemi & Cytochemi 45:1585–1592
Jolly C, Metz A, Govin J, Vigneron M, Turner BM, et al. (2004) Stress-induced transcription of satellite III repeats. J Cell Biol 164:25–33
Kloc A, Zaratiegui M, Nora E, Martienssen R (2008) RNA interference guides histone modification during the S phase of chromosomal replication. Curr Biol 18:490–495
Lee HR, Neumann P, Macas J, Jiang JM (2006) Transcription and evolutionary dynamics of the centromeric satellite repeat CentO in rice. Mol Biol Evol 23:2505–2520
Li XX, Dawe RK (2009) Fused sister kinetochores initiate the reductional division in meiosis I. Nat Cell Biol 11:1103–1108
Li J, Yang F, Zhu J, He SB, Li LJ (2009) Characterization of a tandemly repeated subtelomeric sequence with inverted telomere repeats in maize. Genome 52:286–293
Lu J, Gilbert DM (2007) Proliferation-dependent and cell cycle-regulated transcription of mouse pericentric heterochromatin. J Cell Biol 179:411–421
Luke B, Lingner J (2009) TERRA: telomeric repeat-containing RNA. EMBO J 28:2503–2510
Maison C, Bailly D, Peters AHFM, Quivy JP, Roche D, et al. (2002) Higher-order structure in pericentric heterochromatin involves a distinct pattern of histone modification and an RNA component. Nat Genet 30:329–334
Majerova E, Fojtova M, Mozgova I, Bittova M, Fajkus J (2011) Hypomethylating drugs efficiently decrease cytosine methylation in telomeric DNA and activate telomerase without affecting telomere lengths in tobacco cells. Plant Mol Biol 77:371–380
May BP, Lippman ZB, Fang YD, Spector DL, Martienssen RA (2005) Differential regulation of strand-specific transcripts from Arabidopsis centromeric satellite repeats. PLoS Genet 1:705–714
McClintock B (1929) Chromosome morphology in Zea mays. Science 69:629
McKnight TD, Riha K, Shippen DE (2002) Telomeres, telomerase, and stability of the plant genome. Plant Mol Biol 48:331–337
Peacock WJ, Dennis ES, Rhoades MM, Pryor AJ (1981) Highly repeated DNA sequence limited to knob heterochromatin in maize. Proc Nat Acad Sci USA 78:4490–4494
Pezer Z, Ugarkovic D (2012) Satellite DNA-associated siRNAs as mediators of heat shock response in insects. RNA Biol 9:587–595
Pontes O, Li CF, Nunes PC, Haag J, Ream T, et al. (2006) The Arabidopsis chromatin-modifying nuclear siRNA pathway involves a nucleolar RNA processing center. Cell 126:79–92
Pontes O, Costa-Nunes P, Vithayathil P, Pikaard CS (2009) RNA polymerase V functions in Arabidopsis interphase heterochromatin organization independently of the 24-nt siRNA-directed DNA methylation pathway. Mol Plant 2:700–710
Prieto P, Moore G, Shaw P (2007) Fluorescence in situ hybridization on vibratome sections of plant tissues. Nat Protoc 2:1831–1838
Probst AV, Okamoto I, Casanova M, El Marjou F, Le Baccon P, et al. (2010) A strand-specific burst in transcription of pericentric satellites is required for chromocenter formation and early mouse development. Dev Cell 19:625–638
Rivin CJ, Cullis CA, Walbot V (1986) Evaluating quantitative variation in the genome of Zea mays. Genetics 113:1009–1019
Rosic S, Kohler F, Erhardt S (2014) Repetitive centromeric satellite RNA is essential for kinetochore formation and cell division. J Cell Biol 207:335–349
Santos AP, Wegel E, Allen GC, Thompson WF, Stoger E, et al. (2006) In situ methods to localize transgenes and transcripts in interphase nuclei: a tool for transgenic plant research. Plant Methods 2:18
Schnable PS, Ware D, Fulton RS, Stein JC, Wei FS, et al. (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115
Schoeftner S, Blasco MA (2008) Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nat Cell Biol 10:228–236
Sippel AE, Hynes N, Groner B, Schutz G (1977) Frequency distribution of messenger sequences within polysomal messenger RNA and nuclear RNA from rat liver. Eur J Biochem 77:141–151
Ting DT, Lipson D, Paul S, Brannigan BW, Akhavanfard S, et al. (2011) Aberrant overexpression of satellite repeats in pancreatic and other epithelial cancers. Science 331:593–596
Topp CN, Zhong CX, Dawe RK (2004) Centromere-encoded RNAs are integral components of the maize kinetochore. Proc Natl Acad Sci U S A 101:15986–15991
Trofimova I, Popova D, Vasilevskaya E, Krasikova A (2014) Non-coding RNA derived from a conservative subtelomeric tandem repeat in chicken and Japanese quail somatic cells. Mol Cytogenet 7:102
Ugarkovic D (2005) Functional elements residing within satellite DNAs. EMBO Rep 6:1035–1039
Vrbsky J, Akimcheva S, Watson JM, Turner TL, Daxinger L, et al. (2010) siRNA-mediated methylation of I telomeres. PLoS Genet 6:e1000986
Werner MS, Ruthenburg AJ (2015) Nuclear fractionation reveals thousands of chromatin-tethered noncoding RNAs adjacent to active genes. Cell Rep 12:1089–1098
Wong LH, Brettingham-Moore KH, Chan L, Quach JM, Anderson MA, et al. (2007) Centromere RNA is a key component for the assembly of nucleoproteins at the nucleolus and centromere. Genome Res 17:1146–1160
Wu TY, Wang YX, Wu R (1994) Transcribed repetitive DNA-sequences in telomeric regions of rice (Oryza sativa). Plant Mol Biol 26:363–375
Zhong CX, Marshall JB, Topp C, Mroczek R, Kato A, et al. (2002) Centromeric retroelements and satellites interact with maize kinetochore protein CENH3. Plant Cell 14:2825–2836
Zhu Q, Pao GM, Huynh AM, Suh H, Tonnu N, et al. (2011) BRCA1 tumour suppression occurs via heterochromatin-mediated silencing. Nature 477:179–184
Acknowledgments
This work was supported by grants IOS-0922703 and IOS-1444514 from the National Science Foundation to J.J.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Hans de Jong
Electronic supplementary material
ESM 1
(PDF 1979 kb)
Rights and permissions
About this article
Cite this article
Koo, DH., Zhao, H. & Jiang, J. Chromatin-associated transcripts of tandemly repetitive DNA sequences revealed by RNA-FISH. Chromosome Res 24, 467–480 (2016). https://doi.org/10.1007/s10577-016-9537-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-016-9537-5