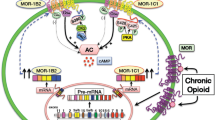
Abstract
The biased signaling has been extensively studied in the original mu opioid receptor (MOR-1), particularly through G protein and β-arrestin2 signaling pathways. The concept that the G protein pathway is often linked to the therapeutic effect of the drug, while the β-arrestin pathway is associated to the side effects has been proposed to develop biased analgesic compounds with limited side-effects associated with traditional opiates. The mu opioid receptor gene, OPRM1, undergoes extensive alternative pre-mRNA splicing, generating multiple splice variants or isoforms that are conserved from rodent to human. One type of the Oprm1 splice variants are the full-length 7 transmembrane (7TM) C-terminal splice variants, which have identical receptor structures including entire binding pocket, but contain a different intracellular C-terminal tail resulted from 3′ alternative splicing. Increasing evidence suggest that these full-length 7TM C-terminal variants play important roles in mu opioid pharmacology, raising questions regarding biased signaling at these multiple C-terminal variants. In the present study, we investigated the effect of different C-terminal variants on mu agonist-induced G protein coupling, β-arrestin2 recruitment, and ultimately, signaling bias. We found that mu agonists produced marked differences in G protein activation and β-arrestin2 recruitment among various C-terminal variants, leading to biased signaling at various level. Particularly, MOR-1O, an exon 7-associated variant, showed greater β-arrestin2 bias for most mu agonists than MOR-1, an exon 4-associated variant. Biased signaling of G protein-coupled receptors has been defined by evidences that different agonists can produce divergent signaling transduction pathways through a single receptor. Our findings that a single mu agonist can induce differential signaling through multiple 7TM splice variants provide a new perspective on biased signaling at least for Oprm1, which perhaps is important for our understanding of the complex mu opioid actions in vivo where all the 7TM splice variants co-exist.





Similar content being viewed by others
Change history
31 October 2020
A Correction to this paper has been published: https://doi.org/10.1007/s10571-020-00990-4
References
Abbadie C, Pan Y-X, Drake CT, Pasternak GW (2000a) Comparative immunhistochemical distributions of carboxy terminus epitopes from the mu opioid receptor splice variants MOR-1D, MOR-1 and MOR-1C in the mouse and rat central nervous systems. Neuroscience 100:141–153
Abbadie C, Pan Y-X, Pasternak GW (2000b) Differential distribution in rat brain of mu opioid receptor carboxy terminal splice variants MOR-1C and MOR-1-like immunoreactivity: evidence for region-specific processing. J Comp Neurol 419:244–256
Abbadie C, Pasternak GW (2001) Differential in vivo internalization of MOR-1 and MOR-1C by morphine. NeuroReport 12:3069–3072
Barak LS, Ferguson SS, Zhang J, Caron MG (1997) A beta-arrestin/green fluorescent protein biosensor for detecting G protein-coupled receptor activation. J Biol Chem 272:27497–27500. https://doi.org/10.1074/jbc.272.44.27497
Bertrand L et al (2002) The BRET2/arrestin assay in stable recombinant cells: a platform to screen for compounds that interact with G protein-coupled receptors (GPCRS). J Recept Signal Transduct Res 22:533–541. https://doi.org/10.1081/rrs-120014619
Bohn LM, Gainetdinov RR, Lin FT, Lefkowitz RJ, Caron MG (2000) Mu-opioid receptor desensitization by beta-arrestin-2 determines morphine tolerance but not dependence. Nature 408:720–723
Bohn LM, Gainetdinov RR, Sotnikova TD, Medvedev IO, Lefkowitz RJ, Dykstra LA, Caron MG (2003) Enhanced rewarding properties of morphine, but not cocaine, in beta(arrestin)-2 knock-out mice. J Neurosci 23:10265–10273
Bohn LM, Lefkowitz RJ, Caron MG (2002) Differential mechanisms of morphine antinociceptive tolerance revealed in (beta)arrestin-2 knock-out mice. J Neurosci 22:10494–10500
Bolan EA, Pan YX, Pasternak GW (2004) Functional analysis of MOR-1 splice variants of the mouse mu opioid receptor gene. Oprm Synapse 51:11–18
Brown TG, Xu J, Hurd YL, Pan YX (2020) Dysregulated expression of the alternatively spliced variant mRNAs of the mu opioid receptor gene, OPRM1, in the medial prefrontal cortex of male human heroin abusers and heroin self-administering male rats. J Neurosci Res. https://doi.org/10.1002/jnr.24640
Celver JP, Lowe J, Kovoor A, Gurevich VV, Chavkin C (2001) Threonine 180 is required for G-protein-coupled receptor kinase 3- and beta -arrestin 2-mediated desensitization of the {micro}-opioid receptor in xenopus oocytes. J Biol Chem 276:4894–4900
Chakrabarti S, Liu NJ, Gintzler AR (2020) Phosphorylation of unique C-terminal sites of the mu-opioid receptor variants 1B2 and 1C1 influences their Gs association following chronic morphine. J Neurochem 152:449–467. https://doi.org/10.1111/jnc.14863
Chavkin C, McLaughlin JP, Celver JP (2001) Regulation of opioid receptor function by chronic agonist exposure: constitutive activity and desensitization. Mol Pharmacol 60:20–25
Dever SM et al (2014) Differential expression of the alternatively spliced OPRM1 isoform mu-opioid receptor-1K in HIV-infected individuals. AIDS 28:19–30. https://doi.org/10.1097/QAD.0000000000000113
Dever SM, Xu R, Fitting S, Knapp PE, Hauser KF (2012) Differential expression and HIV-1 regulation of mu-opioid receptor splice variants across human central nervous system cell types. J Neurovirol 18:181–190. https://doi.org/10.1007/s13365-012-0096-z
Dixon AS et al (2016) NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem Biol 11:400–408. https://doi.org/10.1021/acschembio.5b00753
Doyle GA et al (2007) Identification of five mouse mu-opioid receptor (MOR) gene (Oprm1) splice variants containing a newly identified alternatively spliced exon. Gene 395:98–107
Eglen RM, Singh R (2003) Beta galactosidase enzyme fragment complementation as a novel technology for high throughput screening. Comb Chem High Throughput Screen 6:381–387. https://doi.org/10.2174/138620703106298473
Ehrlich AT et al (2019) Biased signaling of the Mu opioid receptor revealed in native neurons. iScience 14:47–57. https://doi.org/10.1016/j.isci.2019.03.011
Gandini MA, Souza IA, Raval D, Xu J, Pan YX, Zamponi GW (2019) Differential regulation of Cav2.2 channel exon 37 variants by alternatively spliced mu-opioid receptors. Mol Brain 12:98. https://doi.org/10.1186/s13041-019-0524-6
Grim TW, Acevedo-Canabal A, Bohn LM (2020) Toward directing opioid receptor signaling to refine opioid therapeutics. Biol Psychiatry 87:15–21. https://doi.org/10.1016/j.biopsych.2019.10.020
Grinnell SG et al (2016) Mediation of buprenorphine analgesia by a combination of traditional and truncated mu opioid receptor splice variants. Synapse 70:395–407. https://doi.org/10.1002/syn.21914
Hamdan FF, Audet M, Garneau P, Pelletier J, Bouvier M (2005) High-throughput screening of G protein-coupled receptor antagonists using a bioluminescence resonance energy transfer 1-based beta-arrestin2 recruitment assay. J Biomol Screening 10:463–475. https://doi.org/10.1177/1087057105275344
Hurle MA, Sanchez A, Garcia-Sancho J (1999) Effects of k- and m-opioid receptor agonists on Ca2+ channels in neuroblastoma cells: involvement of the orphan opioid receptor. Eur J Pharmacol 379:191–198
Kenakin T, Christopoulos A (2013) Signalling bias in new drug discovery: detection, quantification and therapeutic impact. Nat Rev Drug Discov 12:205–216. https://doi.org/10.1038/nrd3954
Kenakin T, Watson C, Muniz-Medina V, Christopoulos A, Novick S (2012) A simple method for quantifying functional selectivity and agonist bias. ACS Chem Neurosci 3:193–203. https://doi.org/10.1021/cn200111m
Koch T, Schulz S, Pfeiffer M, Klutzny M, Schroder H, Kahl E, Hollt V (2001) C-terminal splice variants of the mouse mu-opioid receptor differ in morphine-induced internalization and receptor resensitization. J Biol Chem 276:31408–31414
Koch T, Schulz S, Schroder H, Wolf R, Raulf E, Hollt V (1998) Carboxyl-terminal splicing of the rat mu opioid receptor modulates agonist-mediated internalization and receptor resensitization. J Biol Chem 273:13652–13657
Koehl A et al (2018) Structure of the mu-opioid receptor-G(i) protein complex. Nature 558:547. https://doi.org/10.1038/s41586-018-0219-7
Majumdar S et al (2011) Truncated G protein-coupled mu opioid receptor MOR-1 splice variants are targets for highly potent opioid analgesics lacking side effects. Proc Natl Acad Sci USA 108:19778–19783. https://doi.org/10.1073/pnas.1115231108
Majumdar S et al (2012) Synthesis and evaluation of aryl-naloxamide opiate analgesics targeting truncated exon 11-associated mu opioid receptor (MOR-1) splice variants. J Med Chem 55:6352–6362. https://doi.org/10.1021/jm300305c
Manglik A et al (2012) Crystal structure of the mu-opioid receptor bound to a morphinan antagonist. Nature 485:321–U170. https://doi.org/10.1038/nature10954
Mann A, Illing S, Miess E, Schulz S (2015) Different mechanisms of homologous and heterologous mu-opioid receptor phosphorylation. Br J Pharmacol 172:311–316. https://doi.org/10.1111/bph.12627
McPherson J et al (2010) mu-opioid receptors: correlation of agonist efficacy for signalling with ability to activate internalization. Mol Pharmacol 78:756–766. https://doi.org/10.1124/mol.110.066613
Molinari P et al (2010) Morphine-like opiates selectively antagonize receptor-arrestin interactions. J Biol Chem 285:12522–12535. https://doi.org/10.1074/jbc.M109.059410
Pan YX (2005) Diversity and complexity of the mu opioid receptor gene: alternative pre-mRNA splicing and promoters. DNA Cell Biol 24:736–750
Pan YX et al (1999) Identification and characterization of three new alternatively spliced mu-opioid receptor isoforms. Mol Pharmacol 56:396–403
Pan YX, Xu J, Bolan E, Moskowitz HS, Xu M, Pasternak GW (2005a) Identification of four novel exon 5 splice variants of the mouse mu-opioid receptor gene: functional consequences of C-terminal splicing. Mol Pharmacol 68:866–875
Pan L, Xu J, Yu R, Xu MM, Pan YX, Pasternak GW (2005b) Identification and characterization of six new alternatively spliced variants of the human mu opioid receptor gene. Oprm Neuroscience 133:209–220
Pan YX, Xu J, Xu M, Rossi GC, Matulonis JE, Pasternak GW (2009) Involvement of exon 11-associated variants of the mu opioid receptor MOR-1 in heroin, but not morphine, actions. Proc Natl Acad Sci 106:4917–4922
Pasternak DA, Pan L, Xu J, Yu R, Xu MM, Pasternak GW, Pan YX (2004) Identification of three new alternatively spliced variants of the rat mu opioid receptor gene: dissociation of affinity and efficacy. J Neurochem 91:881–890
Pasternak GW, Childers SR, Pan YX (2020) Emerging insights into mu opioid pharmacology. Handb Exp Pharmacol 258:89–125. https://doi.org/10.1007/164_2019_270
Pasternak GW, Pan YX (2013) Mu opioids and their receptors: evolution of a concept. Pharmacol Rev 65:1257–1317. https://doi.org/10.1124/pr.112.007138
Pedersen MF et al (2020) Biased agonism of clinically approved mu-opioid receptor agonists and TRV130 is not controlled by binding and signaling kinetics. Neuropharmacology 166:107718. https://doi.org/10.1016/j.neuropharm.2019.107718
Raehal KM, Walker JK, Bohn LM (2005) Morphine side effects in beta-arrestin 2 knockout mice. J Pharmacol Exp Ther 314:1195–1201. https://doi.org/10.1124/jpet.105.087254
Schmid CL et al (2017) Bias factor and therapeutic window correlate to predict safer opioid analgesics. Cell 171:1165. https://doi.org/10.1016/j.cell.2017.10.035
Tanowitz M, Hislop JN (2008) Alternative splicing determines the post-endocytic sorting fate of G-protein-coupled receptors. J Biol Chem 283:35614–35621. https://doi.org/10.1074/jbc.M806588200
van der Westhuizen ET, Breton B, Christopoulos A, Bouvier M (2014) Quantification of ligand bias for clinically relevant beta2-adrenergic receptor ligands: implications for drug taxonomy. Mol Pharmacol 85:492–509. https://doi.org/10.1124/mol.113.088880
Verzillo V, Madia PA, Liu NJ, Chakrabarti S, Gintzler AR (2014) Mu-opioid receptor splice variants: sex-dependent regulation by chronic morphine. J Neurochem 130:790–796. https://doi.org/10.1111/jnc.12768
Williams JT et al (2013) Regulation of mu-opioid receptors: desensitization, phosphorylation, internalization, and tolerance. Pharmacol Rev 65:223–254. https://doi.org/10.1124/pr.112.005942
Xu J, Faskowitz AJ, Rossi GC, Xu M, Lu Z, Pan YX, Pasternak GW (2015) Stabilization of morphine tolerance with long-term dosing: Association with selective upregulation of mu-opioid receptor splice variant mRNAs. Proc Natl Acad Sci USA 112:279–284. https://doi.org/10.1073/pnas.1419183112
Xu J et al (2017) Alternatively spliced mu opioid receptor C termini impact the diverse actions of morphine. J Clin Investig 127:1561–1573. https://doi.org/10.1172/JCI88760
Xu J et al (2014) Differential expressions of the alternatively spliced variant mRNAs of the micro opioid receptor gene, OPRM1, in brain regions of four inbred mouse strains. PLoS ONE 9:e111267. https://doi.org/10.1371/journal.pone.0111267
Xu J et al (2013) Stabilization of the mu-opioid receptor by truncated single transmembrane splice variants through a chaperone-like action. J Biol Chem 288:21211–21227. https://doi.org/10.1074/jbc.M113.458687
Zhou XE et al (2017) Identification of phosphorylation codes for arrestin recruitment by G protein-coupled receptors. Cell 170(457–469):e413. https://doi.org/10.1016/j.cell.2017.07.002
Acknowledgements
This work was supported, in part, by Grants from the National Institute on Drug Abuse of the National Institutes of Health, DA042888, DA046714 and DA007242, the Mayday Foundation and the Peter F. McManus Charitable Trust to YXP, and a core Grant from the National Cancer Institute (CA008748) to Memorial Sloan Kettering Cancer Center.
Author information
Authors and Affiliations
Contributions
AN, GWP and YXP conceived and designed the study. AN, AH, JX, DIB and YXP performed experiments. AN, AH and YXP analyzed data. AN, GWP and YXP wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
YXP is a co-scientific founder of Sparian Biosciences. All other authors declare that there is no conflict of interest.
Additional information
This article is dedicated to the memory of our dear mentor, colleague, and friend, Dr. Gavril W. Pasternak (deceased February 22, 2019), whose pioneering work over four decades contributed enormously to our understanding of opioid pharmacology.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: typo in co-author name Amanda Hunkele has been corrected.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Narayan, A., Hunkele, A., Xu, J. et al. Mu Opioids Induce Biased Signaling at the Full-Length Seven Transmembrane C-Terminal Splice Variants of the mu Opioid Receptor Gene, Oprm1. Cell Mol Neurobiol 41, 1059–1074 (2021). https://doi.org/10.1007/s10571-020-00973-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-020-00973-5