Abstract
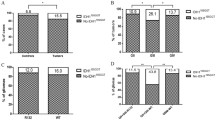
Molecular studies have an important role in the elucidation of the mechanisms involved in Glioblastoma multiforme (GBM) development. The occurrence of FHIT gene alterations, which has an important role in different cancers, has not yet been studied well in GBM. We aimed to investigate the occurrence of alterations of FHIT gene sequence and protein expression in the GBMs. Sequence alterations in exons 5–9 of the FHIT gene were screened in 63 GBMs using the single-strand conformational polymorphism method, followed by DNA sequencing. Additionally, the level of Fhit protein expression in tissues of 48 tumors was assessed by immunohistochemistry (IHC). In our investigation, FHIT gene alterations in the coding region were detected in 11 of the 63 GBM cases (17.5%). Two different sequence variants were determined: one novel missense variant (G→C transition at codon 49) and one previously described silent alteration (C→T transition at codon 88). Using web-based programs, such as SIFT and ESEfinder, it was determined that both alterations might have caused significant modification on protein function. In addition, we identified a previously reported an intronic polymorphism (T→A transition at IVS8-17) in 47.5% of cases as a similar rate (45%) in the control group. Moreover, it was observed that Fhit protein expression was reduced in 87.5% of tumors. In conclusion, the reduction or loss of Fhit protein expression by genetic alterations or epigenetic mechanisms in GBM might be associated with brain tumorigenesis.



Similar content being viewed by others
Abbreviations
- GBM:
-
Glioblastoma multiforme
- IHC:
-
Immunohistochemistry
- PCR:
-
Polymerase chain reaction
- SSCP:
-
Single strand conformational polymorphism
- WHO:
-
World Health Organization
References
Ahmadian M, Wistuba II, Fong KM et al (1997) Analysis of the FHIT Gene and FRA3B region in sporadic breast cancer, preneoplastic lesions, and familial breast cancer probands. Cancer Res 57:3664–3668
Barker FG, Prados MD, Chang SM et al (1996) Radiation response and survival time in patients with glioblastoma multiforme. J Neurosurg 84:442–448
Batchelor TT, Betensky RA, Esposito JM, Pham LD, Dorfman MV, Piscatelli N, Jhung S, Rhee D, Louis DN (2004) Age-dependent prognostic effects of genetic alterations in glioblastoma. Clin Cancer Res 10:228–233
Bekar A, Cecener G, Tunca B, Guler G, Egeli U, Tolunay S (2007) Investigation of mutations and expression of the FHIT gene in Turkish patients with brain metastases derived from non-small cell lung cancer. Tumori 93:604–607
Cecener G, Egeli Ü, Taşdelen İ, Tunca B, Duman H, Kızıl A (1998) Common fragile sites expression and genetic predisposition to breast cancer. Teratog Carcinog Mutagen 18:279–291
Cecener G, Egeli U, Tunca B, Tasdelen I, Tolunay S, Bilgel N (2007) Importance of novel sequence alterations in the FHIT gene on formation of breast cancer. Tumori 93:597–603
Collins VP (2004) Brain tumours: classification and genes. J Neurol Neurosurg Psychiatry 75:2–11
Curran WJ Jr, Scott CB, Horton J et al (1993) Recursive partitioning analysis of prognostic factors in three radiation therapy oncology group malignant glioma trials. J Natl Cancer Inst 85:704–710
Druck T, Hadaczek P, Fu TB et al (1997) Structure and expression of the human FHIT gene in normal and tumor cells. Cancer Res 57:504–512
Egeli Ü, Karadağ M, Tunca B, Özyardımcı N (1997) The expression of common fragile sites and genetic predisposition to squamous cell lung cancers. Cancer Genet Cytogenet 95:153–158
Egeli U, Özkan L, Tunca B, Kahraman S, Cecener G, Ergül E, Engin K (2000) The relationship between genetic susceptibility to head and neck cancer with the expression of common fragile sites. Head Neck 22:591–598
Fong KM, Biesterveld EJ, Virmania A et al (1997) FHIT and FRA3B 3p14.2 Allele loss are common in lung cancer and preneoplastic bronchial lesions and are associated with cancer related FHIT cDNA splicing aberrations. Cancer Res 57:2256–2267
Frank S, Muller J, Plaschke J, Hahn M, Hampl J, Hampl M, Pistorius S, Schackert G, Schackert HK (1997) The putative tumor suppressor gene FHIT at 3p14.2 is rarely affected by loss of heterozygosity in primary human brain tumors. Cancer Res 57:2638–2641
Gemma A, Hagiwarn K, Ke Y, Burke LM, Khan MA, Nagashima M, Bennett WP, Harris CC (1997) FHIT mutations in human primary gastric cancer. Cancer Res 57:1435–1437
Guler G, Uner A, Guler N, Han SY, Iliopoulos D, Hauck WW, McCue P, Huebner K (2004) The fragile genes FHIT and WWOX are inactivated coordinately in invasive breast carcinoma. Cancer 100:1605–1614
Hill C, Hunter SB, Brat DJ (2003) Genetic markers in glioblastoma: prognostic significance and future therapeutic implications. Adv Anat Pathol 10:212–217
Huebner K, Croce C (2001) FRA3B and other common fragile sites: the weakest links. Nat Rev Cancer 1:214–221
Ichimura K, Ohgaki H, Kleihues P, Collins VP (2004) Molecular pathogenesis of astrocytic tumours. J Neurooncol 70:137–160
Iliopoulos D, Guler G, Han SY, Druck T, Ottey M, McCorkell KA, Huebner K (2006) Roles of FHIT and WWOX fragile genes in cancer. Cancer Lett 232:27–36
Jansen M, de Witt Hamer PC, Witmer AN, Troost D, van Noorden CJ (2004) Current perspectives on antiangiogenesis strategies in the treatment of malignant gliomas. Brain Res 45:143–163
Kannan K, Krishnamurthy J, Feng J, Nakajima T, Tsuchida N, Shanmugam G (2000) Mutation profile of the p53, FHIT, p16INK4a/p19ARF and H-ras genes in Indian breast carcinomas. Int J Oncol 17:1031–1035
Karadağ M, Tunca B, Cecener G, Egeli Ü, Özyardımcı N, Ege E, Gözü O (2002) Chromosomal fragile sites and relationship between genetic predisposition to small cell lung cancer. Teratog Carcinog Mutagen 22:31–40
Kleihues P, Burger PC, Scheithauer BW (1993) The new WHO classification of brain tumours. Brain Pathol 3:255–268
Le Beau MM, Drabkin H, Glover TW, Gemmill R, Rassool FV, McKeithan TW, Smith DI (1998) An FHIT tumor suppressor gene? Genes Chromosomes Cancer 21:281–289
Mathe E, Olivier M, Kato S, Ishioka C, Hainaut P, Tavtigian SV (2006) Computational approaches for predicting the biological effect of p53 missense mutations: a comparison of three sequence analysis based methods. Nucleic Acids Res 34:1317–1325
Ohgaki H, Kleihues P (2005) Population-based studies on incidence, survival rates, and genetic alterations in astrocytic and oligodendroglial gliomas. J Neuropathol Exp Neurol 64:479–489
Ohgaki H, Dessen P, Jourde B et al (2004) Genetic pathways to glioblastoma: a population-based study. Cancer Res 64:6892–6899
Ohta M, Inoue H, Cotticelli MG et al (1996) The FHIT gene, spanning the chromosome 3p14.2 fragile site and renal carcinoma associated t(3;8) breakpoint, is abnormal in digestive tract cancers. Cell 84:587–597
Pekarsky Y, Zanesi N, Palamarchuk A, Huebner K, Croce CM (2002) FHIT: from gene discovery to cancer treatment and prevention. Lancet Oncol 3:748–754
Sozzi G, Veronese M, Negrini M et al (1996) The FHIT Gene At 3p14.2 is abnormal in lung cancer. Cell 85:17–26
Tsai TC, Yang HM, Wu YL, Chi CW, Chou MD, Lee LS, Chang TJ (1999) Abnormal transcripts of FHIT gene in Chinese brain tumors. Oncol Rep 6:345–348
Tunca B, Egeli Ü, Zorluoğlu A, Yılmazlar T, Yerci Ö, Kızıl A (2000) The expression frequency of common fragile sites and genetic predisposition to colon cancer. Cancer Genet Cytogenet 119:139–145
Tunca B, Cecener G, Gebitekin C, Egeli U, Ediz B, Ercan I (2002) Investigation of genetic susceptibility to non-small cell lung cancer by fragile site expression. Teratog Carcinog Mutagen 22:205–215
Tunca B, Bekar A, Cecener G, Egeli U, Vatan O, Tolunay S, Kocaeli H, Aksoy K (2007) Impact of novel PTEN mutations in Turkish patients with glioblastoma multiforme. J Neurooncol 82:263–269
Wang J, Smith PJ, Krainer AR, Zang MQ (2005) Distribution of SR protein exonic splicing enhancer motifs human protein-coding genes. Nucleic Acids Res 33:5053–5062
Watson JD, Hopkins NH (1987) The genetic code. In: Watson JD, Hopkins NH (eds) Molecular biology of the gene, 4th edn. The Benjamin/Cummings Publishing Company, California, p 444
Yoshino K, Enomoto T, Nakamura T, Nakashima R, Wada H, Saitoh J, Noda K, Murata Y (1998) Aberrant FHIT transcripts in squamous cell carcinoma of the uterine cervix. Int J Cancer 76:176–181
Zhao XR, Kang LC, Zhou YS et al (2003) Mutations of fragile histidine triad gene in Peutz-Jeghers syndrome and canceration. Ai Zheng 22:50–54
Acknowledgments
We would like to thank Professor Dr. Kay Huebner in the Comprehensive Cancer Center, Ohio State University for her suggestions and kindly providing Fhit antiserum. We thank Dr. Ediz in the Biostatistics Department, Medical Faculty of Uludag University for his expert advice. We also thank Prizma and Elips Ltd for their support in dealing with experimental equipment.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cecener, G., Tunca, B., Egeli, U. et al. FHIT Gene Sequence Variants and Reduced Fhit Protein Expression in Glioblastoma Multiforme. Cell Mol Neurobiol 30, 301–307 (2010). https://doi.org/10.1007/s10571-009-9452-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-009-9452-9