Abstract
There is increasing evidence that microbes play a key role in some plant invasions. A diverse and widespread but little understood group of plant-associated microbes are the fungal root endophytes of the order Sebacinales. They are associated with exotic populations of invasive knotweed (Reynoutria ssp.) in Europe, but their effects on the invaders are unknown. We used the recently isolated Sebacinales root endophyte Serendipita herbamans to experimentally inoculate invasive knotweed and study root colonisation and effects on knotweed growth under different environmental conditions. We verified the inoculation success and fungal colonisation through immunofluorescence microscopy and qPCR. We found that S. herbamans strongly colonized invasive knotweed in low-nutrient and shade environments, but much less under drought or benign conditions. At low nutrients, the endophyte had a positive effect on plant growth, whereas the opposite was true under shaded conditions. Our study demonstrates that the root endophyte S. herbamans has the potential to colonize invasive knotweed fine roots and impact its growth, and it could thus also play a role in natural populations. Our results also show that effects of fungal endophytes on plants can be strongly environment-dependent, and may only be visible under stressful environmental conditions.
Similar content being viewed by others
Introduction
Fungal endophytes are a phylogenetically diverse and widespread group of plant-associated microbes (Rodriguez et al. 2009). They can influence the growth and reproduction of individual plants, or their resistance to abiotic stress or natural enemies (Cosme et al. 2016; Kivlin et al. 2013; Mayerhofer et al. 2013; Oberhofer et al. 2014; Rho et al. 2018; Rodriguez et al. 2008). Some of the positive effects are related to the ability of endophytes to improve the nutrition of their host plants (Behie and Bidochka 2014). There is also evidence that endophytes can influence the diversity and composition of entire plant communities (Afkhami and Strauss 2016; Aguilar-Trigueros and Rillig 2016; Clay and Holah 1999; Rudgers et al. 2004, 2005) as well as their associated ecological networks (e.g. herbivores and their parasitoids; Omacini et al. 2001). However, so far our understanding of fungal endophytes is based on experiments with very few taxa, in particular the genus Neotyphodium and its asexual stage Epichloë. Other fungal systems have been hardly studied, mainly because most fungal endophytes are often difficult to cultivate and thus controlled experiments for testing their ecological functions have so far been impossible.
An important group of fungal endophytes for which this has long been true is the Serendipitaceae family in the order of Sebacinales that contains many species with broad geographic and host ranges (Garnica et al. 2016; Weiss et al. 2011). Previous experimental work has so far been largely restricted to Serendipita indica (Piriformospora indica), and it showed that S. indica stimulates plant growth and influences plant nutrition and tolerances to biotic and abiotic stresses (Achatz et al. 2010; Barazani et al. 2005; Gill et al. 2016; Waller et al. 2005). Our group in Tübingen recently isolated and cultivated another widespread Serendipitaceae species, Serendipita herbamans, which is abundant and associated with a broad range of host species and habitats in Central Europe (Riess et al. 2014).
Soil microbes can influence plant growth and stress tolerance, and these effects are to some extent host plant-specific. As a consequence, plant–microbe interactions play a role in structuring plant communities, and there is increasing evidence that they are also important in the invasion of exotic plant species (Callaway et al. 2004; Dawson and Schrama 2016; Inderjit and van der Putten 2010; Klironomos 2002). In general, plant-associated microbes may have positive or negative feedbacks on plants (Bever et al. 2012; van der Putten et al. 2013). If exotic plants accumulate biota with overall more positive effects, possibly because some of their native pathogens are not present in the introduced range, this may give invaders an advantage over native plants (Callaway et al. 2011; Maron et al. 2014; Mitchell and Power 2003; Reinhart et al. 2003). Alternatively, exotic plants may influence soil biota to the detriment of native plants, e.g. through increasing abundances of their pathogens (Mangla and Callaway 2008) or disrupting interactions with mutualists (Meinhardt and Gehring 2012; Stinson et al. 2006).
Most previous research on plant–microbe interactions and plant invasion has focused on soil-borne microbes rather than endophytes, even though fungal endophytes are clearly abundant and diverse also in invasive plant populations (Clay et al. 2016; Shipunov et al. 2008). Besides an interesting series of studies by Aschehoug et al. (2012, 2014) who demonstrated that the leaf endophyte Alternaria alternata makes invasive knapweed (Centaurea stoebe) more competitive and allelopathic against native North American grasses, there has so far been little experimental work on fungal endophytes and invasive plants.
One of the most problematic plant invaders of temperate Europe and North America is the Japanese knotweed (Reynoutria japonica) and its hybrid R. × bohemica. Their aggressive growth can damage buildings and other structures, and they have huge impacts on native plant communities and ecosystems (Aguilera et al. 2010; Gerber et al. 2008; Hejda et al. 2009). Because of these ecological and economic costs, there is great interest in controlling invasive knotweeds, and in understanding the biological mechanisms contributing to their success. Previous experimental research indicates that allelochemical or microbial processes belowground, or their interplay, may contribute to knotweed invasion success (Murrell et al. 2011; Parepa et al. 2013b; Siemens and Blossey 2007). However, the precise mechanisms and in particular microbial taxa involved in these phenomena are unknown.
In a preliminary screening of some invasive knotweed populations around Tübingen (see Supplementary Information) we had found a large diversity of root-associated fungi, and 40% of the studied fine-root samples also harboured ITS sequences of Sebacinales fungi, including S. herbamans (Table S1). Thus, interactions between invasive knotweed and Sebacinales appeared to be common, and we were curious about the nature of the interaction between the two taxa, in particular whether these hidden and little understood, but very common, fungi influenced the growth and performance of invasive knotweed. We suspected that if S. herbamans had an effect on knotweed, it would depend on environmental conditions, in particular the resource supply of the plants. We tested this through a greenhouse experiment in which we inoculated Japanese knotweed and its hybrid with S. herbamans under benign as well as drought, shading and low-nutrient conditions. Compared to previous studies on plant-Sebacinales interactions, our experiment had a rather realistic set-up, with a natural, non-sterile substrate and a range of environmental conditions. To confirm inoculation success and quantify fungal colonization, we used immunofluorescence microscopy and qPCR. Specifically, we asked the following questions: (1) Does colonization of knotweed by S. herbamans depend on environmental conditions? (2) What effects does the endophyte have on the growth and performance of knotweed, and (3) Are these effects environment-dependent?
Methods
Plant material
Reynoutria japonica and its hybrid Reynoutria × bohemica are large perennial forbs from the Polygonaceae family that have become invasive in riparian and ruderal habitats in the temperate regions of Europe and North America. They are clonal plants with extensive rhizome networks, and in their invasive range they often form dense monoclonal stands and become extremely dominant (Aguilera et al. 2010). In our experiment, we used plant material from a live collection of knotweed clones that had originally been collected across seven regions in Switzerland and Germany (Krebs et al. 2010) and that had been cultivated in a common garden for several years. We used rhizome cuttings from 20 R. japonica clones and 13 R. × bohemica clones, with approximately ten rhizome fragments, each containing two nodes and thus one intact internode, from each clone. After removal of all fine roots, the rhizome fragments were surface-sterilized using the method described by Huang et al. (2014).
Endophyte material
We worked with the endophyte Serendipita herbamans (DSM 27,534), a member of the order Sebacinales, whose discovery and isolation was described in Riess et al. (2014). Prior to the experiment, we grew the endophyte for 14 days in Petri dishes with MEA medium containing 2% malt extract and 1.5% agar at 20 °C in the dark. We then used 5 mm plugs from these plates, containing media and mycelia, to inoculate 0.5 L Erlenmeyer flasks with 250 ml malt extract (2.0%) liquid medium. The inoculated flasks were incubated in the dark on a rotary shaker (47–52 rpm) at 20 °C. After two weeks of incubation, the resulting mycelium was separated from the media and washed five times with sterile distilled water.
Experimental set-up
We set up a greenhouse experiment at the University of Tübingen in which we tested the effects of endophyte inoculation on knotweed growth in four different environments: control, drought, low nutrients and shade. Except for the control environment, all conditions were expected to be stressful for the plants. We planted individual rhizome fragments 3 cm deep in 1.5 L pots filled with a 1:3 mixture of field soil and sand (Sand- und Kieswerk Bischoff, Rottenburg, Germany). Prior to planting, we measured the length and diameter of each rhizome fragment. All pots were placed on individual saucers and watered as needed. After two weeks, when all aboveground shoots had appeared, we inoculated half of the pots with 0.5 g of fresh S. herbamans mycelium which were applied in small pits close to the center of the rhizomes (Fig. 1a). For non-inoculated plants we also created the same pits and applied a similar volume of distilled water. Another two weeks after the inoculation, we started the environmental treatments. In the shade treatment, the plants were covered individually with shading mesh bags that reduced light levels to approximately 20%. The low-nutrient plants did not receive any fertilizer throughout the experiment, whereas all others received 7:3:6 N:P:K fertilizer (b1 Universal-Flüssigdünger, toom Baumarkt GmbH) equivalent to 150 kg N/ha distributed across 15 applications at seven-day intervals during the treatments. The drought plants generally received only a third of the regular watering amount and, in contrast to all other plants, regularly showed signs of wilting (loss of turgor). A total of 288 plants (160 R. japonica and 128 R. x bohemica) were randomly assigned to the eight treatment combinations (four environmental conditions, with or without endophytes), with approximately equal representation of the two taxa in each treatment. Throughout the experiment, the plants were grown in a climate-controlled greenhouse, in a completely randomized order, with supplemental lighting at a 14:10 h light:dark cycle at 20/18 °C.
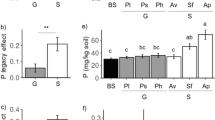
Experimental inoculation of Reynoutria plants with Serendipita herbamans, and the resulting fungal colonization. a An experimental pot right after inoculation, with freshly regenerated knotweed and the pit through which S. herbamans mycelium was added. b Fluorescence microscopic image of a root section of an inoculated Reynoutria plant, stained with WGA-AF 488. c Relative colonization rates of S. herbamans in the different experimental treatments, based on qPCR analyses. Points = individual observations; boxes = 25th—75th percentiles; thick horizontal lines = medians; whiskers = 10th—90th percentiles
Data collection
15 weeks after the start of the treatments, we measured leaf chlorophyll content on four leaves per plant using a handheld chlorophyll meter (SPAD 502Plus, Konica Minolta, Osaka, Japan). We then removed all leaves from the stem, measured their area with a LI-3100C leaf area meter (LI-COR Environmental, Lincoln, Nebraska, USA) and dried the leaves and stems of each plant separately at 80 °C for 3 days. We used the leaf area and leaf dry mass to calculate the specific leaf area (SLA) of each plant, and we combined the leaf and stem dry mass to total aboveground biomass. Finally, we carefully washed the roots of a subset of the plants (see next section) and took 10 fine root samples from different parts of the root system that we mixed and immediately stored at − 20 °C for subsequent DNA extraction, or placed in fixing solution (0.15% (wt/vol) trichloroacetic acid in 4:1 (vol/vol) ethanol/chloroform) for microscopy.
Assessment of endophyte colonization
We assessed the fungal colonization of knotweed roots qualitatively through fluorescence microscopy and quantitatively through qPCR. Both analyses were done for randomly selected subsets of the plants. For the microscopy, we collected roots from three pots of each treatment by species combination, including both inoculated and non-inoculated samples (altogether 16 × 3 = 48 samples). The root samples were stained with Wheat Germ Agglutinin-Alexa Fluor 488 (WGA488; Thermo Fisher, Waltham MA, USA), which specifically stains fungal cell walls. The staining procedure was as described in Deshmukh et al. (2006), and the images were recorded on a Leica TCS SP5 2 confocal microscope using the bright field channel and a GFP filter set for detection of WGA488.
For the qPCR, we analysed 10 plants from each treatment by species combination, i.e. a total of 80 plants, with three replicates of non-inoculated plants and seven replicates for inoculated plants. We ground fine roots to a fine powder in liquid nitrogen using a sterile mortar and pestle, and we used 500 mg of this material to isolate DNA using the DNAeasy Plant Mini Kit (Qiagen, Hilden, Germany). The relative amounts of S. herbamans DNA in the samples were determined through qPCR reactions with a S. herbamans-specific and a Reynoutria-specific primer pair. The S. herbamans-specific primer pair SerhaITS binds to the ITS region of the 5.8 S rDNA sequence of S. herbamans (SerhaITSfw199: 5’-AGCCTTGTGCGGTAAAGCGA-3’, SerhaITSrev199: 5’-TGTATTCCGGCACCTTAACCTC-3’). The Reynoutria-specific primer pair FallCHS binds to the genomic DNA of the Chalcone synthase gene EF090266.2 of Fallopia japonica (now Reynoutria japonica) (FallCHSfwd: 5’-GGAGATGCGTGTATATTCTT-3’, FallCHSrev: 5’-CCAAAGATGAAGCCATGTAG-3’. The PCR primers were designed using Primer-BLAST (Ye et al. 2012). For the PCR amplification we ran real-time PCRs on a Biorad CFX96 Thermocycler (BioRad, Hercules CA, USA) using the ABsolute SYBR Capillary Mix (Thermo Fisher, Waltham MA, USA) in a final volume of 20 μl, and the following cycler programmes: 95 °C for 15 min followed by 45 cycles of 95 °C for 15 s, 55 °C for 20 s and 72° C for 20 s for the FallCHS primer pair, and 95 °C for 15 min followed by 45 cycles of 95 °C for 15 s, 60 °C for 15 s and 72 °C for 10 s for the Serha199 primer pair. To calculate relative amounts of S. herbamans DNA, we used the 2−deltaCt method (Livak and Schmittgen 2001) using the raw threshold cycle (Ct) values determined for the S. herbamans- and the Reynoutria-specific primer pairs.
Statistical analyses
To test for species differences in, and effects of environmental conditions on endophyte colonization, we analysed the relative S. herbamans densities, as determined by qPCR, with a linear model that included the effects of Reynoutria species, environmental treatment, and their interaction as fixed factors. We analysed knotweed responses to endophytes and environments with regard to three variables: aboveground biomass, leaf chlorophyll content, and specific leaf area. For each response variable we fitted a linear mixed model with fungal inoculation, environmental treatments, knotweed species, and interactions included as fixed factors, and clone identity included as random factor. To account for possible influences of initial size differences, we included the volume of the planted rhizome as a covariate in all three analyses. Prior to the analyses, the biomass and specific leaf area data were log-transformed to achieve homoscedasticity. All linear models were fitted with the lmer function in the lme4 package (Bates et al. 2015) in R (R Core Team 2018). In addition to testing for overall effects of endophyte inoculation in the linear models, we also conducted pairwise comparisons of inoculated vs. inoculated plants in each environmental treatment, using Tukey tests in the lsmeans package (Lenth 2016). We used the effects (Fox 2003) and ggplot2 (Wickham 2009) packages to visualize results.
Results
The experimental inoculation of knotweed plants with Serendipita herbamans was successful, but colonization rates were strongly environment-dependent (Fig. 1c; main effect of environmental treatment in the linear model: F = 27.12, P < 0.001). The average colonization level was lowest – albeit not zero – in the non-inoculated samples, and it remained low in the inoculated control and drought treatments. Under shaded and low nutrient conditions, however, colonization rates increased four-fold and eight-fold, respectively (Fig. 1c). There were no differences among the two Reynoutria species in terms of fungal colonisation (F < 1 and P > 0.5 for species main effect and species x treatment interaction; Fig. S1A). The colonization of Reynoutria roots by S. herbamans was confirmed by fluorescence microscopy. In all root samples from inoculated plants we detected hyphal structures typical for S. herbamans on root surfaces, between the outer cell layers, and inside of some cortical root cells (Fig. 1b). We also observed some hyphal structures in the roots of non-inoculated plants.
As expected, the stress treatments in our experiment strongly impacted the growth of knotweed (Table 1, Fig. 2). Compared to control plants, the biomass was reduced in all three stress treatments, but particularly strongly under low-nutrient conditions. There were also strong treatment effects on chlorophyll content and SLA, with a particularly low chlorophyll content at low nutrient availability, and the highest SLA under shaded conditions (Fig. 2). There were also differences between the two knotweed taxa (Table 1). The hybrid Reynoutria x bohemica was generally larger (+ 25%) and had a higher SLA (+ 5%) than R. japonica, and its biomass was less sensitive to drought and shading than that of R. japonica (Fig. S1B).
Effects of inoculation with Serendipita herbamans on invasive knotweed aboveground biomass, leaf chlorophyll content, and specific leaf area under different types of stress and in control conditions. The disc areas are proportional to the estimated means without endophyte inoculation, and the arrows show the log-responses of plants to endophyte inoculation in each treatment. Black arrows represent significant endophyte effects within a particular treatment, as indicated by Tukey’s HSD test, grey arrows are non-significant
Inoculation with Serendipita herbamans influenced the growth of the knotweed plants, but again in a strongly treatment-dependent manner, with significant fungus by treatment interactions (but no fungus main effects) for aboveground biomass and chlorophyll content, and a marginally significant interaction for SLA (Table 1, Fig. 2). Under low-nutrient conditions, addition of the endophyte increased knotweed biomass by 15% (Tukey’s HSD, P < 0.001), but decreased it by 10% (P = 0.021) in the shade, or had no effect at all under drought or control conditions. Similarly, endophyte inoculation increased chlorophyll content by 13% (P = 0.006) under low nutrients but decreased it by 5% (P = 0.020) under shaded conditions, and had no significant effects in the other two treatments. Finally, the SLA was positively affected (+ 5%, P = 0.044) in the shade but negatively (− 6%, P = 0.042) under low-nutrient conditions, with no effects under drought or control conditions.
Discussion
Plant–microbe interactions play an important role in natural ecosystems (Bever et al. 2012; Klironomos 2002; van der Putten et al. 2013). However, the ecological function of endophytic microbes that live within plants is so far little understood. In this study we show that the fungal root endophyte Serendipita herbamans can rapidly colonize invasive knotweed (Reynoutria ssp.) and influence its growth, with detrimental effects in the shade but promotion of growth under low-nutrient conditions. Our study thus demonstrates that this widespread endophyte interacts with an important invasive plant, and it also highlights the environment-dependency of plant-endophyte interactions.
Endophyte colonization
Compared to previous studies on plant-Sebacinales interactions, our experiment had a rather realistic set-up, with fungi inoculated into a non-sterile natural soil that presumably already contained a resident microbial community. Microscopy and qPCR confirmed that our inoculations were successful and that S. herbamans was able to colonize knotweed plants, which confirms field observations in the Tübingen area where Sebacinales including S. herbamans are frequent endophytes of invasive knotweed populations (Table S1). This is not surprising, given the broad host range of Serendipita herbamans, and of Sebacinales in general, which also includes native Polygonaceae (Garnica et al. 2013; Riess et al. 2014). Although exotic species are known to lose specialised biotic interactions, they often interact with generalist enemies and mutualists in the introduced range (Mitchell and Power 2003; Richardson et al. 2007; van Kleunen et al. 2018). However, so far we do not know how novel the interaction between knotweed and S. herbamans really is because there are no data on endophyte diversity from the native East Asian range.
We found that the relative colonization of knotweed plants by S. herbamans was generally much stronger under low-nutrients or shade conditions than under control or drought conditions. Thus, Reynoutria plants appear to actively regulate their interactions with S. herbamans in an environment-specific fashion. It is known that plants can control fungal colonization, e.g. through the production of defense compounds or secondary metabolites inhibiting microbial growth (Zipfel and Oldroyd 2017), or by diverting more carbohydrates to fungal symbionts (Carbonnel and Gutjahr 2014; Martin et al. 2017). This has also been shown for the closely related S. indica which interferes with the immune system of host plants (Jacobs et al. 2011) and influences sugar concentrations in their roots (Opitz et al. 2021). The functional and adaptive explanation for this is usually that plant benefits from interactions with fungi are environment-dependent, and therefore plants stimulate or restrict fungal access depending on these benefits. For instance, mycorrhizal colonisation is often triggered by low-nutrient conditions (Bueno de Mesquita et al. 2018). We also found that relative fungal colonization was highest under low-nutrient conditions, which is in line with the idea that S. herbamans improves the nutrition of Reynoutria plants. It is less clear why relative colonization was also increased in shaded plants because these should have been mainly carbon-limited, and under such conditions plant–microbe interactions often turn parasitic, as has been shown e.g. for interactions with mycorrhiza or rhizobia (Ballhorn et al. 2016; Lau et al. 2012).
We also detected S. herbamans in some non-inoculated plants. The sources of this could be external, e.g. fungi spores present in the potting soil, or splash dispersal from adjacent pots. However, the most likely explanation seems that S. herbamans was already present in some of the planted rhizomes. We know that some invasive knotweed populations are naturally colonized by S. herbamans, and we therefore cannot rule out that some surface-sterilized rhizomes still harboured the fungus.
Endophyte effects on plant growth
The inoculated S. herbamans fungi not only successfully colonized the knotweed plants in our experiment, but they also significantly impacted their growth. The magnitude and direction of these effects were strongly environment-dependent. Under benign or drought conditions, endophyte effects on plants were small and non-significant, whereas under low-nutrient conditions inoculation had strong positive effects, and under shade conditions strong negative effects on knotweed performance. Similar context-dependent effects of endophytes have been found in other study systems (Davitt et al. 2010; Laitinen et al. 2016; Shaffer et al. 2018).
Low-nutrient conditions greatly reduced knotweed biomass, and here relative S. herbamans colonization was strong and the fungus increased plant growth. This suggests an active promotion of endophyte access by the plants because the fungi improve plant nutrition under these conditions. The observed increase of leaf chlorophyll content, which strongly correlates with leaf nitrogen content (Evans 1989), supports this idea. We know that S. herbamans improves plant growth under lab conditions (Riess et al. 2014), and that the closely related Serendipita indica can improve the nutrient acquisition and growth of many plant species (Achatz et al. 2010; Barazani et al. 2005; Giauque et al. 2019; Varma et al. 1999; Waller et al. 2005). Thus, it seems very likely that S. herbamans also improved the nutrition, and as a consequence biomass growth, of invasive knotweed in our experiment.
Under shade conditions, the effects of S. herbamans were reversed, and inoculation negatively affected knotweed biomass as well as leaf chlorophyll content, suggesting that under these conditions the fungus indeed turned parasitic and compromised plant nutrition. Similar shifts in the directions of plant–microbe interactions have been observed in other studies (Ballhorn et al. 2016; Lau et al. 2012), and the likely explanation is that the typical ‘trade logic’ of mutualistic plant–microbe interactions – microbes receive photosynthates in exchange for improved nutrient uptake – only works where soil nutrients are limiting, but under carbon-limited shade conditions, it does not.
In the control and drought treatments, colonization and growth effects of endophytes were very low, indicating that under these conditions the host plants limited fungi access, similar to what is known from plant-mycorrhiza interactions (Averill et al. 2019; Carbonnel and Gutjahr 2014). For the drought treatment, with episodes of plant wilting, it is also possible that the spread of fungi was simply limited by the lack of moisture.
Our results that S. herbamans can promote or weaken knotweed growth depending on environmental context raises intriguing questions about the habitat preferences of invasive knotweeds. Across their invasive range in Europe and North America, the species mostly thrive in open and nutrient-rich habitats, and benefit in particular from fluctuating nutrient supply (Parepa et al. 2013a), but they rarely spread under closed canopy (Beerling 1991; Pyšek et al. 2009). It is possible that interactions with S. herbamans or other microbes contribute to these habitat preferences, by facilitating nutrient uptake in open habitats but limiting knotweed under shaded conditions. Further research—in particular field experiments—is needed to test these hypotheses.
Conclusions
Our study demonstrates that the common fungal endophyte Serendipita herbamans can rapidly colonize fine roots of invasive knotweed and influence its growth both positively or negatively, depending on the environmental context. As S. herbamans is present in at least some invasive knotweed populations, the fungus could play a role in the growth and success of knotweed in some invaded habitats. However, understanding the true significance of this plant-fungus interaction requires further data, because ecological communities are of course more complex than our experimental set-up. In its natural habitat, invasive knotweed also interacts with competitors, herbivores and other enemies and mutualists, and some of these might be interacting with S. herbamans, too. Thus, the next step should be multi-species experiments, in the field or using mesocosm approaches, that evaluate the impact of S. herbamans on knotweed and other plants in a community context.
Data availability
All data from our experiment will be made available through Dryad. The sequencing data from the supplement are stored at GenBank under MZ650923—MZ651047.
References
Achatz B, von Rüden S, Andrade D, Neumann E, Pons-Kühnemann J, Kogel K-H, Franken P, Waller F (2010) Root colonization by Piriformospora indica enhances grain yield in barley under diverse nutrient regimes by accelerating plant development. Plant Soil 333(1):59–70
Afkhami ME, Strauss SY (2016) Native fungal endophytes suppress an exotic dominant and increase plant diversity over small and large spatial scales. Ecology 97(5):1159–1169
Aguilar-Trigueros CA, Rillig MC (2016) Effect of different root endophytic fungi on plant community structure in experimental microcosms. Ecol Evol 6(22):8149–8158
Aguilera AG, Alpert P, Dukes JS, Harrington R (2010) Impacts of the invasive plant Fallopia japonica (Houtt) on plant communities and ecosystem processes. Biol Invasions 12(5):1243–1252
Aschehoug ET, Metlen KL, Callaway RM, Newcombe G (2012) Fungal endophytes directly increase the competitive effects of an invasive forb. Ecology 93(1):3–8
Aschehoug ET, Callaway RM, Newcombe G, Tharayil N, Chen S (2014) Fungal endophyte increases the allelopathic effects of an invasive forb. Oecologia 175(1):285–291
Averill C, Bhatnagar JM, Dietze MC, Pearse WD, Kivlin SN (2019) Global imprint of mycorrhizal fungi on whole-plant nutrient economics. Proc Natl Acad Sci USA 116(46):23163–23168
Ballhorn DJ, Schädler M, Elias JD, Millar JA, Kautz S (2016) Friend or foe-light availability determines the relationship between mycorrhizal fungi, rhizobia and lima bean (Phaseolus lunatus L). PLoS ONE 11(5):e0154116
Barazani O, Benderoth M, Groten K, Kuhlemeier C, Baldwin IT (2005) Piriformospora indica and Sebacina vermifera increase growth performance at the expense of herbivore resistance in Nicotiana attenuata. Oecologia 146(2):234–243
Bates D, Mächler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67(1):1–48
Beerling DJ (1991) The effect of Riparian land use on the occurrence and abundance of Japanese knotweed Reynoutria japonica on selected rivers in South Wales. Biol Cons 55(3):329–337
Behie SW, Bidochka MJ (2014) Nutrient transfer in plant-fungal symbioses. Trends Plant Sci 19(11):734–740
Bever JD, Platt TG, Morton ER (2012) Microbial population and community dynamics on plant roots and their feedbacks on plant communities. Annu Rev Microbiol 66:265–283
Bueno de Mesquita CP, Sartwell SA, Ordemann EV, Porazinska DL, Farrer EC, King AJ, Spasojevic MJ, Smith JG, Suding KN, Schmidt SK (2018) Patterns of root colonization by arbuscular mycorrhizal fungi and dark septate endophytes across a mostly-unvegetated, high-elevation landscape. Fungal Ecol 36:63–74
Callaway RM, Thelen GC, Rodriguez A, Holben WE (2004) Soil biota and exotic plant invasion. Nature 427(6976):731–733
Callaway RM, Bedmar EJ, Reinhart KO, Silvan CG, Klironomos J (2011) Effects of soil biota from different ranges on Robinia invasion: acquiring mutualists and escaping pathogens. Ecology 92(5):1027–1035
Carbonnel S, Gutjahr C (2014) Control of arbuscular mycorrhiza development by nutrient signals. Front Plant Sci 5:462
Clay K, Holah J (1999) Fungal endophyte symbiosis and plant diversity in successional fields. Science 285(5434):1742–1745
Clay K, Shearin ZRC, Bourke KA, Bickford WA, Kowalski KP (2016) Diversity of fungal endophytes in non-native Phragmites australis in the Great Lakes. Biol Invasions 18(9):2703–2716
Cosme M, Lu J, Erb M, Stout MJ, Franken P, Wurst S (2016) A fungal endophyte helps plants to tolerate root herbivory through changes in gibberellin and jasmonate signaling. New Phytol 211(3):1065–1076
Davitt AJ, Stansberry M, Rudgers JA (2010) Do the costs and benefits of fungal endophyte symbiosis vary with light availability? New Phytol 188(3):824–834
Dawson W, Schrama M (2016) Identifying the role of soil microbes in plant invasions. J Ecol 104(5):1211–1218
Deshmukh S, Hückelhoven R, Schäfer P, Imani J, Sharma M, Weiss M, Waller F, Kogel K-H (2006) The root endophytic fungus Piriformospora indica requires host cell death for proliferation during mutualistic symbiosis with barley. Proc Natl Acad Sci USA 103(49):18450–18457
Evans JR (1989) Photosynthesis and nitrogen relationships in leaves of C3 plants. Oecologia 78(1):9–19
Fox J (2003) Effect displays in R for generalised linear models. J Stat Softw 8(15):1–27
Garnica S, Riess K, Bauer R, Oberwinkler F, Weiß M (2013) Phylogenetic diversity and structure of sebacinoid fungi associated with plant communities along an altitudinal gradient. FEMS Microbiol Ecol 83(2):265–278
Garnica S, Riess K, Schön ME, Oberwinkler F, Setaro SD (2016) Divergence times and phylogenetic patterns of Sebacinales, a highly diverse and widespread fungal lineage. PLoS ONE 11(3):e0149531
Gerber E, Krebs C, Murrell C, Moretti M, Rocklin R, Schaffner U (2008) Exotic invasive knotweeds (Fallopia spp) negatively affect native plant and invertebrate assemblages in European riparian habitats. Biol Conserv 141(3):646–654
Giauque H, Connor EW, Hawkes CV (2019) Endophyte traits relevant to stress tolerance, resource use and habitat of origin predict effects on host plants. New Phytol 221(4):2239–2249
Gill SS, Gill R, Trivedi DK, Anjum NA, Sharma KK, Ansari MW, Ansari AA, Johri AK, Prasad R, Pereira E, Varma A, Tuteja N (2016) Piriformospora indica: potential and significance in plant stress tolerance. Front Microbiol 7:332
Hejda M, Pyšek P, Jarošík V (2009) Impact of invasive plants on the species richness, diversity and composition of invaded communities. J Ecol 97(3):393–403
Huang B, Lin H, Yan C, Qiu H, Qiu L, Yu R (2014) Optimal inductive and cultural conditions of Polygonum multiflorum transgenic hairy roots mediated with Agrobacterium rhizogenes R1601 and an analysis of their anthraquinone constituents. Pharmacogn Mag 10(37):77–82
Inderjit and van der Putten WH (2010) Impacts of soil microbial communities on exotic plant invasions Trends Ecol Evol 25(9), 512–519
Jacobs S, Zechmann B, Molitor A, Trujillo M, Petutschnig E, Lipka V, Kogel K-H, Schäfer P (2011) Broad-spectrum suppression of innate immunity is required for colonization of Arabidopsis roots by the fungus Piriformospora indica. Plant Physiol 156(2):726–740
Kivlin SN, Emery SM, Rudgers JA (2013) Fungal symbionts alter plant responses to global change. Am J Bot 100(7):1445–1457
Klironomos JN (2002) Feedback with soil biota contributes to plant rarity and invasiveness in communities. Nature 417(6884):67–70
Krebs C, Mahy G, Matthies D, Schaffner U, Tiébré M-S, Bizoux J-P (2010) Taxa distribution and RAPD markers indicate different origin and regional differentiation of hybrids in the invasive Fallopia complex in central-western Europe. Plant Biol 12(1):215–223
Laitinen RK, Hellström KO, Wäli PR (2016) Context-dependent outcomes of subarctic grass-endophyte symbiosis. Fungal Ecol 23:66–74
Lau JA, Bowling EJ, Gentry LE, Glasser PA, Monarch EA, Olesen WM, Waxmonsky J, Young RT (2012) Direct and interactive effects of light and nutrients on the legume-rhizobia mutualism. Acta Oecologica 39:80–86
Lenth RV (2016) Least-squares means: the R package lsmeans. J Stat Softw 69:1–33
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25(4):402–408
Mangla S, Callaway RM (2008) Exotic invasive plant accumulates native soil pathogens which inhibit native plants. J Ecol 96(1):58–67
Maron JL, Klironomos J, Waller L, Callaway RM (2014) Invasive plants escape from suppressive soil biota at regional scales. J Ecol 102(1):19–27
Martin FM, Uroz S, Barker DG (2017) Ancestral alliances: plant mutualistic symbioses with fungi and bacteria. Science 356(6340):eaad4501
Mayerhofer MS, Kernaghan G, Harper KA (2013) The effects of fungal root endophytes on plant growth: a meta-analysis. Mycorrhiza 23(2):119–128
Meinhardt KA, Gehring CA (2012) Disrupting mycorrhizal mutualisms: a potential mechanism by which exotic tamarisk outcompetes native cottonwoods. Ecol Appl 22(2):532–549
Mitchell CE, Power AG (2003) Release of invasive plants from fungal and viral pathogens. Nature 421(6923):625–627
Murrell C, Gerber E, Krebs C, Parepa M, Schaffner U, Bossdorf O (2011) Invasive knotweed affects native plants through allelopathy. Am J Bot 98(1):38–43
Oberhofer M, Güsewell S, Leuchtmann A (2014) Effects of natural hybrid and non-hybrid Epichloë endophytes on the response of Hordelymus europaeus to drought stress. New Phytol 201(1):242–253
Omacini M, Chaneton EJ, Ghersa CM, Müller CB (2001) Symbiotic fungal endophytes control insect host-parasite interaction webs. Nature 409(6816):78–81
Opitz MW, Daneshkhah R, Lorenz C, Ludwig R, Steinkellner S, Wieczorek K (2021) Serendipita indica changes host sugar and defense status in Arabidopsis thaliana: cooperation or exploitation? Planta 253(3):74
Parepa M, Fischer M, Bossdorf O (2013a) Environmental variability promotes plant invasion. Nat Commun 4:1604
Parepa M, Schaffner U, Bossdorf O (2013b) Help from underground: soil biota facilitate knotweed invasion. Ecosphere 4(2):art31
Pyšek P, Jarošík V, Pergl J, Randall R, Chytrý M, Kühn I, Tichý L, Danihelka J, Chrtek Jun J, Sádlo J (2009) The global invasion success of Central European plants is related to distribution characteristics in their native range and species traits. Divers Distrib 15(5):891–903
R Core Team (2018) R: a language and environment for statistical computing R Foundation for Statistical Computing
Reinhart KO, Packer A, Van der Putten WH, Clay K (2003) Plant-soil biota interactions and spatial distribution of black cherry in its native and invasive ranges. Ecol Lett 6(12):1046–1050
Rho H, Hsieh M, Kandel SL, Cantillo J, Doty SL, Kim S-H (2018) Do endophytes promote growth of host plants under stress? A meta-analysis on plant stress mitigation by endophytes. Microb Ecol 75(2):407–418
Richardson DM, Allsopp N, D’antonio CM, Milton SJ, Rejmánek M (2007) Plant invasions - the role of mutualisms. Biol Rev 75(1):65–93
Riess K, Oberwinkler F, Bauer R, Garnica S (2014) Communities of endophytic sebacinales associated with roots of herbaceous plants in agricultural and grassland ecosystems are dominated by Serendipita herbamans sp nov. PLoS ONE 9(4):e94676
Rodriguez RJ, Henson J, Van Volkenburgh E, Hoy M, Wright L, Beckwith F, Kim Y-O, Redman RS (2008) Stress tolerance in plants via habitat-adapted symbiosis. ISME J 2(4):404–416
Rodriguez RJ, White JF Jr, Arnold AE, Redman RS (2009) Fungal endophytes: diversity and functional roles. New Phytol 182(2):314–330
Rudgers JA, Koslow JM, Clay K (2004) Endophytic fungi alter relationships between diversity and ecosystem properties. Ecol Lett 7(1):42–51
Rudgers JA, Mattingly WB, Koslow JM (2005) Mutualistic fungus promotes plant invasion into diverse communities. Oecologia 144(3):463–471
Shaffer JP, Zalamea P-C, Sarmiento C, Gallery RE, Dalling JW, Davis AS, Baltrus DA, Arnold AE (2018) Context-dependent and variable effects of endohyphal bacteria on interactions between fungi and seeds. Fungal Ecol 36:117–127
Shipunov A, Newcombe G, Raghavendra AKH, Anderson CL (2008) Hidden diversity of endophytic fungi in an invasive plant. Am J Bot 95(9):1096–1108
Siemens TJ, Blossey B (2007) An evaluation of mechanisms preventing growth and survival of two native species in invasive Bohemian knotweed (Fallopia x bohemica, Polygonaceae). Am J Bot 94(5):776–783
Stinson KA, Campbell SA, Powell JR, Wolfe BE, Callaway RM, Thelen GC, Hallett SG, Prati D, Klironomos JN (2006) Invasive plant suppresses the growth of native tree seedlings by disrupting belowground mutualisms. PLoS Biol 4(5):e140
van der Putten WH, Bardgett RD, Bever JD, Bezemer TM, Casper BB, Fukami T, Kardol P, Klironomos JN, Kulmatiski A, Schweitzer JA, Suding KN, Van de Voorde TFJ, Wardle DA (2013) Plant-soil feedbacks: the past, the present and future challenges. J Ecol 101(2):265–276
van Kleunen M, Bossdorf O, Dawson W (2018) The ecology and evolution of alien plants. Annu Rev Ecol Evol Syst 49(1):25–47
Varma A, Savita V, Sudha, Sahay N, Butehorn B, Franken P (1999) Piriformospora indica, a cultivable plant-growth-promoting root endophyte Appl Environ Microbiol 65(6), 2741–2744
Waller F, Achatz B, Baltruschat H, Fodor J, Becker K, Fischer M, Heier T, Hückelhoven R, Neumann C, von Wettstein D, Franken P, Kogel K-H (2005) The endophytic fungus Piriformospora indica reprograms barley to salt-stress tolerance, disease resistance, and higher yield. Proc Natl Acad Sci USA 102(38):13386–13391
Weiss M, Sýkorová Z, Garnica S, Riess K, Martos F, Krause C, Oberwinkler F, Bauer R, Redecker D (2011) Sebacinales everywhere: previously overlooked ubiquitous fungal endophytes. PLoS ONE 6(2):e16793
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer, New york
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13:134
Zipfel C, Oldroyd GED (2017) Plant signalling in symbiosis and immunity. Nature 543(7645):328–336
Acknowledgements
We thank Christiane Karasch-Wittmann, Sabine Silberhorn, Julia Rafalski, Mirjam Rieger, Mohamed Osman, and several student helpers for their assistance with the greenhouse experiment and/or molecular work. This project was supported by the German Research Foundation (DFG grant BO 3241/8-1 to OB), and by a scholarship of the China Scholarship Council (Grant No. 201504910498) to ZL.
Funding
Open Access funding enabled and organized by Projekt DEAL. Deutsche Forschungsgemeinschaft BO 3241/8–1,Oliver Bossdorf. China Scholarship Council, 201504910498, Zhiyong Liao.
Author information
Authors and Affiliations
Contributions
SG, MP and OB conceived and designed the experiment, SG, ZL and SH carried out the experiment, FW performed microscopy and qPCR analyses, MP and OB analyzed the data, SG, FW, MP and OB drafted the manuscript, all authors contributed to its revision and approved the version to be submitted for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Garnica, S., Liao, Z., Hamard, S. et al. Environmental stress determines the colonization and impact of an endophytic fungus on invasive knotweed. Biol Invasions 24, 1785–1795 (2022). https://doi.org/10.1007/s10530-022-02749-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-022-02749-y