Abstract
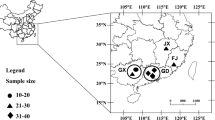
Gynaephora qinghaiensis (Lepidoptera: Lymantriidae: Gynaephora), a serious economic pest in alpine meadows, is mainly distributed in Yushu prefecture, Qinghai province, China. In this study, we aimed to investigate the genetic diversity and population structure of G. qinghaiensis through analyzing the sequence of 194 mitochondrial cytochrome oxidase subunit (COI) genes (658 bp in length) identified from 10 geographic populations located in three different countries, including Zhiduo, Zaduo, and Chengduo, of Yushu prefecture. Eleven haplotypes were identified from all populations of G. qinghaiensis with high levels of haplotype diversity (0.78500) and low levels of nucleotide diversity (0.00511). High levels of genetic differentiation and low levels of gene flow were also detected among the populations of G. qinghaiensis. Analysis of molecular variance (AMOVA) showed that 90.13% of the variation was attributed to distribution among groups (Chengduo, Zhiduo, and Zaduo), and 5.22% and 4.65% were, respectively, attributed to distribution among populations, within group, and within populations. The result of mantel test showed a highly significant positive correlation (P < 0.01) between FST and geographical distance. A maximum likelihood tree showed that most haplotypes were grouped into three clusters corresponding to the three counties, suggesting a significant phylogeographic structure in the populations of G. qinghaiensis. The haplotype networks revealed that H2 may be the most primitive haplotype and the most adaptable in nature. Populations 7# and 8# had haplotype H2 and higher haplotype diversity; therefore, we speculated that the G. qinghaiensis in both populations were more adaptable to the environment and had greater outbreak potential and, therefore, should be focused on in terms of prevention and control. Our findings provide valuable information for further study of the population structure and phylogeny of G. qinghaiensis and provide a theoretical basis for the control of G. qinghaiensis.





Similar content being viewed by others
Abbreviations
- AMOVA:
-
Analysis of molecular variance
- COI:
-
Cytochrome oxidase subunit I
- F ST :
-
Genetic differentiation coefficient
- H :
-
Number of haplotypes per population
- h :
-
Haplotype diversity
- mv:
-
Possible mutation sites
- ML:
-
The maximum likelihood method
- Nm :
-
Gene flow
- n :
-
Number of individuals per sample site
- QTP:
-
The Qinghai-Tibetan Plateau
- π :
-
Nucleotide diversity
- PCR:
-
Polymerase chain reaction
References
Alyokhin A, Baker M, Mota-Sanchez D, Dively G, Grafius E (2008) Colorado potato beetle resistance to insecticides. Am J Potato Res 85:395–413
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecifific phylogenies. Mol Biol Evol 16:37–48
Bergl RA, Bradley BJ, Nsubuga AM, Vigilant L (2008) Effects of habitat fragmentation, population size and demographic history on genetic diversity: the cross river Gorilla in a comparative context. Am J Primatol 70:848–859
Chen HQ, Fan YQ, Chen KX (2014) Present situation and protection measures of grassland ecological environment in Yushu prefecture. Mod Agric Sci Technol 13:280–282
Chou Y, Ying XC (1979) A taxonomic study on the steppe caterpillars (Lepidoptera: Lymantriidae). Entomotaxonomia 1:23–28
Djebbi S, Amara WB, Bouktila D, Makni H, Makni M, Mezghani-Khemakhem M (2018) Assessment of Pea Weevil Bruchus pisorum (Coleoptera: Bruchidae) genetic diversity based on mitochondrial COI gene sequences. Afr Entomol 26:95–103
Excoffifier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Gallo-Franco JJ, Velasco-Cuervo SM, Aguirre-Ramirez E, Obando RG, Carrejo NS, Toro-Perea N (2017) Genetic diversity and population structure of Anastrepha striata (Diptera: Tephritidae) in three natural regions of southwestern Colombia using mitochondrial sequences. Genetica 145:79–89
Goldberg TL, Ruvolo M (1997) The geographic apportionment of mitochondrial genetic diversity in east African chimpanzees, Pantroglodytes schweinfurthii. Mol Biol Evol 14:976–984
Grant W, Bowen BW (1998) Shallow population histories in deep evolutionary lineages of marine fishes: insights from sardines and anchovies and lessons for conservation. J Hered 89:415–426
Islam SU, Qasim M, Ali H, Islam W, Arif M, Dash CK, Lin WZ, Du ZG, Wu ZJ (2008) Genetic diversity of the families Aeshnidae, Gomphidae and Libellulidae through COI gene from South China. Acta Trop 185:273–279
Kandul NP, Lukhtanov VA, Dantchenko AV, Coleman JWS, Sekercioglu CH, Haig D, Pierce NE (2004) Phylogeny of Agrodiaetus Hübner 1822 (Lepidoptera: Lycaenidae) inferred from mtDNA sequences of COI and COII and nuclear sequences of EF1-α: karyotype diversification and species radiation. Syst Biol 53:278–298
Kim M, Wan XL, Kim MJ, Jeong HC, Ahn NH, Kim KG (2010) Phylogenetic relationships of true butterflies (Lepidoptera: Papilionoidea) inferred from COI, 16S rRNA and EF-1α sequences. Mol Cells 30:409–425
Knaepkens G, Bervoets L, Verheyen E, Eens M (2004) Relationship between population size and genetic diversity in endangered populations of the European bullhead (Cottus gobio): implications for conservation. Biol Conserv 3:403–410
Kumar NR, Chang JC, Narayanan MB, Ramasamy S (2016) Phylogeographical structure in mitochondrial DNA of whitefly, Bemisia tabaci Gennadius (Hemiptera: Aleyrodidae) in southern India and Southeast Asia. Mitochondrial DNA Part A 5:621–631
Li JJ, Ye YY, Wu CW, Guo BY, Gul Y (2014) Genetic diversity and population structure of Sepiella japonica (Mollusca: Cephalopoda: Decapoda) inferred by mitochondrial DNA (COI) variations. Biochem Syst Ecol 56:8–15
Liang HW, Meng Y, Luo XZ, Li Z, Zou GW (2018) Genetic diversity of six Monopterus albus populations based on COI gene sequences. J Fish Sci China 25:837–846
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Liu X, Zhang GR (2017) Study on the sustainable development of cordyceps sinensis resources. Science Press, Beijing
Liu ZK, Yan L, Mei JR, Huo KK (1994) Investigation on species of grassland caterpillar in Qinghai province. J Qinghai Anim Husb Vet Med coll 1:26–28
Liu YQ, Zhou DY, Gong XY, Feng YN, He SY, Zhao WZ (2018) Mt DNA variation of Apis cerana populations in the Qinghai-Tibet plateau. J Shanxi Agric Univ 38:60–76
Ma GL (2007) The degradation of alpine grassland and its countermeasures in Yushu prefecture. Prataculture Anim Husb 6:33–35
Madsen T, Olsson M, Wittzell H, Stille B, Gullberg A, Shine R, Andersson S, Tegelström H (2000) Population size and genetic diversity in sand lizards (Lacerta agilis) and adders (Vipera berus). Biol Conserv 2:257–262
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Matern A, Desender K, Drees C, Gaublomme E, Paill W, Assmann T (2009) Genetic diversity and population structure of the endangered insect species Carabus variolosus in its western distribution range: Implications for conservation. Conserv Genet 10:391–405
Palraju M, Paulchamy R, Sundaram J (2018) Population genetic structure and molecular diversity of Leucinodes orbonalis based on mitochondrial COI gene sequences. Mitochondrial DNA Part A 29:1231–1239
Pérez-Asso AR, Núñez-Aguila R, Genaro JA (2016) Morphology and COI barcodes reveal four new species in the lycieus group of Calisto (Lepidoptera, Nymphalidae, Satyrinae). Zootaxa 4170:401–450
Roderick GK (1996) Geographic structure of insect populations: gene flow, phylogeography, and their uses. Annu Rev Entomol 41:325–352
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Shen H, Liu DY (2001) Summary of genetic diversity. J Biol 18:5–7
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 23:585–595
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Wan XL, Zhang WG (2006) Feeding habit and spatial pattern of Gynaephora Alpherakii larvae. Acta Agrestia Sinica 14:84–88
Wang LY (2013) Occurrence and control of grassland caterpillar. Grass Anim Husb 11:31–33
Wang HZ, Zhong X, Gu L, Li SS, Zhang GR, Liu X (2019a) Analysis of the Gynaephora qinghaiensis pupae immune transcriptome in response to parasitization by Thektogaster sp. Arch Insect Biochem Physiol 100:e21553
Wang SH, Zhang C, Shang M, Wu XG, Cheng YX (2019b) Genetic diversity and population structure of native mitten crab (Eriocheir sensu stricto) by microsatellite markers and mitochondrial COI gene sequence. Gene 693:101–113
Wright S (1931) Evolution in mendelian populations. Genetics 16:97–159
Xu DD, Lou B, Shi HL, Geng Z, Li SL, Zhang YR (2012) Genetic diversity and population structure of Nibea albiflflora in the China Sea revealed by mitochondrial COI sequences. Biochem Syst Ecol 45:158–165
Yan L (2006) Studies of taxonomy, geographic distribution in Gynaephora genus and life-history strategies on Gynaephora menyuanensis. PhD Thesis. Lanzhou: Lanzhou Universtiy, pp 15–22
Yuan ML, Zhang QL (2014) Phylogenetic relationships of Gynaephora (Lepidoptera: Lymantriidae) endemic to the Qinghai-Tibetan plateau inferred from the complete mitochondrial genomes [DB/OL]. Genbank. https://www.ncbi.nlm.nih.gov/nucleotide/KJ507133.1?report=genb. Accessed 20 Dec 2019
Yuan ML, Zhang QL, Wang ZF, Guo ZL, Bao GS (2015) Molecular phylogeny of grassland caterpillars (Lepidoptera: Lymantriinae: Gynaephora) endemic to the Qinghai-Tibetan Plateau. PLoS ONE 10:e0127257
Zhang QL, Yuan ML (2013) Research status and prospect of grassland caterpillars (Lepidoptera: Lymantriidae). Prataculture Sci 30:638–646
Zheng JH, Nie HT, Yang F, Yan XW (2019) Genetic variation and population structure of different geographical populations of Meretrix petechialis based on mitochondrial gene COI. J Genet 98:68
Acknowledgements
Thanks to Dr. Xin Zhong, Dr. Shaosong Li, Dr. Li Gu, Dr. Jiequn Yi, and Professor Guren Zhang for their guidance in data analysis. This work was supported by the Research Start-Up fund of Luliang University and the National Natural Science Foundation (Grant Number: 81503333).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no financial and nonfinancial conflicts of interests in this study.
Ethical approval
G.qinghaiensis is a widely distributed pest in the Qinghai-Tibetan Plateau, so this study does not involve any ethical issues.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, H., Zhong, X., Lin, H. et al. Genetic Diversity and Population Structure of Gynaephora qinghaiensis in Yushu Prefecture, Qinghai Province Based on the Mitochondrial COI Gene. Biochem Genet 59, 1396–1412 (2021). https://doi.org/10.1007/s10528-021-10065-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-021-10065-8