Abstract
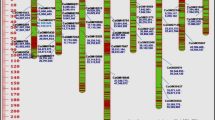
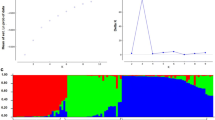
Genetic diversity and relationships within and among members of the primary gene pool of chickpea, including 38 accessions of Cicer arietinum, six of C. reticulatum,, and four of C. echinospermum, were investigated using 31 ISSR markers. The study revealed moderate diversity, detecting 141 fragments, of which 79 (56%) were polymorphic. Averages were 0.125 for polymorphic information content, 0.350 for marker index, and 0.715 for resolving power. The UPGMA dendrogram and the principal coordinate analysis revealed a clear differentiation between wild and cultivated accessions. The clustering pattern did not strictly follow the grouping of accessions by geographic origin but was in good agreement with the pedigree data and the seed type. The study demonstrates that ISSRs provide promising marker tools in revealing genetic diversity and relationships in chickpea and can contribute to efficient identification, conservation, and utilization of germplasm for plant improvement through conventional as well as molecular breeding approaches.






Similar content being viewed by others
References
Abbo S, Berger J, Turner NC (2003) Evolution of cultivated chickpea: four bottlenecks limit diversity and constrain adaptation. Funct Plant Biol 30:1081–1087
Ahmad F, Khan AI, Awan FS, Sadia B, Sadaqat HA, Bahadur S (2010) Genetic diversity of chickpea (Cicer arietinum L.) germplasm in Pakistan as revealed by RAPD analysis. Genet Mol Res 9:1414–1420
Berger J, Abbo S, Turner NC (2003) Ecogeography of annual Cicer species: the poor state of the world collection. Crop Sci 43:1076–1090
Bhagyawant SS, Srivastava N (2008) Genetic fingerprinting of chickpea (Cicer arietinum L.) germplasm using ISSR markers and their relationships. Afr J Biotechnol 7:4428–4431
Bohn MH, Utz F, Melchinger AE (1999) Genetic similarities among winter wheat cultivars determined on the basis of RFLPs, AFLPs and SSRs and their use for predicting progeny variance. Crop Sci 39:228–237
Choudhary P, Khanna SM, Jain PK, Bharadwaj C, Kumar J, Lakhera PC, Srinivasan R (2012) Genetic structure and diversity analysis of the primary gene pool of chickpea using SSR markers. Genet Mol Res 11:891–905
Choumane W, Winter P, Weigand F, Kahl G (2000) Conservation and variability of sequence tagged microsatellite sites (STMSs) from chickpea (Cicer aerietinum L.) within the genus Cicer. Theor Appl Genet 101:269–278
Chowdhury MA, Vandenberg B, Warkentin T (2002) Cultivar identification and genetic relationship among selected breeding lines and cultivars in chickpea (Cicer arietinum L.). Euphytica 127:317–325
Croser JS, Ahmad F, Clarke HJ, Siddique KHM (2003) Utilization of wild Cicer in chickpea improvement–progress, constraints and prospects. Aust J Agri Res 54:429–444
dos Santos LF, de Oliveira EJ, dos Santos Silva A, de Carvalho FM, Costa JL, Pádua JG (2011) ISSR markers as a tool for the assessment of genetic diversity in Passiflora. Biochem Genet 49:540–554
FAO (2010) Agriculture Data. United Nations Food and Agriculture Organization. http://faostat.fao.org/site/567/default.aspx. Accessed 1 Jan 2012
Fernandez E, Figueiras M, Benito C (2002) The use of ISSR and RAPD markers for detecting DNA polymorphism, genotype identification and genetic diversity among barley cultivars with known origin. Theor Appl Genet 104:845–851
Glaszmann JC, Kilian B, Upadhyaya HD, Varshney RK (2010) Accessing genetic diversity for crop improvement. Curr Opin Plant Biol 13:1–7
Grativol C, da Fonseca Lira-Medeiros C, Hemerly AS, Ferreira PCG (2011) High efficiency and reliability of inter-simple sequence repeats (ISSR) markers for evaluation of genetic diversity in Brazilian cultivated Jatropha curcas L. accessions. Mol Biol Rep 38:4245–4256
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Millan T, Clarke HJ, Siddique KHM, Buhariwalla HK, Gaur PM, Kumar J, Kahl G, Winter P (2006) Chickpea molecular breeding: new tools and concepts. Euphytica 147:81–103
Muminovic J, Melchinger AE, Lübberstedt T (2004) Genetic diversity in corn salad (Valerianella locusta) and related species as determined by AFLP markers. Plant Breed 123:460–466
Nguyen TT, Taylor PWJ, Redden RJ, Ford R (2004) Genetic diversity estimates in Cicer using AFLP analysis. Plant Breed 123:173–179
Powell W, Margenta M, Andre C, Hanfrey M, Vogel J, Tingey S, Rafalsky A (1996) The utility of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112
Rafalski JA, Vogel JM, Morgante M, Powell W, Andre C, Tingey SV (1996) Generating and using DNA markers in plants. In: Lai E (ed) Brain B. Nonmammalian Genomic Analysis, A Practical Guide, pp 75–134
Rao LS, Rani PU, Deshmukh PS, Kumar PA, Panguluri SK (2007) RAPD and ISSR fingerprinting in cultivated chickpea (Cicer arietinum L.) and its wild progenitor Cicer reticulatum Ladizinsky. Genet Resour Crop Evol 54:1235–1244
Ratnaparkhe MB, Santra DK, Tullu A, Muehlbauer FJ (1998) Inheritance of inter-simple-sequence-repeat polymorphisms and linkage with a fusarium wilt resistance gene in chickpea. Theor Appl Genet 96:348–353
Reddy MP, Sarla N, Siddiq EA (2002) Inter simple sequence repeat (ISSR) polymorphism and its application in plant breeding. Euphytica 128:9–17
Rohlf FJ (2000) NTsys-PC version 2.1: Numerical Taxonomy and Multivariate Analysis System. Exeter Publications, New York
Roldan-Ruiz I, Dendauw J, VanBockstaele E, Depicker A, De Loose M (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Mol Breed 6:125–134
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length in barley: mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sefera T, Abebie B, Gaur PM, Assefa K, Varshney RK (2011) Characterisation and genetic diversity analysis of selected chickpea cultivars of nine countries using simple sequence repeat (SSR) markers. Crop Pasture Sci 62:177–187
Singh R, Sharma P, Varshney RK, Sharma SK, Singh NK (2008) Chickpea improvement: role of wild species and genetic markers. Biotechnol Genet Eng Rev 25:267–314
Sudupak MA (2004) Inter- and intraspecies inter simple sequence repeat (ISSR) variation in the genus Cicer. Euphytica 135:229–238
Sudupak MA, Akkaya MS, Kence A (2002) Analysis of genetic relationships among perennial and annual Cicer species growing in Turkey using RAPD markers. Theor Appl Genet 105:1220–1228
Talebi R, Naji AM, Fayaz F (2008) Geographical patterns of genetic diversity in cultivated chickpea (Cicer arietinum L.) characterized by amplified fragment length polymorphism. Plant Soil Environ 54:447–452
Udupa SM, Sharma A, Sharma RP, Pai RA (1993) Narrow genetic variability in Cicer arietinum L. as revealed by RFLP analysis. J Plant Biochem Biotechol 2:83–86
Upadhyaya HD, Dwivedi SL, Baum M, Varshney RK, Udupa SM, Gowda CLL, Hoisington DA, Singh S (2008) Genetic structure, diversity, and allelic richness in composite collection and reference set in chickpea (Cicer arietinum L.). BMC Plant Biol 8:106
Upadhyaya HD, Thudi M, Dronavalli N, Gujaria N, Singh S, Sharma S, Varshney RK (2011) Genomic tools and germplasm diversity for chickpea improvement. Plant Genet Resour 9:45–58
Varshney RK, Horres R, Molina C, Nayak S, Jungmann R, Swamy P, Winter P, Jayashree B, Kahl G, Hoisington DA (2007) Extending the repertoire of microsatellite markers for genetic linkage mapping and germplasm screening in chickpea. J SAT Agri Res 5(1):3
Acknowledgments
The authors gratefully acknowledge the Indian Council of Agricultural Research (ICAR) Network Project on Transgenics in Crops (Functional Genomics component) for providing the financial support for this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Choudhary, P., Khanna, S.M., Jain, P.K. et al. Molecular Characterization of Primary Gene Pool of Chickpea Based on ISSR Markers. Biochem Genet 51, 306–322 (2013). https://doi.org/10.1007/s10528-012-9564-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-012-9564-7