Abstract
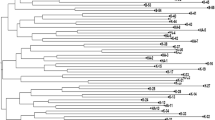
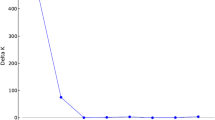
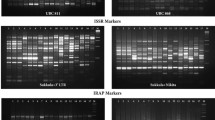
We report here on the phylogenetic analysis, population substructure, and identification of molecular tags of 25 popular rice varieties and four landraces from different ecological belts of India employing a set of 52 simple sequence repeat (SSR) markers. Genetic analysis using the SSR markers categorized the genotypes into two major clusters, distributed according to their pedigree. Population structure analysis suggested that the optimum number of subpopulations was three (K = 3) in the popular varieties and landraces. At K = 5 the allelic distribution was much more similar to the phylogenetic dendrogram. The molecular diversity and population structure analysis indicated that there is not much variation among the popular rice cultivars of India. The study has identified SSR markers producing unique alleles, which should aid in the precise identification, maintenance, and genetic purity analysis of rice varieties.

Similar content being viewed by others
References
Agrama HA, Eizenga GC (2008) Molecular diversity and genome-wide linkage disequilibrium pattern in worldwide rice and its wild relatives. Euphytica 160:339–355
Agrama HA, Eizenga GC, Yan W (2007) Association mapping of yields and its components in rice cultivars. Mol Breed 19:341–356
Aranzana MJ, Abbasi E, Howard W, Arus P (2010) Genetic variation, population structure and linkage disequilibrium in peach commercial varieties. BMC Genet 11:69
Bao JS, Corke H, Sun M (2006) Analysis of genetic diversity and relationship in waxy rice (Oryza sativa L.) using AFLP and ISSR marker. Genet Resor Crop Evol 53:323–330
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software Structure: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Garris A, McCouch S, Kresovich S (2003) Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.). Genetics 165:759–769
Giarrocco LE, Marassi MA, Salerno GL (2007) Assessment of the genetic diversity in Argentine rice cultivars with SSR markers. Crop Sci 47:853–860
Hartl DL, Clark AG (1997) Principles of population genetics, 3rd edn. Sinauer Associates, Sunderland
Huang M, Xie F, Chen L, Zhao X, Jojee L, Madonna D (2010) Comparative analysis of genetic diversity and structure in rice using ILP and SSR markers. Rice Sci 17(4):257–268
Jin L, Yan L, Peng X, Mei S, Harold C, Jinsong B (2010) Genetic diversity and population structure of a diverse set of rice germplasm for association mapping. Theor Appl Genet 121:175–487
Liu K, Muse S (2005) PowerMarker: new genetic data analysis software, version 2.7. http://www.powermarker.net. Accessed 15 Jan 2011
Loveless MD, Hamrick JL (1984) Ecological determinants of genetic structure in plant populations. Ann Rev Ecol Syst 15:65–95
Lu BR, Snow AA (2005) Gene flow from genetically modified rice and its environmental consequences. Bioscience 55:669–678
Mackill DJ (1995) Classifying japonica rice cultivars with RAPD markers. Crop Sci 35(3):889–894
McNally KL, Childs KL, Bohnert R, Davidson RM, Zhao K, Ulat VJ, Zeller G, Clark RM, Hoen DR, Bureau T, Stokowshi R, Ballinger DG, Frazer KA, Cox DR, Padhukasahasram B, Bustamante CD, Weigel D, Mackill DJ, Bruskiewich CD, Ratsch G, Buell CR, Leung H, Leach JE (2009) Genomewide SNP variation reveals relationships among landraces and modern varieties of rice. Proc Natl Acad Sci USA 106:12273–12278
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/darwin. Accessed 20 Jan 2011
Prabhu KV, Somers DJ, Rakow G, Gugel RK (1998) Molecular markers linked to white rust resistance in mustard Brassica juncea. Theor Appl Genet 97:865–870
Pritchard JK, Wen W (2003) Documentation for the Structure software Version 2. Chicago. http://www.pritch.bsd.uchicagoedu/software/structure2_1.html. Accessed 11 Feb 2011
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rabbani MA, Iwabuchi A, Murakami Y, Suzuki T, Takayanagi K (1998) Genetic diversity in mustard (Brassica juncea L.) germplasm from Pakistan as determined by RAPDs. Euphytica 103:235–242
Singh RK, Singh US, Khush GS, Rohilla R, Singh JP, Singh G, Shekhar KS (2000) Small and medium grained aromatic rices of India. In: Singh RK, Singh US, Khush GS (eds) Aromatic rices. Oxford & IBH, New Delhi, pp 155–177
Singh VK, Upadhyay P, Sinha P, Mall AK, Jaiswal SK, Singh A, Ellur RK, Biradar S, Sundaram RM, Singh S, Ahmed I, Mishra B, Singh AK, Kole CR (2010) Determination of genetic relationships among elite thermo-sensitive genic male sterile lines (TGMS) of rice (Oryza sativa L.) employing morphological and simple sequence repeat (SSR) markers. J Genet 90(1):11–19
Stich B, Melchinger AE, Frisch M, Maurer HP, Heckenberger M, Reif JC (2005) Linkage disequilibrium in European elite maize germplasm investigated with SSRs. Theor Appl Genet 111:723–730
Sun CQ, Wang XK, Li ZC, Yoshimura A, Iwata N (2001) Comparison of the genetic diversity of common wild rice (Oryza rufipogan Griff.) and cultivated rice (Oryza sativa L.) using RFLP markers. Theor Appl Genet 102:152–162
Sundaram RM, Naveenkumar B, Biradar SK, Balachandran SM, Mishra B, IlyasAhmed M, Viraktamath BC, Ramesha MS, Sarma NP (2008) Identification of informative SSR markers capable of distinguishing hybrid rice parental lines and their utilization in seed purity assessment. Euphytica 163:215–224
Tam SM, Mhiri C, Vogelaar A, Kerkveld M, Pearce SR, Grandbastein MA (2005) Comparative analysis of genetic diversities within tomato and pepper collections detected by retrotransposon-based SSAP, AFLP and SSR. Theor Appl Genet 110:819–831
Upadhyay P, Singh VK, Neeraja CN (2011) Identification of genotype specific alleles and molecular diversity assessment of popular rice (Oryza sativa L.) varieties of India. Int J Plant Breed Genet 5(2):130–140
Yu J, Hu S, Wang J (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296:79–92
Zhao WG, Chung JW, Ma KH, Kim TS, Kim SM, Shin DI, Kim CH, Koo HM, Park YJ (2009) Analysis of genetic diversity and population structure of rice cultivars from Korea, China and Japan using SSR markers. Gene Genome 31:283–292
Zhu MY, Wang YY, Zhu YY, Lu BR (2004) Estimating genetic diversity of rice landraces from Yunnan by SSR assay and its implication for conservation. Acta Bot Sin 46:1458–1467
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Upadhyay, P., Neeraja, C.N., Kole, C. et al. Population Structure and Genetic Diversity in Popular Rice Varieties of India as Evidenced from SSR Analysis. Biochem Genet 50, 770–783 (2012). https://doi.org/10.1007/s10528-012-9519-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-012-9519-z