Abstract
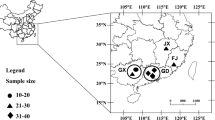
Genetic diversity and population structure of Sipunculus nudus were evaluated using a 652 base pair fragment of the mitochondrial cytochrome oxidase I gene. The populations were collected from Beihai, Sanya, and Xiamen. A total of 71 polymorphic sites defined 16 distinct haplotypes. The mean haplotype diversity and nucleotide diversity of the three populations were 0.9354 ± 0.0168 and 0.0035 ± 0.0018, respectively. Analysis at the intrapopulation level showed that the Beihai population had the greatest haplotype and nucleotide diversity, followed by the Xiamen and Sanya populations. Analysis of molecular variance showed significant genetic differentiation among the three populations (Fst = 0.0796, P < 0.05). The present results revealed that S. nudus populations had a high level of genetic diversity and distinct population structures.


Similar content being viewed by others
References
Avise JC (1994) Molecular markers, natural history and evolution. Chapman & Hall, New York
Avise JC, Hamrick JL (1996) Conservation genetics: case histories from nature. Chapman & Hall, New York
Brown WM, George M Jr, Wilson AC (1979) Rapid evolution of animal mitochondrial DNA. Proc Natl Acad Sci USA 76:1967–1971
Cassone BJ, Boulding EG (2006) Genetic structure and phylogeography of the lined shore crab, Pachygrapsus crassipes, along the northeastern and western Pacific Coasts. Mar Biol 149:213–226
Du XD, Chen ZA, Deng YW, Wang QH, Huang RL (2008) Genetic diversity of three wild populations of Sipunculus nudus in South China Sea as inferred from mitochondrial 16S rRNA sequences. Israeli J Aquac 60(4):237–242
Excoffier LG, Laval G, Schneider S (2005) Arlequin version 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome coxidase subunit I from diverse metozoan invertebrates. Mol Mar Biol Biotechnol 3:294–299
Hare MP, Avise JC (1996) Molecular genetic analysis of a stepped multilocus cline in the American oyster (Crassostrea virginica). Evolution 50:2305–2315
Inoue N, Watanabe H, Kojima S, Sekiguchi H (2007) Population structure of Japanese spiny lobster Panulirus japonicus inferred by nucleotide sequence analysis of mitochondrial COI gene. Fish Sci 73(3):550–556
Kumar S, Tamura K, Nei M (2004) Mega 3.0: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Kyle CJ, Boulding EG (2000) Comparative population genetic structure of marine gastropods (Littorina spp.) with and without pelagic larval dispersal. Mar Biol 137:835–845
Lan GB, Liao SM, Yan B (2007) Effect of water temperature on larval development and metamorphosis of Sipunculus nudus. J Fish China 31(5):633–638 (In Chinese)
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Tajima F (1981) DNA polymorphism detectable by restriction endonucleases. Genetics 97(1):145–163
Palumbi SR, Wilson AC (1990) Mitochondrial DNA diversity in the sea urchins Strongycogentrotus purpuratus and S. droebachiensis. Evolution 44:403–415
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wang QH, Du XD, Li K (2006) Genetic diversity of Sipunculus nudus as revealed by RAPD. Mar Fish Res 27(3):57–61 (In Chinese)
Wörheide G (2006) Low variation in partial cytochrome oxidase subunit I (COI) mitochondrial sequences in the coralline demosponge Astrosclera willeyana across the Indo-Pacific. Mar Biol 148:907–912
Wright S (1978) Evolution and the genetics of populations: variability within and among natural populations, vol 4. University of Chicago Press, Chicago
Yu ZN, Kong XY, Guo TH, Jiang YY, Zhuang ZM, Jin XS (2005) Mitochondrial DNA sequence variation of Japanese anchovy Engraulis japonicus from the Yellow Sea and East China Sea. Fish Sci 71:299–307
Yu ZN, Wei XH, Kong XY, Yu SS (2007) Application of the first internal transcribed spacer (ITS-1) of ribosomal DNA as a molecular marker to population analysis in Farrer’s scallop Chlamys farreri. Acta Oceanol Sin 26(1):93–100
Acknowledgments
The work was financially supported by the Natural Science Foundation of Guangdong Province under contract No. 30371026. The authors are grateful to the reviewers for critical comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Du, X., Chen, Z., Deng, Y. et al. Comparative Analysis of Genetic Diversity and Population Structure of Sipunculus nudus as Revealed by Mitochondrial COI Sequences. Biochem Genet 47, 884–891 (2009). https://doi.org/10.1007/s10528-009-9291-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-009-9291-x