Abstract
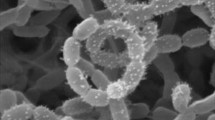
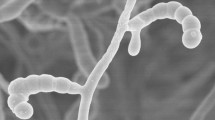
The taxonomic position of a Streptomyces strain isolated from a hyper-arid desert soil was established using a polyphasic approach. The organism had chemical and morphological properties typical of the genus Streptomyces and formed a phyletic line at the periphery of the Streptomyces coeruleorubidus subcluster in the 16S rRNA gene tree. DNA:DNA relatedness values between the isolate and its nearest phylogenetic neighbours, Streptomyces lomondensis NRRL 3252T and Streptomyces lusitanus NRRL B-12501T were 42.5 (±0.48)% and 25.0 (±1.78)%, respectively. The isolate was readily distinguished from these organisms using a combination of morphological and phenotypic properties. On the basis of these results, it is proposed that isolate C63T (CGMCC 4.6997T, = KACC 15425T) be classified as the type strain of Streptomyces deserti sp. nov.


Similar content being viewed by others
References
Chun J, Goodfellow M (1995) A phylogenetic analysis of the genus Nocardia with 16S rRNA gene sequences. Int J Syst Bacteriol 45:240–245
Chun J, Lee J-H, Jung Y, Kim M, Kim S, Kim B-K, Lim Y-W (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Collins MD (1985) Isoprenoid quinone analysis in bacterial classification and identification. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic Press, London, pp 267–287
De Ley J, Cattour H, Reynaerts A (1970) The qualitative measurement of DNA hybridization from renaturation rates. J Biochem 12:133–142
Euzéby JP (2011) List of bacterial names with standing in nomenclature: a folder available on the Internet. Last full update 9 Sep 2011. http://www.bacterio.cict.fr/
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Goodfellow M, Fiedler HP (2010) A guide to successful bioprospecting: informed by actinobacterial systematics. Antonie van Leeuwenhoek 98:119–142
Hasegawa T, Takizawa M, Tanida S (1983) A rapid analysis for chemical grouping of aerobic actinomycetes. J Gen Appl Microbiol 29:319–322
Jones KL (1949) Fresh isolates of actinomycetes in which the presence of sporogenous aerial mycelia is a fluctuating characteristic. J Bacteriol 57:141–145
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, London, pp 21–132
Kawato M, Shinobu R (1959) On Streptomyces herbaricolor sp. nov., supplement: a simple technique for microscopical observation. Mem Osaka Univ Lib Arts Educ B 8:114–119
Kumar Y, Goodfellow M (2010) Reclassification of Streptomyces hygroscopicus strains as Streptomyces aldersoniae sp. nov., Streptomyces angustmyceticus sp. nov., comb. nov., Streptomyces ascomycinicus sp. nov., Streptomyces decoyicus sp. nov., comb. nov., Streptomyces milbemycinicus sp. nov. and Streptomyces wellingtoniae sp. nov. Int J Syst Evol Microbiol 60:769–775
Labeda DP, Goodfellow M, Brown R, Ward AC, Lanoot C, Vanncanneyt M, Swings J, Kim S-B, Liu Z, Chun J, Tamura T, Oguchi A, Kikuchi T, Kikuchi H, Nishii T, Tsuji K, Yamaguchi Y, Tase A, Takahashi M, Sakane T, Suzuki KI, Hatano K (2011) Phylogenetic study of the species within the family Streptomycetaceae. Antonie van Leeuwenhoek (in press)
Manfio GP, Zakrzewska-Czerwinska J, Atalan E, Goodfellow M (1995) Towards minimal standards for the description of Streptomyces species. Biotechnologia 7–8:242–253
Mao J, Wang J, Dai H-Q, Zhang Z-D, Tang Q-Y, Ren B, Yang N, Goodfellow M, Zhang L-X, Liu Z-H (2011) Yuhushiella deserti gen. nov., sp. nov., a new member of the suborder Pseudonocardineae. Int J Syst Evol Microbiol 61:621–630
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Nachtigall J, Kulik A, Bull AT, Goodfellow M, Arenjo JA, Maier A, Wiese J, Inhoff JF, Süssmuth R-D, Fiedler H-P (2011) Atacamycins A-C, 22 membered antitumor macrolide derivatives produced by Streptomyces sp. C38. J Antibiot (in press)
O’Donnell AG, Falconer C, Goodfellow M, Ward AC, Williams E (1993) Biosystematics and diversity amongst novel carboxydotrophic actinomycetes. Antonie van Leeuwenhoek 64:325–340
Okoro CK, Brown R, Jones AL, Andrews BA, Asenjo JA, Goodfellow M, Bull AT (2009) Diversity of culturable actinomycetes in hyper-arid soils of the Atacama Desert, Chile. Antonie van Leeuwenhoek 95:121–133
Okoro CK, Bull AT, Mutreja A, Rong X, Huang Y, Goodfellow M (2010) Lechevalieria atacamensis sp. nov., Lechevalieria deserti sp. nov. and Lechevalieria roselyniae sp. nov., isolated from hyper-arid soils. Int J Syst Evol Microbiol 60:296–300
Rateb ME, Houssen WE, Arnold M, Abdelrahman M-H, Deng H, Harrison WTA, Okoro CK, Asenjo JA, Andrews BA, Ferguson G, Bull AT, Goodfellow M, Ebel R, Jaspars M (2011) Chaxamycins A–D, bioactive ansamycins from a hyper-arid desert Streptomyces sp. J Nat Prod 74:1491–1499
Rzhetsky A, Nei M (1992) A simple method for estimating and testing minimum evolution trees. Mol Biol Evol 9:945–967
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids, MIDI Technical Note 101. MIDI Inc, Newark
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Evol Microbiol 16:313–340
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Vickers JC, Williams ST, Ross GW (1984) A taxonomic approach to selective isolation of streptomycetes from soil. In: Ortiz-Ortiz L, Bojalil LF, Yakoleff V (eds) Biological, biochemical and biomedical aspects of actinomycetes. Academic Press, Orlando, pp 553–561
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray R-G-E et al (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Williams ST, Goodfellow M, Alderson G, Wellington EMH, Sneath PHA, Sackin MJ (1983) Numerical classification of Streptomyces and related genera. J Gen Microbiol 129:1743–1813
Williams ST, Goodfellow M, Alderson G (1989) Genus Streptomyces Waksman and Henrici 1943, 339AL. In: Williams ST, Sharpe ME, Holt JG (eds) Bergey’s manual of systematic bacteriology, vol 4. Williams & Wilkins, Baltimore, pp 2452–2492
Acknowledgments
A.T.B. thanks the Leverhulme Trust for an Emeritus Fellowship, and A.T.B. and J.A.S. thank the Royal Society for International Joint Project Grant JP100654 and funded in part by the National Science Foundation of China (grant 31100003).
Author information
Authors and Affiliations
Corresponding author
Additional information
The GenBank accession number for the 16S rRNA gene sequence of Streptomyces deserti C63T is HE577172.
Rights and permissions
About this article
Cite this article
Santhanam, R., Okoro, C.K., Rong, X. et al. Streptomyces deserti sp. nov., isolated from hyper-arid Atacama Desert soil. Antonie van Leeuwenhoek 101, 575–581 (2012). https://doi.org/10.1007/s10482-011-9672-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-011-9672-0