Abstract
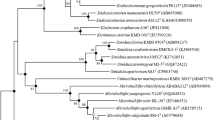
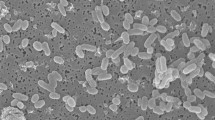
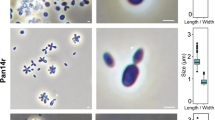
A novel species in the genus Candida was obtained from deep-sea hydrothermal fields on the Mid-Atlantic Ridge. Strains Mo39, MARY089 and CBS 5307, respectively, isolated from an unidentified deep-sea coral collected near Rainbow hydrothermal vent, from water samples near Menez Gwen hydrothermal field and from the stomach of a marine fish are considered as a novel taxon. Sequence similarities in the D1/D2 region of the 26S rRNA gene indicated that strains Mo39, MARY089 and CBS 5307 have for closest neighbors Candida spencermartinsiae, Candida taylorii, Candida atmosphaerica and Candida atlantica. The strains, respectively, differ from C. spencermartinsiae, C. taylorii, C. atmosphaerica andCandida atlantica by 4, 4.3, 4.3 and 4.7% in the D1/D2 domain. Strains Mo39, MARY089 and CBS 5307 were differentiated from others by differences in the ability to assimilate d-Gluconate and in the ability to grow at relatively high temperature. Only strain Mo39 displays an optimal growth at 3% sea salts, indicating that this strain is clearly adapted to live in marine conditions. Sequence similarities between strains Mo39, MARY089 and CBS 5307 and related species and differences in the ability to utilize specific carbon compounds revealed that these strains represent a hitherto unknown species. Sexual reproduction was not observed in strains Mo39, MARY089 and CBS 5307. An anamorphic name Candida oceani sp. nov. is proposed for the type strain Mo39T (= CBS 11857T = DSM 23777T) and the two other strains MARY089 and CBS 5307. To our knowledge, this is the first description of a micro-eukaryotic organism including a strain isolated from a deep-sea coral near a hydrothermal ecosystem.



Similar content being viewed by others
References
Burgaud G, Le Calvez T, Arzur D, Vandenkoornhuyse P, Barbier G (2009) Diversity of culturable marine filamentous fungi from deep-sea hydrothermal vents. Environ Microbiol 11:1588–1600
Burgaud G, Arzur D, Durand L, Cambon-Bonavita MA, Barbier G (2010) Marine culturable yeasts in deep-sea hydrothermal vents: species richness and association with fauna. FEMS Microbiol Ecol 73:121–133
Gadanho M, Sampaio J (2005) Occurrence and diversity of yeasts in the Mid-Atlantic Ridge hydrothermal fields near the azores archipelago. Microb Ecol 50:408–417
Guindon S, Lethiec F, Duroux P, Gascuel O (2005) PHYML online—a web server for fast maximum likelihood-based phylogenetic inference. Nucleic Acids Res 33:557–559
Kohlmeyer J, Kohlmeyer E (1979) Marine Mycology: the Higher Fungi. Academic Press, New York, USA
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 73:331–371
Kurtzman CP, Suzuki M (2010) Phylogenetic analysis of ascomycete yeasts that form coenzyme Q-9 and the proposal of the new genera Babjeviella, Meyerozyma, Millerozyma, Priceomyces, and Scheffersomyces. Mycoscience 51:2–14
Lachance MA, Starmer WT (1998) Ecology and yeasts. In: Kurtzman CP, Fell JW (eds) The yeasts, a taxonomic study, 4th edn. Elsevier Science B.V, Amsterdam, pp 21–30
Meyer SA, Payne RW, Yarrow D (1998) Candida Berkhout. In: Kurtzman CP, Fell JW (eds) The yeasts, a taxonomic study, 4th edn. Elsevier, Amsterdam, pp 454–573
Nagahama T (2005) Yeast biodiversity in freshwater, marine and deep-sea environments. In: Rosa C and Peter G (eds) The yeasts handbook. Springer, Berlin, Heidelberg, New York, pp 241–262
Nagahama T, Hamamoto M, Nakase T (1999) Kluyveromyces nonfermentans sp. nov. a new yeast species isolated from the deep sea. Int J of Syst Evol Micr 49:1899–1905
Nagahama T, Hamamoto M, Nakase T, Horikoshi K (2001) Rhodotorula lamellibrachii sp. nov. a new yeast species from a tubeworm collected at the deep-sea floor in Sagami Bay and its phylogenetic analyses. Antonie Van Leeuwenhoek 80:317–323
Nagahama T, Hamamoto M, Nakase T, Takaki Y, Horikoshi K (2003a) Cryptococcus surugaensis sp. nov. a novel yeast species from sediment collected on the deep-sea floor of Suruga Bay. Int J of Syst Evol Micr 53:2095–2098
Nagahama T, Hamamoto M, Nakase T, Horikoshi K (2003b) Rhodotorula benthica sp. nov. and Rhodotorula calyptogenae sp. nov. novel yeast species from animals collected from the deep-sea floor and Rhodotorula lysiniphila sp. nov. which is related phylogenetically. Int J of Syst Evol Micr 53:897–903
Nagahama T, Hamamoto M, Horikoshi K (2006) Rhodotorula pacifica sp. nov. a novel yeast species from sediment collected on the deep-sea floor of the north-west Pacific Ocean. Int J of Syst Evol Micr 56:295–299
Nagahama T, Abdel-Wahab MA, Nogi Y, Miyazaki M, Uematsu K, Hamamoto M, Horikoshi K (2008) Dipodascus tetrasporeus sp. nov. an ascosporogenous yeast isolated from deep-sea sediments in the Japan Trench. Int J of Syst Evol Micr 58:1040–1046
Nguyen HV, Gaillardin C, Neuvéglise C (2009) Differentiation of Debaryomyces hansenii and Candida famata by rRNA gene intergenic spacer fingerprinting and reassessment of phylogenetic relationships among D. hansenii, C. famata, D. fabryi, C. flareri (= D. subglobosus) and D. prosopidis: description of D. vietnamensis sp. nov. closely related to D. nepalensis. FEMS Yeast Res 9:641–662
Posada D, Crandall K (1998) Applications note. MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Ronquist F, Huelsenbeck J (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Statzell-Tallman A, Scorzetti G, Fell JW (2010) Candida spencermartinsiae sp. nov., Candida taylorii sp. nov. and Pseudozyma abaconensis sp. nov., novel yeasts from mangrove and coral reef ecosystems. Int J of Syst Evol Micr 60:1978–1984
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) Software version 4.0. Mol Biol Evol 24:1596
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
White T, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. Chap 38. PCR Protocols: a guide to methods and applications. Orlando, Florida: Academic Press. pp 315–322
Yarrow D (1998) Methods for the isolation, maintenance and identification of yeasts. In: Kurtzman CP, Fell JW (eds) The yeasts, a taxonomic study, 4th edn. Elsevier B.V, Amsterdam, The Netherlands, pp 77–100
Acknowledgments
We thank the chief scientist of the MoMARDREAM-Naut cruise, pilots and support crews of oceanographic vessels and Deep Submergence Vehicles of Ifremer. We greatly acknowledge Justine Amborski for valuable help for latin diagnosis. We thank all members of the GDR Ecchis for discussions and suggestions. We would like to thank the editor and the anonymous reviewers who have provided helpful comments on the refinement of this manuscript. We also thank ANR Deep-Oases and French Research Ministry for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Burgaud, G., Arzur, D., Sampaio, J.P. et al. Candida oceani sp. nov., a novel yeast isolated from a Mid-Atlantic Ridge hydrothermal vent (−2300 meters). Antonie van Leeuwenhoek 100, 75–82 (2011). https://doi.org/10.1007/s10482-011-9566-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-011-9566-1