Abstract
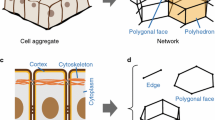
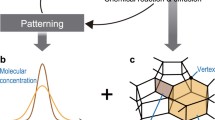
Morphogenesis of tissues in organ development is accompanied by large three-dimensional (3D) deformations, in which mechanical interactions among multiple cells are spatiotemporally regulated. To reveal the deformation mechanisms, in this study, we developed the reversible network reconnection (RNR) model. The model is developed on the basis of 3D vertex model, which expresses a multicellular aggregate as a network composed of vertices. 3D vertex models have successfully simulated morphogenetic dynamics by expressing cellular rearrangements as network reconnections. However, the network reconnections in 3D vertex models can cause geometrical irreversibility, energetic inconsistency, and topological irreversibility, therefore inducing unphysical results and failures in simulating large deformations. To resolve these problems, we introduced (1) a new definition of the shapes of polygonal faces between cellular polyhedrons, (2) an improved condition for network reconnections, (3) a new condition for potential energy functions, and (4) a new constraint condition for the shapes of polygonal faces that represent cell–cell boundaries. Mathematical and computational analyses demonstrated that geometrical irreversibility, energetic inconsistency, and topological irreversibility were resolved by suppressing the geometrical gaps in the network and avoiding the generation of irreversible network patterns in reconnections. Lastly, to demonstrate the applicability of the RNR model, we simulated tissue deformation of growing cell sheets and showed that our model can simulate large tissue deformations, in which large changes occur in the local curvatures and layer formations of tissues. Thus, the RNR model enables in silico recapitulation of complex tissue morphogenesis.
Similar content being viewed by others
References
Bellaiche Y, Segalen M (2009) Cell division orientation and planar cell polarity pathways. Semin Cell Dev Biol 20(8): 972–977
Blanchard GB, Murugesu S et al (2010) Cytoskeletal dynamics and supracellular organisation of cell shape fluctuations during dorsal closure. Development 137(16): 2743–2752
Bragg L, Nye JF (1947) A dynamical model of a crystal structure. Proc R Soc Lond A Math Phys Sci 190(1023): 474–481
Davies JA (2005) Mechanisms of morphogenesis: the creation of biological form. Elsevier Academic Press, Burlington
Eiraku M, Takata N et al (2011) Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472(7341): 51–56
Eiraku M, Adachi T et al (2012) Relaxation-expansion model for self-driven retinal morphogenesis. Bioessays 34(1): 17–25
Farhadifar R, Röper JC et al (2007) The influence of cell mechanics, cell–cell interactions, and proliferation on epithelial packing. Curr Biol 17(24): 2095–2104
Forgacs G, Newman SA (2005) Biological physics of the developing embryo. Cambridge University Press, New York
Friedlander DR, Mège RM et al (1989) Cell sorting-out is modulated by both the specificity and amount of different cell-adhesion molecules (CAMs) expressed on cell-surfaces. Proc Natl Acad Sci USA 86(18): 7043–7047
Fuchizaki K, Kusaba T et al (1995) Computer modelling of three-dimensional cellular pattern growth. Philos Mag B 71(3): 333–357
Heisenberg CP, Tada M et al (2000) Silberblick/Wnt11 mediates convergent extension movements during zebrafish gastrulation. Nature 405(6782): 76–81
Honda H, Ogita Y et al (1982) Cell movements in a living mammalian tissue: long-term observation of individual cells in wounded corneal endothelia of cats. J Morphol 174(1): 25–39
Honda H, Tanemura M et al (2004) A three-dimensional vertex dynamics cell model of space-filling polyhedra simulating cell behavior in a cell aggregate. J Theor Biol 226(4): 439–453
Honda H, Motosugi N et al (2008) Computer simulation of emerging asymmetry in the mouse blastocyst. Development 135(8): 1407–1414
Honda H, Nagai T et al (2008) Two different mechanisms of planar cell intercalation leading to tissue elongation. Dev Dyn 237(7): 1826–1836
Ingber DE, Mammoto T (2010) Mechanical control of tissue and organ development. Development 137(9): 1407–1420
Inoue Y, Adachi T (2011) Coarse-grained Brownian ratchet model of membrane protrusion on cellular scale. Biomech Model Mechanobiol 10(4): 495–503
Jamali Y, Azimi M, et al (2010) A sub-cellular viscoelastic model for cell population mechanics. PLoS One 5(8)
Kikuchi R (1956) Shape distribution of two-dimensional soap froths. J Chem Phys 24(4): 861–867
Lecuit T, Lenne PF (2007) Cell surface mechanics and the control of cell shape, tissue patterns and morphogenesis. Nat Rev Mol Cell Biol 8(8): 633–644
Lecuit T, Rauzi M et al (2008) Nature and anisotropy of cortical forces orienting Drosophila tissue morphogenesis. Nat Cell Biol 10(12): 1401–1410
Lecuit T, Rauzi M et al (2010) Planar polarized actomyosin contractile flows control epithelial junction remodelling. Nature 468(7327): 1110–1114
Letizia A, Sotillos S et al (2011) Regulated Crb accumulation controls apical constriction and invagination in Drosophila tracheal cells. J Cell Sci 124(2): 240–251
Martin AC, Kaschube M et al (2009) Pulsed contractions of an actin-myosin network drive apical constriction. Nature 457(7228): 495–499
Nagai T, Honda H (2001) A dynamic cell model for the formation of epithelial tissues. Philos Mag B 81(7): 699–719
Nagai T, Honda H (2009) Computer simulation of wound closure in epithelial tissues: Cell–basal-lamina adhesion. Phys Rev E Stat Nonlin Soft Matter Phys 80(6): 061903
Nagai T, Kawasaki K et al (1988) Vertex dynamics of two-dimensional cellular-patterns. J Phys Soc Jpn 57(7): 2221–2224
Nagai T, Ohta S et al (1990) Computer simulation of cellular pattern growth in two and three dimensions. Phase Transit 28: 177–211
Odell GM, Oster G et al (1981) The mechanical basis of morphogenesis. 1. Epithelial folding and invagination. Dev Biol 85(2): 446–462
Rauzi M, Verant P et al (2008) Nature and anisotropy of cortical forces orienting Drosophila tissue morphogenesis. Nat Cell Biol 10(12): 1401–1410
Solon J, Kaya-Copur A et al (2009) Pulsed forces timed by a ratchet-like mechanism drive directed tissue movement during dorsal closure. Cell 137(7): 1331–1342
Staple DB, Farhadifar R et al (2010) Mechanics and remodelling of cell packings in epithelia. Eur Phys J E Soft Matter 33(2): 117–127
Taniguchi K, Maeda R et al (2011) Chirality in planar cell shape contributes to left-right asymmetric epithelial morphogenesis. Science 333(6040): 339–341
VanEssen DC (1997) A tension-based theory of morphogenesis and compact wiring in the central nervous system. Nature 385(6614): 313–318
Weaire D, Kermode JP (1983) Computer-simulation of a two-dimensional soap froth. 1. Method and motivation. Philos Mag B 48(3): 245–259
Weaire D, Kermode JP (1984) Computer-simulation of a two-dimensional soap froth. 2. Analysis of results. Philos Mag B 50(3): 379–395
Weliky M, Oster G (1990) The mechanical basis of cell rearrangement. 1. Epithelial morphogenesis during fundulus epiboly. Development 109(2): 373–386
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
The Below is the Electronic Supplementary Material.
Rights and permissions
About this article
Cite this article
Okuda, S., Inoue, Y., Eiraku, M. et al. Reversible network reconnection model for simulating large deformation in dynamic tissue morphogenesis. Biomech Model Mechanobiol 12, 627–644 (2013). https://doi.org/10.1007/s10237-012-0430-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10237-012-0430-7