Abstract
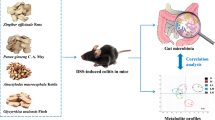
Ulcerative colitis (UC) and Crohn’s disease (CD) are two major forms of inflammatory bowel disease (IBD). The disease has been linked with gut microbiota dysbiosis in which the balance of commensal communities is disrupted. Accumulating evidence demonstrates that treatment with biologically active compounds can modulate gut microbiota composition in animal models. Our previous work has also shown the beneficial effect of Luem Pua (LP) rice extract, which is rich in anthocyanins, on inflammation. However, its effect on gut microbiota is yet to be explored. In this study, we profiled fecal microbiota of acetic acid (AA)–induced UC and indomethacin (ID)–induced CD rat models with and without pretreatment with LP rice extract by 16S rRNA gene sequencing. The results showed that gut microbiota communities of rats were altered by both AA-induced UC and ID-induced CD. The relative abundances of beneficial bacteria, especially the Lachnospiraceae NK4A136 group and Lactobacillus, were decreased in the AA-induced UC model, while some opportunistic pathogens (Bacteroides, Escherichia/Shigella, Fusobacterium, and Veillonella) were raised by ID-induced CD. Interestingly, pretreatment with LP rice extract before AA-inducing UC in rats increased the proportion of the butyrate-producing bacteria (Lachnospiraceae NK4A136 group). The abundances of these beneficial bacteria and other SCFA-producing bacteria were unaffected by the indomethacin treatment with LP. Overall, our study revealed different impacts of AA-induced UC and ID-induced CD on changes in community composition and hinted at how LP may protect against UC by modifying the gut microbiota.




Similar content being viewed by others
Data availability
The raw sequence data generated during the current study are available in the NCBI Sequence Read Archive (SRA) repository under the BioProject accession number PRJNA785748 (BioSample accession numbers SAMN23575674–SAMN23575697) (https://www.ncbi.nlm.nih. gov/biopro ject/PRJNA785748).
Abbreviations
- CD:
-
Crohn’s disease
- IBD:
-
Inflammatory bowel disease
- LP:
-
Luem Pua rice extract
- UC:
-
Ulcerative colitis
- AA-induce UC:
-
Acetic acid–induced ulcerative colitis
- D0C:
-
Control group on day 0 of acetic acid–induced ulcerative colitis
- D7C:
-
Control group on day 7 of acetic acid–induced ulcerative colitis
- D7AA:
-
Acute acetic acid–induced ulcerative colitis
- D7AALP:
-
Acute acetic acid–induced ulcerative colitis pretreated with Luem Pua rice extract
- ID-induce CD:
-
Indomethacin-induced Crohn’s disease
- D0DW:
-
Control group on day 0 of chronic indomethacin-induced Crohn’s disease
- D11DW:
-
Control group on day 11 of chronic indomethacin-induced Crohn’s disease
- D11ID:
-
Chronic indomethacin-induced Crohn’s disease
- D11IDLP:
-
Chronic indomethacin-induced Crohn’s disease pretreated with Luem Pua rice extract
References
Anderson MJ (2006) Distance-based tests for homogeneity of multivariate dispersions. Biom 62:245–253. https://doi.org/10.1111/j.1541-0420.2005.00440.x
Anderson MJ, Ellingsen KE, McArdle BH (2006) Multivariate dispersion as a measure of beta diversity. Ecol Lett 9:683–693. https://doi.org/10.1111/j.1461-0248.2006.00926.x
Bokulich NA, Subramanian S, Faith JJ et al (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59. https://doi.org/10.1038/nmeth.2276
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Caporaso JG, Lauber CL, Walters WA et al (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Dey YN, Sharma G, Wanjari MM et al (2017) Beneficial effect of Amorphophallus paeoniifolius tuber on experimental ulcerative colitis in rats. Pharm Biol 55:53–62. https://doi.org/10.1080/13880209.2016.1226904
Edgar RC, Haas BJ, Clemente JC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinforma 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
El-Akabawy G, El-Sherif NM (2019) Zeaxanthin exerts protective effects on acetic acid-induced colitis in rats via modulation of pro-inflammatory cytokines and oxidative stress. Biomed Pharmacother 111:841–851. https://doi.org/10.1016/j.biopha.2019.01.001
Fox J, Weisberg S (2019) An R companion to applied regression, Third edit. Sage, Thousand Oaks CA. Available online: https://socialsciences.mcmaster.ca/jfox/Books/Companion/. Accessed 8 Nov 2021
Glassner KL, Abraham BP, Quigley EMM (2020) The microbiome and inflammatory bowel disease. J Allergy Clin Immunol 145:16–27. https://doi.org/10.1016/j.jaci.2019.11.003
Guan Q (2019) A comprehensive review and update on the pathogenesis of inflammatory bowel disease. J Immunol Res 2019:1–16. https://doi.org/10.1155/2019/7247238
Guo X, Huang C, Xu J, Xu H, Liu L, Zhao H, Wang J, Huang W, Peng W, Chen Y, Nie Y, Zhou Y, Zhou Y (2022) Gut Microbiota is a potential biomarker in inflammatory bowel disease. Front Nutr 8. https://doi.org/10.3389/fnut.2021.818902
Han Y, Song M, Gu M et al (2019) Dietary intake of whole strawberry inhibited colonic inflammation in dextran-sulfate-sodium-treated mice via restoring immune homeostasis and alleviating gut microbiota dysbiosis. J Agric Food Chem 67:9168–9177. https://doi.org/10.1021/acs.jafc.8b05581
Hu S, Wang J, Xu Y et al (2019) Anti-inflammation effects of fucosylated chondroitin sulphate from Acaudina molpadioides by altering gut microbiota in obese mice. Food Funct 10:1736–1746. https://doi.org/10.1039/C8FO02364F
Huang K, Dong W, Liu W et al (2019) 2- o-β- d -Glucopyranosyl- l -ascorbic acid, an ascorbic acid derivative isolated from the fruits of Lycium barbarum L., modulates gut microbiota and palliates colitis in dextran sodium sulfate-induced colitis in mice. J Agric Food Chem 67:11408–11419. https://doi.org/10.1021/acs.jafc.9b04411
Huangfu L, Cai X, Yang J et al (2021) Irisin attenuates inflammation in a mouse model of ulcerative colitis by altering the intestinal microbiota. Exp Ther Med 22:1433. https://doi.org/10.3892/etm.2021.10868
Khan I, Ullah N, Zha L et al (2019) Alteration of gut microbiota in inflammatory bowel disease (IBD): cause or consequence? IBD treatment targeting the gut microbiome. Pathog 8:126
Khor B, Gardet A, Xavier RJ (2011) Genetics and pathogenesis of inflammatory bowel disease. Nat 474:307–317. https://doi.org/10.1038/nature10209
Klindworth A, Pruesse E, Schweer T et al (2013) Evaluation of general 16s ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1–e1. https://doi.org/10.1093/nar/gks808
Lee WT, Tung YT, Wu CC et al (2018) Camellia oil (Camellia oleifera Abel.) modifies the composition of gut microbiota and alleviates acetic acid-induced colitis in rats. J Agric Food Chem 66:7384–7392. https://doi.org/10.1021/acs.jafc.8b02166
Li M, Liu Y, Cui J et al (2019) Ulcerative colitis with mucosal lesions in duodenum. Med (Baltimore) 98:e15035. https://doi.org/10.1097/MD.0000000000015035
Mak WY, Zhao M, Ng SC, Burisch J (2020) The epidemiology of inflammatory bowel disease: east meets west. J Gastroenterol Hepatol 35:380–389. https://doi.org/10.1111/jgh.14872
McMurdie PJ, Holmes S (2013) Phyloseq: an r package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8(4). https://doi.org/10.1371/journal.pone.0061217
Morgan XC, Tickle TL, Sokol H et al (2012) Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol 13:R79. https://doi.org/10.1186/gb-2012-13-9-r79
Ng SC, Shi HY, Hamidi N et al (2017) Worldwide incidence and prevalence of inflammatory bowel disease in the 21st century: a systematic review of population-based studies. Lancet 390:2769–2778. https://doi.org/10.1016/S0140-6736(17)32448-0
Oksanen J, Kindt R, Pierre L, O’Hara B, Simpson GL, Solymos P, Stevens MH. HH, Wagner H, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MH. HH, Wagner H (2016) Vegan: community ecology package, R package version 2.4-0. R package version 2.2-1. The Comprehensive R Archive Network. Available online: https://CRAN.R-project.org/package=vegan. Accessed 30 Oct 2021
Pawar A, Ghodasara J, Anap R, Kuchekar B (2011) Protective effect of hydroalcoholic root extract of Rubia cordifolia in indomethacin-induced enterocolitis in rats. Indian J Pharm Sci 73:250. https://doi.org/10.4103/0250-474X.91577
Pawłowska B, Prof Sobieszczańska B (2017) Intestinal epithelial barrier: the target for pathogenic Escherichia coli. Adv Clin Exp Med 26:1437–1445
Quast C, Pruesse E, Yilmaz P et al (2013) The Silva ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
R Core Team (2020) R: a language and environment for statistical computing. R foundation for statistical computing, Vienna, Austria
Schroeder BO (2019) Fight them or feed them: how the intestinal mucus layer manages the gut microbiota. Gastroenterol Rep 7:3–12. https://doi.org/10.1093/gastro/goy052
Segata N, Izard J, Waldron L et al (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. https://doi.org/10.1186/gb-2011-12-6-r60
Shang L, Liu H, Yu H et al (2021) Core altered microorganisms in colitis mouse model: a comprehensive time-point and fecal microbiota transplantation analysis. Antibiot 10:643. https://doi.org/10.3390/antibiotics10060643
Srisuwan S, Arkaravichien T, Mahatheeranont S et al (2015) Effects of aqueous extract of unpolished dark purple glutinous rice, Var Luem Pua, on ROS in SK-N-SH cells and scopolamine-induced memory deficit in mice. Trop J Pharm Res 14:1635–1641. https://doi.org/10.4314/tjpr.v14i9.13
Suwannalert P, Rattanachitthawat S (2011) High levels of phytophenolics and antioxidant activities in Oryza sativa - unpolished Thai rice strain of Leum Phua. Trop J Pharm Res 10:431–436. https://doi.org/10.4314/tjpr.v10i4.8
Terán-Ventura E, Aguilera M, Vergara P, Martínez V (2014) Specific changes of gut commensal microbiota and TLRS during indomethacin-induced acute intestinal inflammation in rats. J Crohn’s Colitis 8:1043–1054. https://doi.org/10.1016/j.crohns.2014.02.001
Thipart K, Phunikhom K, Noenplab ANL, Sattayasai J (2020) Dark purple Luem Pua rice extract reduces disease severity in rat models of inflammatory bowel disease. Trop J Pharm Res 19:629–636. https://doi.org/10.4314/tjpr.v19i3.25
Thipart K, Phunikhom K, Noenplab ANL, Sattayasai J (2021) Luem Pua rice extract attenuates the severity of dextran sulfate sodium-induced ulcerative colitis in rats through cholinomimetic effects. Int J Appl Pharm 13:5–8. https://doi.org/10.22159/ijap.2021.v13s1.Y0029
Thomas S, Svetlana L, Julie L et al (2017) Development of inflammatory bowel disease is linked to a longitudinal restructuring of the gut metagenome in mice. mSystems 2:e00036-17. https://doi.org/10.1128/mSystems.00036-17
Venegas DP, De La Fuente MK, Landskron G, González MJ, Quera R, Dijkstra G, Harmsen HJM, Faber KN, Hermoso MA (2019) Short chain fatty acids (SCFAs) mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Front Immunol 10(MAR). https://doi.org/10.3389/fimmu.2019.00277
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer-Verlag, New York
Xu H-M, Huang H-L, Liu Y-D et al (2021) Selection strategy of dextran sulfate sodium-induced acute or chronic colitis mouse models based on gut microbial profile. BMC Microbiol 21:279. https://doi.org/10.1186/s12866-021-02342-8
Yeom Y, Kim B-S, Kim S-J, Kim Y (2016) Sasa quelpaertensis leaf extract regulates microbial dysbiosis by modulating the composition and diversity of the microbiota in dextran sulfate sodium-induced colitis mice. BMC Complement Altern Med 16:481. https://doi.org/10.1186/s12906-016-1456-7
Zou S, Fang L, Lee M-H (2018) Dysbiosis of gut microbiota in promoting the development of colorectal cancer. Gastroenterol Rep 6(1). https://doi.org/10.1093/gastro/gox031
Zuo T, Ng SC (2018) The gut microbiota in the pathogenesis and therapeutics of inflammatory bowel disease. Front Microbiol 9. https://doi.org/10.3389/fmicb.2018.02247
Acknowledgements
We thank Mae Fah Luang University for supporting the Gut Microbiome Research Group.
Funding
This work was funded by Mae Fah Luang University.
Author information
Authors and Affiliations
Contributions
Kornsuda Thipart, Kutcharin Phunikhom, Jintana Sattayasai, and Siam Popluechai conceived and designed the study. Kornsuda Thipart performed the experiments. Lucsame Gruneck analyzed the data and prepared the figures. Kornsuda Thipart, Lucsame Gruneck, and Siam Popluechai wrote the original draft. Kutcharin Phunikhom, Thomas J. Sharpton, Jintana Sattayasai, Lucsame Gruneck, and Siam Popluechai reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
All procedures were complied with the standards for the care and use of experimental animals and approved by the Animal Ethics Committee of Khon Kaen University according to the Ethics of Animal Experimentation of National Research Council of Thailand (Ethics Registry: ACUC-KKU-21/2560). Animal use was constrained to abide with the Buddhist moral of the country.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thipart, K., Gruneck, L., Phunikhom, K. et al. Dark-purple rice extract modulates gut microbiota composition in acetic acid– and indomethacin-induced inflammatory bowel disease in rats. Int Microbiol 26, 423–434 (2023). https://doi.org/10.1007/s10123-022-00309-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-022-00309-x