Abstract
Substance P is a neurotransmitter or modulator in both the central and peripheral nervous systems. In this work, modifications of the lysine in SP by homocysteine and an acetyl group as well as the conformational dynamics of the native and modified SP peptides and their complexes with the NK1 receptor were studied via MD simulation. It was found that modifying SP stabilizes the peptide structure, but the modified SP peptides are less likely to bind to the NK1 receptor, so the resulting complexes are less stable. The RMSD of native SP (~0.33 nm) is about twice as large as that of the modified SP peptides (~0.18 nm), while the RMSD for the receptor complexed with native SP is ~0.3 nm, and that for the receptor complexed with either of the modified peptides is ~0.35 nm, which demonstrates the high stability of the modified SP peptides as well as the receptor complexed with native SP. Such behavior was also observed in other structural analyses. The binding free energies of the native and modified SP peptides with the NK1 receptor were also compared. The ΔGbind values for the binding of homocysteinylated SP to the NK1 receptor and the binding of the acetylated SP and native SP to the NK1 receptor were −38.89, −64.46, and − 264.52 kJ mol−1, respectively. Modification of the lysine of SP decreases the binding affinity of the peptide to the NK1 receptor. In other words, homocysteinylation or acetylation of SP leads to weaker interactions of the peptide with the NK1 receptor compared to those between native SP and NK1. We propose that this phenomenon leads to increased levels of homocysteinylated SP in plasma in many diseases such as breast cancer.

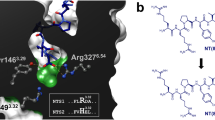
Substance P (SP) is a neuropeptide which binds to the NK1 receptor. SP is of great pharmacological interest, as agonists and antagonists of SP can potentially be used to treat many chronic diseases. Therefore, in this work, the lysine (LYS) in SP was theoretically modified with a homocysteine or acetyl group to explore the effects of such a modification on the binding affinity of this peptide with the NK1 receptor and the structural dynamics of the resulting complex










Similar content being viewed by others
Abbreviations
- SP:
-
Substance P
- SP1:
-
Substance P (Lys-homo)
- SP2:
-
Substance P (Lys-acetyl)
- NK1:
-
Neurokinin 1
- RMSD:
-
Root mean square deviation
- RMSF:
-
Root mean square fluctuation
- R g :
-
Radius of gyration
- DSSP:
-
Secondary structure analysis
- EGFR:
-
Epidermal growth factor receptor
- MD:
-
Molecular dynamics simulation
- CD:
-
Circular dichroism
- NMR:
-
Nuclear magnetic resonance
- SDS:
-
Sodium dodecyl sulfate
- MM/PBSA:
-
Molecular mechanics/Poisson–Boltzmann surface area
- PDB:
-
Protein Data Bank
- DOPC:
-
1,2-Dioleoyl-sn-glycero-3-phosphocholine
- VMD:
-
Visual molecular dynamics
- H-bond:
-
Hydrogen bond
- CBS:
-
Cystathionine beta-synthase
- MTHFR:
-
Methylene tetrahydrofolate reductase
References
Hökfelt T, Pernow B, Wahren J (2001) Substance P: a pioneer amongst neuropeptides. J Intern Med 249(1):27–40. https://doi.org/10.1046/j.0954-6820.2000.00773.x
Reiser G, Hamprecht B (1988) Characterization of a substance P receptor activating a cation permeability in neuronal cell lines. Eur J Pharmacol 145(3):273–280. https://doi.org/10.1016/0014-2999(88)90430-X
Kohara H, Tajima S, Yamamoto M, Tabata Y (2010) Angiogenesis induced by controlled release of neuropeptide substance P. Biomaterials 31(33):8617–8625. https://doi.org/10.1016/j.biomaterials.2010.07.079
Rizo J, Gierasch LM (1992) Constrained peptides: models of bioactive peptides and protein substructures. Annu Rev Biochem 61(1):387–416. https://doi.org/10.1146/annurev.bi.61.070192.002131
Jensen J (1989) Substance P and other tachykinins. In: Holmgren S (ed) The comparative physiology of regulatory peptides. Springer, Berlin, p 130–149. https://doi.org/10.1007/978-94-009-0835-2_6
Pennefather JN et al (2004) Tachykinins and tachykinin receptors: a growing family. Life Sci 74(12):1445–1463. https://doi.org/10.1016/j.lfs.2003.09.039
Steinhoff MS, von Mentzer B, Geppetti P, Pothoulakis C, Bunnett NW (2014) Tachykinins and their receptors: contributions to physiological control and the mechanisms of disease. Physiol Rev 94(1):265–301. https://doi.org/10.1152/physrev.00031.2013
Garcia-Recio S, Gascón P (2015) Biological and pharmacological aspects of the NK1-receptor. Biomed Res Int 2015:495704. https://doi.org/10.1155/2015/495704
Tansky MF, Pothoulakis C, Leeman SE (2007) Functional consequences of alteration of N-linked glycosylation sites on the neurokinin 1 receptor. Proceedings of the National Academy of Sciences of the United States of America. 104 (25):10691–10696. https://doi.org/10.1073/pnas.0703394104
Mantyh PW (2001) Neurobiology of substance P and the NK1 receptor. J Clin Psychiatry 63:6–10 http://www.psychiatrist.com/jcp/article/pages/2002/v63s11/v63s1102
Otsuka M, Yoshioka K (1993) Neurotransmitter functions of mammalian tachykinins. Physiol Rev 73(2):229–308. https://doi.org/10.1152/physrev.1993.73.2.229
Krause JE, Takeda Y, Hershey AD (1992) Structure, functions, and mechanisms of substance P receptor action. J Investig Dermatol 98(6):S2–S7. https://doi.org/10.1111/1523-1747.ep12462082
Satake H, Kawada T (2006) Overview of the primary structure, tissue-distribution, and functions of tachykinins and their receptors. Curr Drug Targets 7(8):963–974
Blier P et al (2004) Impact of substance P receptor antagonism on the serotonin and norepinephrine systems: relevance to the antidepressant/anxiolytic response. J Psychiatry Neurosci 29(3):208
Arora P, Cuevas B, Russo A, Johnson GL, Trejo J (2008) Persistent transactivation of EGFR and ErbB2/HER2 by protease-activated receptor-1 promotes breast carcinoma cell invasion. Oncogene 27(32):4434–4445. https://doi.org/10.1038/onc.2008.84
Mayordomo C et al (2012) Targeting of substance P induces cancer cell death and decreases the steady state of EGFR and Her2. J Cell Physiol 227(4):1358–1366. https://doi.org/10.1002/jcp.22848
Garcia-Recio S, Pastor-Arroyo EM, Marín-Aguilera M, Almendro V, Gascón P (2015) The transmodulation of HER2 and EGFR by substance P in breast cancer cells requires c-Src and metalloproteinase activation. PLoS One 10(6):e0129661. https://doi.org/10.1371/journal.pone.0129661
Dutta AS et al (1986) Analogues of substance P. Peptides containing D-amino acid residues in various positions of substance P and displaying agonist or receptor selective antagonist effects. J Med Chem 29(7):1163–1171. https://doi.org/10.1021/jm00157a008
Engberg G, Svensson TH, Rosell S, Folkers K (1981) A synthetic peptide as an antagonist of substance P. Nature 293(5829):222–223. https://doi.org/10.1038/293222a0
Carrey E (1990) Peptide mapping. In: Creighton TE (ed) Protein function: a practical approach. Oxford University Press, Oxford
Khodadadi S et al (2012) Effect of N-homocysteinylation on physicochemical and cytotoxic properties of amyloid β-peptide. FEBS Lett 586(2):127–131. https://doi.org/10.1016/j.febslet.2011.12.018
Feeney RE (1987) Chemical modification of proteins: comments and perspectives. Chem Biol Drug Des 29(2):145–161. https://doi.org/10.1111/j.1399-3011.1987.tb02241.x
Cascieri M et al (1986) Conformationally constrained tachykinin analogs which are selective ligands for the eledoisin binding site. Mol Pharmacol 29(1):34–38
Karima O et al (2012) Altered tubulin assembly dynamics with N-homocysteinylated human 4R/1N tau in vitro. FEBS Lett 586(21):3914–3919. https://doi.org/10.1016/j.febslet.2012.09.024
Teleman O, Von Der Lieth CW (1990) Molecular dynamics simulation provides a possible structure for substance P-like peptides in aqueous solution. Biopolymers 30(1–2):13–23. https://doi.org/10.1002/bip.360300104
Corcho FJ, Canto J, Perez JJ (2004) Comparative analysis of the conformational profile of substance P using simulated annealing and molecular dynamics. J Comput Chem 25(16):1937–1952. https://doi.org/10.1002/jcc.20114
Prabhu A, Malde A, Coutinho E, Srivastava S (2005) Solution conformation of substance P antagonists—[D-Arg1, D-Trp7,9, Leu11]-SP, [D-Arg1, D-Pro2, D-Trp7,9, Leu11]-SP and [D-Pro2, D-Trp7,9]-SP by CD, NMR and MD simulations. Peptides 26(5):875–885. https://doi.org/10.1016/j.peptides.2004.12.001
Wymore T, Wong TC (1999) Molecular dynamics study of substance P peptides partitioned in a sodium dodecylsulfate micelle. Biophys J 76(3):1213–1227. https://doi.org/10.1016/S0006-3495(99)77285-1
Gayen A, Goswami SK, Mukhopadhyay C (2011) NMR evidence of GM1-induced conformational change of substance P using isotropic bicelles. Biochim Biophys Acta Biomembr 1808(1):127–139. https://doi.org/10.1016/j.bbamem.2010.09.023
Hypercube Inc. (2002) Hyperchem 7.0. Hypercube Inc., Gainesville
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461. https://doi.org/10.1002/jcc.21334
Lee J et al (2016) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12(1):405. https://doi.org/10.1021/acs.jctc.5b00935
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Morris GM et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791. https://doi.org/10.1002/jcc.21256
Weiner SJ et al (1984) A new force field for molecular mechanical simulation of nucleic acids and proteins. J Am Chem Soc 106(3):765–784. https://doi.org/10.1021/ja00315a051
Abraham M, van der Spoel D, Lindahl E, Hess B, The GROMACS Development Team (2016) GROMACS user manual, version 5.1.2. University of Groningen, Groningen
Duan Y et al (2003) A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J Comput Chem 24(16):1999–2012. https://doi.org/10.1002/jcc.10349
Case D et al (2005) AMBER 10. University of California, San Francisco
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935. https://doi.org/10.1063/1.445869
Aaqvist J (1990) Ion-water interaction potentials derived from free energy perturbation simulations. J Phys Chem 94(21):8021–8024. https://doi.org/10.1021/j100384a009
Nosé S (1984) A unified formulation of the constant temperature molecular dynamics methods. J Chem Phys 81(1):511–519. https://doi.org/10.1063/1.447334
Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys Rev A 31(3):1695. https://doi.org/10.1103/PhysRevA.31.1695
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52(12):7182–7190. https://doi.org/10.1063/1.328693
Cheatham TI, Miller J, Fox T, Darden T, Kollman P (1995) Molecular dynamics simulations on solvated biomolecular systems: the particle mesh Ewald method leads to stable trajectories of DNA, RNA, and proteins. J Am Chem Soc 117(14):4193–4194. https://doi.org/10.1021/ja00119a045
Ryckaert J-P, Ciccotti G, Berendsen HJ (1977) Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341. https://doi.org/10.1016/0021-9991(77)90098-5
Massova I, Kollman PA (1999) Computational alanine scanning to probe protein–protein interactions: a novel approach to evaluate binding free energies. J Am Chem Soc 121(36):8133–8143. https://doi.org/10.1021/ja990935j
Rastelli G, Rio AD, Degliesposti G, Sgobba M (2010) Fast and accurate predictions of binding free energies using MM-PBSA and MM-GBSA. J Comput Chem 31(4):797–810. https://doi.org/10.1002/jcc.21372
Wang J, Hou T, Xu X (2006) Recent advances in free energy calculations with a combination of molecular mechanics and continuum models. Curr Comput Aided Drug Des 2(3):287–306. https://doi.org/10.2174/157340906778226454
Gohlke H, Kiel C, Case DA (2003) Insights into protein–protein binding by binding free energy calculation and free energy decomposition for the Ras–Raf and Ras–RalGDS complexes. J Mol Biol 330(4):891–913. https://doi.org/10.1016/S0022-2836(03)00610-7
Gohlke H, Case DA (2004) Converging free energy estimates: MM-PB (GB) SA studies on the protein–protein complex Ras–Raf. J Comput Chem 25(2):238–250. https://doi.org/10.1002/jcc.10379
Tan C, Tan Y-H, Luo R (2007) Implicit nonpolar solvent models. J Phys Chem B 111(42):12263–12274. https://doi.org/10.1021/jp073399n
Sitkoff D, Sharp KA, Honig B (1994) Accurate calculation of hydration free energies using macroscopic solvent models. J Phys Chem 98(7):1978–1988. https://doi.org/10.1021/j100058a043
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22(12):2577–2637. https://doi.org/10.1002/bip.360221211
Jakubowski H (1997) Metabolism of homocysteine thiolactone in human cell cultures possible mechanism for pathological consequences of elevated homocysteine levels. J Biol Chem 272(3):1935–1942
Jakubowski H, Zhang L, Bardeguez A, Aviv A (2000) Homocysteine thiolactone and protein homocysteinylation in human endothelial cells. Circ Res 87(1):45–51. https://doi.org/10.1161/01.RES.87.1.45
Jakubowski H (1999) Protein homocysteinylation: possible mechanism underlying pathological consequences of elevated homocysteine levels. FASEB J 13(15):2277–2283. https://doi.org/10.1096/fasebj.13.15.2277
Jakubowski H (2000) Homocysteine thiolactone: metabolic origin and protein homocysteinylation in humans. J Nutr 130(2):377S–381S. https://doi.org/10.1093/jn/130.2.377S
Jakubowski H (2000) Calcium-dependent human serum homocysteine thiolactone hydrolase. A protective mechanism against protein N-homocysteinylation. J Biol Chem 275(6):3957–3962
Refsum H, Ueland PM, Nygård O, Vollset SE (1998) Homocysteine and cardiovascular disease. Annu Rev Med 49(1):31–62. https://doi.org/10.1146/annurev.med.49.1.31
Medina MÁ, Urdiales JL, Amores-Sánchez MI (2001) Roles of homocysteine in cell metabolism. Eur J Biochem 268(14):3871–3882. https://doi.org/10.1046/j.1432-1327.2001.02278.x
Polevoda B, Sherman F (2003) N-terminal acetyltransferases and sequence requirements for N-terminal acetylation of eukaryotic proteins. J Mol Biol 325(4):595–622. https://doi.org/10.1016/S0022-2836(02)01269-X
Hollebeke J, Van Damme P, Gevaert K (2012) N-terminal acetylation and other functions of Nα-acetyltransferases. Biol Chem 393(4):291–298. https://doi.org/10.1515/hsz-2011-0228
Hershko A, Heller H, Eytan E, Kaklij G, Rose IA (1984) Role of the alpha-amino group of protein in ubiquitin-mediated protein breakdown. Proc Natl Acad Sci 81(22):7021–7025. https://doi.org/10.1073/pnas.81.22.7021
Behnia R, Panic B, Whyte JR, Munro S (2004) Targeting of the Arf-like GTPase Arl3p to the Golgi requires N-terminal acetylation and the membrane protein Sys1p. Nat Cell Biol 6(5):405–413. https://doi.org/10.1038/ncb1120
Acknowledgements
The authors appreciate the financial support provided by the Research Council of the Malek Ashtar University of Technology, Tehran, Iran for the present study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(TXT 288 bytes)
Rights and permissions
About this article
Cite this article
Davoudmanesh, S., Mosaabadi, J.M. Investigation of the effect of homocysteinylation of substance P on its binding to the NK1 receptor using molecular dynamics simulation. J Mol Model 24, 177 (2018). https://doi.org/10.1007/s00894-018-3695-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3695-7