Abstract
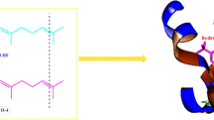
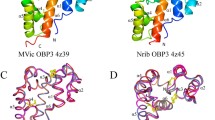
Odorant-binding proteins (OBPs) play an important role as ligand-transfer filters in olfactory recognition in insects. (E)-β-farnesene (EBF) is the main component of the aphid alarm pheromone and could keep aphids away from crops to prevent damage. Computational insight into the molecular binding mode of EBF analogs containing a heterocycle based on the structure of Megoura viciae OBP 3 (MvicOBP3) was obtained by molecular docking and molecular dynamics simulations. The results showed that high affinity EBF analogs substituted with an aromatic ring present a unique binding conformation in the surface cavity of MvicOBP3. A long EBF chain was located inside the cavity and was surrounded by many hydrophobic residues, while the substituted aromatic ring was exposed to the outside due to limitations from the formation of multiple hydrogen bonds. However, the low activity EBF analogs displayed an exactly inverted binding pose, with EBF loaded on the external side of the protein cavity. The affinity of the recently synthesized EBF analogs containing a triazine ring was evaluated in silico based on the binding modes described above and in vitro through fluorescence competitive binding assay reported later. Compound N1 not only showed a similar binding conformation to that of the high affinity analogs but was also found to have a much higher docking score and binding affinity than the other analogs. In addition, the docking score results correlated well with the predicted logP values for these EBF analogs, suggesting highly hydrophobic interactions between the protein and ligand. These studies provide an in silico screening model for the binding affinity of EBF analogs in order to guide their rational design based on aphid OBPs.





Similar content being viewed by others
References
Leal WS (2013) Odorant reception in insects: roles of receptors, binding proteins, and degrading enzymes. Annu Rev Entomol 58:373–391
Crasto CJ (2009) Computational biology of olfactory receptors. Curr Bioinform 4:8–15
Harini K, Kannan S, Nemoto W, Fukui K, Sowdhaminil R (2012) Residue conservation and dimer-interface analysis of olfactory receptor molecular models. J Mol Biochem 1:161–170
Harini K, Sowdhamini R (2012) Molecular modelling of oligomeric states of DmOR83b, an olfactory receptor in D. melanogaster. Bioinform Biol Insights 6:33–47
Sun YF, Qiao HL, Ling Y, Yang SX, Rui CH, Pelosi P, Yang XL (2011) New analogues of (E)-β-farnesene with insecticidal activity and binding affinity to aphid odorant-binding proteins. J Agric Food Chem 59:2456–2461
Wang SS, Sun YF, Du SQ, Qin YG, Duan HX, Yang XL (2016) Computer-aided rational design of novel EBF analogues with an aromatic ring. J Mol Model 22(144):1–9
Zhong T, Yin J, Deng SS, Li KB, Cao YZ (2012) Fluorescence competition assay for the assessment of green leaf volatiles and trans-β-farnesene bound to three odorant-binding proteins in the wheat aphid Sitobion avenae (Fabricius). J Insect Physiol 58:771–781
Qiao HL, Tuccori E, He XL, Gazzano A, Field L, Zhou JJ, Pelosi P (2009) Discrimination of alarm pheromone (E)-β-farnesene by aphid odorant-binding proteins. Insect Biochem Mol Biol 39:414–419
Mao Y, Xu XZ, Xu W, Ishida Y, Leal WS, Ames JB, Clardy J (2010) Crystal and solution structures of an odorant-binding protein from the southern house mosquito complexed with an oviposition pheromone. Proc Natl Acad Sci USA 107:19102–19107
Leite NR, Krogh R, Xu W, Ishida Y, Iulek J, Leal WS, Oliva G (2009) Structure of an odorant-binding protein from the mosquito Aedes aegypti suggests a binding pocket covered by a pH-sensitive “lid”. PLoS One 4:e8006
Lagarde A, Spinelli S, Tegoni M, He XL, Field L, Zhou JJ, Cambillau C (2011) The crystal structure of odorant binding protein 7 from Anopheles gambiae exhibits an outstanding adaptability of its binding site. J Mol Biol 414:401–412
Spinelli S, Lagarde A, Iovinella I, Legrand P, Tegoni M, Pelosi P, Cambillau C (2012) Crystal structure of Apis mellifera OBP14, a C-minus odorant-binding protein, and its complexes with odorant molecules. Insect Biochem Mol Biol 42:41–50
Zheng JG, Li JR, Han L, Wang Y, Wu W, Qi XX, Tao Y, Zhang L, Zhang ZD, Chen ZZ (2015) Crystal structure of the Locusta migratoria odorant binding protein. Biochem Biophys Res Commun 456:737–742
Northey T, Venthur H, De Biasio F, Chauviac FX, Cole A, Ribeiro Junior KAL, Grossi G, Falabella P, Field LM, Keep NH, Zhou JJ (2016) Crystal structures and binding dynamics of odorant-binding protein 3 from two aphid species Megoura viciae and Nasonovia ribisnigri. Sci Rep 6:24739. https://doi.org/10.1038/srep24739
Vanderveken JJ (1977) Oils and other inhibitors of nonpersistent virus transmission. In: Harris KF, Maramorosch K (eds) Aphids as virus vectors. Academic, New York pp 435–454
Edwards LJ, Siddall JB, Dunham LL, Uden P, Kislow CJ (1973) Trans-β-farnesene, alarm pheromone of the green peach aphid, Myzus persicae (Sulzer). Nature 241:126–127
Francis F, Vandermoten S, Verheggen F, Lognay G, Haubruge E (2005) Is the (E)-β-farnesene only volatile terpenoid in aphids? J Appl Entomol 129(1):6–11
Bowers WS, Nishino C, Montgomery ME, Nault LR, Nielson MW (1977) Sesquiterpene progenitor, germacrene a: an alarm pheromone in aphids. Science 196(4290):680–681
Gibson R, Pickett J (1983) Wild potato repels aphids by release of aphid alarm pheromone. Nature 302:608–609
Mondor EB, Baird DS, Slessor K, Roitberg BD (2000) Ontogeny of alarm pheromone secretion in pea aphid, Acyrthosiphon pisum. J Chem Ecol 26(12):2875–2882
Gut J, van Oosten AM, Harrewijn P, van Rheenen B (1988) Additional functions of alarm pheromones in development processes of aphids. Agric Ecosyst Environ 21:125–127
Cui LL, Dong J, Francis F, Liu YJ, Heuskin S, Lognay G, Chen JL, Bragard C, Tooker JF, Liu Y (2012) E-β-farnesene synergizes the influence of an insecticide to improve control of cabbage aphids in China. Crop Prot 35:91–96
Francis F, Lognay G, Haubruge E (2004) Olfactory responses to aphid and host plant volatile releases: (E)-β-farnesene an effective kairomone for the predator Adalia bipunctata. J Chem Ecol 30(4):741–755
Francis F, Martin T, Lognay G, Eric H (2005) Role of (E)-beta-farnesene in systematic aphid prey location by Episyrphus balteatus larvae (Diptera: Syrphidae). Eur J Entomol 102(3):431–436
Foster S, Denholm I, Thompson R, Poppy GM, Powell W (2005) Reduced response of insecticide-resistant aphids and attraction of parasitoids to aphid alarm pheromone; a potential fitness trade-off. B Entomol Res 95(1):37–46
Micha SG, Wyss U (1996) Aphid alarm pheromone (E)-β-farnesene: a host finding kairomone for the aphid primary parasitoid Aphidius uzbekistanicus (hymenoptera: Aphidiinae). Chemoecology 7(3):132–139
Dawson GW, Giffiths DC, Pickett JA, Plumb RT, Woodcock CM, Zhang ZN (1988) Structure-activity studies on aphid alarm pheromone derivatives and their field use against transmission of barley yellow dwarf virus. Pestic Sci 22:17–30
Li ZM, Wang TS, Yao EY (1987) Researches on insect pheromone III studies on aphid alarm pheromone mimics. Acta Chem Sin 45:1124–1128
Zhang ZN, Liu X, Pickett JA (1988) Several aphids alarm pheromone analogues possessing biological activity. Acta Entomol Sin 31:435–438
Sun L, Ling Y, Wang C, Sun YF, Rui CH, Yang XL (2011) Synthesis and biological activity of E-β-farnesene analogues containing substituent nitroguanidine. Chin J Org Chem 31:2061–2066
Kang TN, Ling Y, Rui CH, Yang XL, Fan XL, Chen FH (2008) Synthesis of E-β-farnesene analogues containing five-membered azaheterocycles and their biological activity. Chin J Org Chem 28:617–621
Sun YF, Li YQ, Ling Y, Yu HL, Yang SX, Yang XL (2011) Design, synthesis and biological activity of E-β-farnesene analogues containing pyrazole-carboxamide. Chin J Org Chem 31:1425–1432
Qin YG, Zhang JP, Song DL, Duan HX, Li WH, Yang XL. (2016) Novel (E)-β-Farnesene analogues containing 2-nitro iminohexahydro-1,3,5-triazine: synthesis and biological activity evaluation. Molecules 21(825):1–14
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JAJR, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2003) GAUSSIAN03, revision D.01. Gaussian, Inc., Wallingford, CT
Ruppert J, Welch W, Jain AN (1997) Automatic identification and representation of protein binding sites for molecular docking. Protein Sci 6:524–533
SYBYL Molecular Modeling Software, 7.3 (2006) Tripos, St. Louis, MO
Jain AN (2003) Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J Med Chem 46:499–511
Kellenberger E, Rodrigo J, Muller P, Rognan D (2004) Comparative evaluation of eight docking tools for docking and virtual screening accuracy. Proteins 57:225–242
Jain AN (1996) Scoring noncovalent protein-ligand interactions: a continuous differentiable function tuned to compute binding affinities. J Comput Aided Mol Des 10:427–440
Case DA, Babin V, Berryman JT, Betz RM, Cai Q, Cerutti DS, Cheatham III TE, Darden TA, Duke RE, Gohlke H, Goetz AW, Gusarov S, Homeyer N, Janowski P, Kaus J, Kolossváry I, Kovalenko A, Lee TS, LeGrand S, Luchko T, Luo R, Madej B, Merz KM, Paesani F, Roe DR, Roitberg A, Sagui C, Salomon-Ferrer R, Seabra G, Simmerling CL, Smith W, Swails J, Walker RC, Wang J, Wolf RM, Wu X, Kollman PA (2014) AMBER 14. University of California, San Francisco
Maier JA, Martinez C, Kasavajhala K, Wickstrom L, Hauser KE, Simmerling C (2015) ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J Chem Theory Comput 11:3696–3713
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 26(1):1157–1174
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N log (N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H et al (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Zhou JJ, Vieira FG, He XL, Smadja C, Liu R, Rozas J, Field LM (2010) Genome annotation and comparative analyses of the odorant-binding proteins and chemosensory proteins in the pea aphid Acyrthosiphon pisum. Insect Mol Biol 19(Suppl 2):113–122
Lautenschlager C, Leal WS, Clardy J (2007) Bombyx mori pheromone-binding protein binding nonpheromone ligands: implications for pheromone recognition. Structure 15:1148–1154
Tsitsanou KE, Thireou T, Drakou CE, Koussis K, Keramioti MV, Leonidas DD, Eliopoulos E, Iatrou K, Zographos SE (2012) Anopheles gambiae odorant binding protein crystal complex with the synthetic repellent DEET: implications for structure-based design of novel mosquito repellents. Cell Mol Life Sci 69:283–297
Acknowledgments
This study was supported by the National Key Research and Development Plan (2017YFD0200504) and the National Natural Science Foundation of China (NSFC) (No. 31772207, 31371946, 31272075) and State Key Laboratory Open Fund for Biology of Plant Diseases and Insect Pests (SKLOF201709).
Author information
Authors and Affiliations
Contributions
S.Q. Du, Z.K. Yang, S.S. Wang and H.X. Duan conceived and designed the experiments. Y.G. Qin synthesized the EBF compounds containing a triazine ring; S.Q. Du and H.X. Duan wrote the first draft of the manuscript. H.X. Duan and X.L. Yang made critical revisions and approved the final manuscript. All the authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Du, S., Yang, Z., Qin, Y. et al. Computational investigation of the molecular conformation-dependent binding mode of (E)-β-farnesene analogs with a heterocycle to aphid odorant-binding proteins. J Mol Model 24, 70 (2018). https://doi.org/10.1007/s00894-018-3612-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3612-0