Abstract
The experimentally-observed stable, electrically-conducting interface formed between hydrophobin protein HFBII and silicon provides a model system for the Bio/ICT interfaces required for bionanoelectronics. The present work used molecular dynamics (MD) computer simulations to investigate the atom-scale details of the assembly and structure of the HFBII/silicon interface, using models on the order of 40,000 atoms to compute energy profiles for the full protein interacting with a bare Si(111) substrate in aqueous solution. Five nanoseconds of free, equilibrated dynamics were performed for six models with initial protein:silicon separations ranging from 1.2 to 0.2 nanometers in steps of 0.2 nm. Three of the models formed extensive protein:silicon van der Waals’s interfacial contacts. The model with 0.2 nm starting separation serves as an illustrative example of the dynamic interface created, whereby hydrophobic patch residues cycle between flat and more protruding patch conformations that favor respectively close inter-patch and close patch-surface contacts, with protein:surface separations cycling between 0.2 and 0.4 nm over the 5 ns of dynamics. Analysis of residue-based binding energies at the interface reveal three leucines Leu19, Leu21 and Leu63, together with isoleucine Ile22 and alanine Ala61, as the primary drivers towards adhesion on bare silicon, providing the atom-scale details of HFBII’s hydrophobic patch which in turn provides leads for the engineering of more tightly-coupled interfaces.

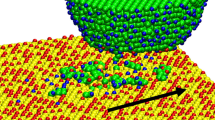
Atom-scale computer simulations reveal the structure, dynamics and energetics of hydrophobin protein HFBII immobilisation on bare silicon, a prototype for ordered organic/inorganic interfaces in future bionanoelectronics devices.





Similar content being viewed by others
References
Aggeli A, Bell M, Boden N, Keen JN, Knowles PF, McLeish TC, Pitkeathly M, Radford SE (1997) Nature 386:259–262
Colombo G, Soto P, Gazit E (2007) Trends Biotechnol 25:211–218
Whitesides GM, Lipomi DJ (2009) Faraday Discuss 143:373–384
Nosonovsky M, Bhushan B (2008) Adv Func Mater 18:843–855
Pum D, Neubauer A, Györvary E, Sára M, Sleytr UB (2000) Nanotechnol 11:100–107
Talbot NJ (1999) Nature 398:295–296
Wösten HA, van Wetter MA, Lugones LG, van der Mei HC, Busscher HJ, Wessels JG (1999) Curr Biol 9:85–88
Hakanpaa J, Paananen A, Askolin S, Nakari-Setala T, Parkkinen T, Penttila M, Linder MB, Rouvinen J (2004) J Biol Chem 279:534–539
Szilvay GR, Paananen A, Laurikainen K, Vuorimaa E, Lemmetyinen H, Peltonen J, Linder MB (2007) Biochemistry 46:2345–2354
Kivioja JM, Kurppa K, Kainlauri M, Linder MB, Ahopelto J (2009) Appl Phys Lett 94:183901
Zhao ZX, Qiao MQ, Yin F, Shao B, Wu BY, Wang YY, Wang XS, Qin X, Li S, Yu L, Chen Q (2007) Biosens Bioelectron 22:3021–3027
Laaksonen P, Kainlauri M, Laaksonen T, Shchepetov A, Jiang H, Ahopelto J, Linder M (2010) Angew Chem Int Ed 49:4946–4949
Lin Z, Strother T, Cai W, Cao X, Smith LM, Hamers RJ (2002) Langmuir 18:788–796
Streifer JA, Kim H, Nichols BM, Hamers RJ (2005) Nanotechnology 16:1868–1873
Nishiyama K, Hoshino T (2007) Appl Phys Lett 90:213901
Ziegler KJ (2005) Trends Biotechnol 23:440–428
Jorgensen W, Chandrasekhar J, Madura J, Impey R, Klein M (1983) J Chem Phys 79:926–935
MacKerell AD, Bashford D, Bellott M, Dunbrack RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S et al (1998) J Phys Chem B 102:3586–3616
Lopes PEM, Murashov V, Tazi M, Demchuk E, MacKerell AD Jr (2006) J Phys Chem B 110:2782–2792
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kale L, Schulten K (2005) J Comput Chem 26:1781–1802
Fan H, Wang X, Zhu J, Robillard GT, Mark AE (2006) Proteins 64:863–873
Kwan AH, Macindoe I, Vukasin PV, Morris VK, Kass I, Gupte R, Mark AE, Templeton MD, Mackay JP, Sunde M (2008) J Mol Biol 382:708–720
Humphrey W, Dalke A, Schulten K (1996) J Mol Graph 14:33–38
Iori F, Di Felice R, Molinari E, Corni S (2009) J Comput Chem 30:1465–1476
Love JC, Estroff LA, Kriebel JK, Nuzzo RG, Whitesides GM (2005) Chem Rev 105:1103–1170
Huskens J (2006) Curr Opin Chem Bio 10:537–543
Kivlehan F, Lefoix M, Moynihan HA, Thompson D, Ogurtsov VI, Herzog G, Arrigan DWM (2010) Electrochim Acta 55:3348–3354
Chan G, Mooney DJ (2008) Trends Biotechnol 26:382–392
Hearn EM, Patel DK, van der Berg B (2008) Proc Natl Acad Sci USA 105:8601–8606
Valo HK, Laaksonen PH, Peltonen LJ, Linder MB, Hirvonen JT, Laaksonen TJ (2010) ACS Nano 4:1750–1758
Zheng G, Patolsky F, Cui Y, Wang WU, Lieber CM (2005) Nature Biotechnol 23:1294–1301
Hiratsuka Y, Miyata M, Tada T, Uyeda TQP (2006) Proc Natl Acad Sci USA 103:13618–13623
Carter EA (2008) Science 321:800–803
Klein MK, Shinoda W (2008) Science 321:798–800
Ander M, Luzhkov VB, Aqvist J (2008) Biophys J 94:820–831
Meinhold L, Smith JC, Kitao A, Zewail AH (2007) Proc Nat Acad Sci 104:17261–17265
Thompson D (2007) Langmuir 23:8441–8451
Thompson D, Miller C, McCarthy FO (2008) Biochemistry 47:10333–10344
Gannon G, Greer JC, Larsson JA, Thompson D (2010) ACS Nano 4:921–932
Acknowledgments
We wish to acknowledge support for this research from Science Foundation Ireland (SFI) under the UREKA Programme and also Enterprise Ireland Innovation Partnership project ORD3D. We acknowledge SFI for computing resources at Tyndall National Institute and SFI/ Higher Education Authority for computing time at the Irish Centre for High-End Computing (ICHEC).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Supporting information
Internal hydrophobic pocket structures for each model to accompany the radial distribution function RDF plots in Fig. 2, residue-based components for hydrophobic patch immobilization on silicon to support the full patch binding energies and interface structures in Figs. 3 and 4, together with a movie showing the dynamics of HFBII/Si(111) interface formation to accompany Figs. 4 and 5.
ESM 1
(DOC 3.16 MB)
Supplementary Fig. S1
(GIF 51.7 MB)
Rights and permissions
About this article
Cite this article
Moldovan, C., Thompson, D. Molecular dynamics of the “hydrophobic patch” that immobilizes hydrophobin protein HFBII on silicon. J Mol Model 17, 2227–2235 (2011). https://doi.org/10.1007/s00894-010-0887-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-010-0887-1