Abstract
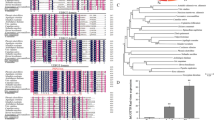
In plants, two types of methionine sulfoxide reductase (MSR) exist, namely methionine-S-sulfoxide reductase (MSRA) and methionine-R-sulfoxide reductase (MSRB). These enzymes catalyze the reduction of methionine sulfoxides (MetO) back to methionine (Met) by a catalytic cysteine (Cys) and one or two resolving Cys residues. Interestingly, a group of MSRA encoded by plant genomes does not have a catalytic residue. We asked that if this group of MSRA did not have any function (as fitness), why it was not lost during the evolutionary process. To challenge this question, we analyzed the gene family encoding MSRA in soybean (GmMSRAs). We found seven genes encoding GmMSRAs, which included three segmental duplicated pairs. Among them, a pair of duplicated genes, namely GmMSRA1 and GmMSRA6, was without a catalytic Cys residue. Pseudogenes were ruled out as their transcripts were detected in various tissues and their Ka/Ks ratio indicated a negative selection pressure. In vivo analysis in Δ3MSR yeast strain indicated that the GmMSRA6 did not have activity toward MetO, contrasting to GmMSRA3 which had catalytic Cys and had activity. When exposed to H2O2-induced oxidative stress, GmMSRA6 did not confer any protection to the Δ3MSR yeast strain. Overexpression of GmMSRA6 in Arabidopsis thaliana did not alter the plant’s phenotype under physiological conditions. However, the transgenic plants exhibited slightly higher sensitivity toward salinity-induced stress. Taken together, this data suggested that the plant MSRAs without the catalytic Cys are not enzymatically active and their existence may be explained by a role in regulating plant MSR activity via dominant-negative substrate competition mechanism.




Similar content being viewed by others
References
Achilli C, Ciana A, Minetti G (2015) The discovery of methionine sulfoxide reductase enzymes: an historical account and future perspectives. BioFactors 41:135–152. https://doi.org/10.1002/biof.1214
Belamkar V, Weeks NT, Bharti AK, Farmer AD, Graham MA, Cannon SB (2014) Comprehensive characterization and RNA-Seq profiling of the HD-zip transcription factor family in soybean (Glycine max) during dehydration and salt stress. BMC Genomics 15:1–25. https://doi.org/10.1186/1471-2164-15-950
Boschi-Muller S, Olry A, Antoine M, Branlant G (2005) The enzymology and biochemistry of methionine sulfoxide reductases. Biochim Biophys Acta 1703:231–238. https://doi.org/10.1016/j.bbapap.2004.09.016
Chatelain E, Satour P, Laugier E, Ly Vu B, Payet N, Rey P, Montrichard F (2013) Evidence for participation of the methionine sulfoxide reductase repair system in plant seed longevity. Proc Natl Acad Sci U S A 110:3633–3638. https://doi.org/10.1073/pnas.1220589110
Chu HD, Le QN, Nguyen HQ, Le DT (2016a) Genome-wide analysis of genes encoding methionine-rich proteins in Arabidopsis and soybean suggesting their roles in the adaptation of plants to abiotic stress. Int J Genomics 2016(8):1–8. https://doi.org/10.1155/2016/5427062
Chu HD, Nguyen K-L, Watanabe Y, Le DT, Tran LS (2016b) Expression analyses of soybean genes encoding methionine-R-sulfoxide reductase under various conditions suggest a possible role in the adaptation to stress. J Korean Soc Appl Biol Chem 59:681–687. https://doi.org/10.1007/s13765-016-0213-4
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Dai C, Wang MH (2012) Characterization and functional analysis of methionine sulfoxide reductase a gene family in tomato. Mol Biol Rep 39:6297–6308. https://doi.org/10.1007/s11033-012-1451-0
Emanuelsson O, Nielsen H, Brunak S, von Heijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300:1005–1016. https://doi.org/10.1006/jmbi.2000.3903
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186. https://doi.org/10.1093/nar/gkr944
Guo X, Wu Y, Wang Y, Chen Y, Chu C (2009) OsMSRA4.1 and OsMSRB1.1, two rice plastidial methionine sulfoxide reductases, are involved in abiotic stress responses. Planta 230:227–238. https://doi.org/10.1007/s00425-009-0934-2
Hansel A, Kuschel L, Hehl S, Lemke C, Agricola HJ, Hoshi T, Heinemann SH (2002) Mitochondrial targeting of the human peptide methionine sulfoxide reductase (MSRA), an enzyme involved in the repair of oxidized proteins. FASEB J 16:911–913. https://doi.org/10.1096/fj.01-0737fje
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinforma 2008(420747):1–5. https://doi.org/10.1155/2008/420747
Hurst LD (2002) The Ka/Ks ratio: diagnosing the form of sequence evolution. Trends Genet 18:486–487
Hussein KA, Joo JH (2015) Isolation and characterization of rhizomicrobial isolates for phosphate solubilization and indole acetic acid production. J Korean Soc Appl Biol Chem 58:847–855. https://doi.org/10.1007/s13765-015-0114-y
Jacques S, Ghesquiere B, De Bock PJ, Demol H, Wahni K, Willems P, Messens J, Van Breusegem F, Gevaert K (2015) Protein methionine sulfoxide dynamics in Arabidopsis thaliana under oxidative stress. Mol Cell Proteomics 14:1217–1229. https://doi.org/10.1074/mcp.M114.043729
Kwon EJ, Lee MK, Jeon H, Hwang JM, Kim J-K, Kim M (2016) Analysis of OsmiR399 expression and down-regulation of LTN1 by the multiple members of OsmiR399 family in rice. J Korean Soc Appl Biol Chem 59:623–630. https://doi.org/10.1007/s13765-016-0203-6
Le DT, Lee BC, Marino SM, Zhang Y, Fomenko DE, Kaya A, Hacioglu E, Kwak GH, Koc A, Kim HY, Gladyshev VN (2009) Functional analysis of free methionine-R-sulfoxide reductase from Saccharomyces cerevisiae. J Biol Chem 284:4354–4364. https://doi.org/10.1074/jbc.M805891200
Le DT, Nishiyama R, Watanabe Y, Tanaka M, Seki M, Ham LH, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2012a) Differential gene expression in soybean leaf tissues at late developmental stages under drought stress revealed by genome-wide transcriptome analysis. PLoS One 7:e49522. https://doi.org/10.1371/journal.pone.0049522
Le DT, Nishiyama R, Watanabe Y, Vankova R, Tanaka M, Seki M, Ham LH, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2012b) Identification and expression analysis of cytokinin metabolic genes in soybean under normal and drought conditions in relation to cytokinin levels. PLoS One 7:e42411. https://doi.org/10.1371/journal.pone.0042411
Le DT, Tarrago L, Watanabe Y, Kaya A, Lee BC, Tran UT, Nishiyama R, Fomenko DE, Gladyshev VN, Tran LS (2013) Diversity of plant methionine sulfoxide reductases B and evolution of a form specific for free methionine sulfoxide. PLoS One 8:e65637. https://doi.org/10.1371/journal.pone.0065637
Lee BC, Le DT, Gladyshev VN (2008) Mammals reduce methionine-S-sulfoxide with MsrA and are unable to reduce methionine-R-sulfoxide, and this function can be restored with a yeast reductase. J Biol Chem 283:28361–28369. https://doi.org/10.1074/jbc.M805059200
Lee Y, Krishnamoorthy R, Selvakumar G, Kim K, Sa T (2015) Alleviation of salt stress in maize plant by co-inoculation of arbuscular mycorrhizal fungi and Methylobacterium oryzae CBMB20. J Korean Soc Appl Biol Chem 58:533–540. https://doi.org/10.1007/s13765-015-0072-4
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. https://doi.org/10.1093/nar/30.1.325
Li CW, Lee SH, Chieh PS, Lin CS, Wang YC, Chan MT (2012) Arabidopsis root-abundant cytosolic methionine sulfoxide reductase B genes MsrB7 and MsrB8 are involved in tolerance to oxidative stress. Plant Cell Physiol 53:1707–1719. https://doi.org/10.1093/pcp/pcs114
Libault M, Farmer A, Joshi T, Takahashi K, Langley RJ, Franklin LD (2010) An integrated transcriptome atlas of the crop model Glycine max, and its use in comparative analyses in plants. Plant J 63:86–99. https://doi.org/10.1111/j.1365-313X.2010.04222.x
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Lin Z, Johnson LC, Weissbach H, Brot N, Lively MO, Lowther WT (2007) Free methionine-(R)-sulfoxide reductase from Escherichia coli reveals a new GAF domain function. Proc Natl Acad Sci U S A 104:9597–9602. https://doi.org/10.1073/pnas.0703774104
Moskovitz J, Flescher E, Berlett BS, Azare J, Poston JM, Stadtman ER (1998) Overexpression of peptide-methionine sulfoxide reductase in Saccharomyces cerevisiae and human T cells provides them with high resistance to oxidative stress. Proc Natl Acad Sci U S A 95:14071–14075
Mumberg D, Muller R, Funk M (1995) Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene 156:119–122. https://doi.org/10.1016/0378-1119(95)00037-7
Oh SK, Baek KH, Seong ES, Joung YH, Choi GJ, Park JM, Cho HS, Kim EA, Lee S, Choi D (2010) CaMsrB2, pepper methionine sulfoxide reductase B2, is a novel defense regulator against oxidative stress and pathogen attack. Plant Physiol 154:245–261. https://doi.org/10.1104/pp.110.162339
Roje S (2006) S-Adenosyl-L-methionine: beyond the universal methyl group donor. Phytochemistry 67:1686–1698. https://doi.org/10.1016/j.phytochem.2006.04.019
Romero HM, Berlett BS, Jensen PJ, Pell EJ, Tien M (2004) Investigations into the role of the plastidial peptide methionine sulfoxide reductase in response to oxidative stress in Arabidopsis. Plant Physiol 136:3784–3794. https://doi.org/10.1104/pp.104.046656
Rouhier N, Vieira Dos Santos C, Tarrago L, Rey P (2006) Plant methionine sulfoxide reductase a and B multigenic families. Photosynth Res 89:247–262. https://doi.org/10.1007/s11120-006-9097-1
Ruan H, Tang XD, Chen ML, Joiner ML, Sun G, Brot N, Weissbach H, Heinemann SH, Iverson L, Wu CF, Hoshi T (2002) High-quality life extension by the enzyme peptide methionine sulfoxide reductase. Proc Natl Acad Sci U S A 99:2748–2753. https://doi.org/10.1073/pnas.032671199
Sadanandom A, Poghosyan Z, Fairbairn DJ, Murphy DJ (2000) Differential regulation of plastidial and cytosolic isoforms of peptide methionine sulfoxide reductase in Arabidopsis. Plant Physiol 123:255–264. https://doi.org/10.1104/pp.123.1.255
Sanchez J, Nikolau BJ, Stumpf PK (1983) Reduction of N-acetyl methionine sulfoxide in plants. Plant Physiol 73:619–623
Siddikee MA, Sundaram S, Chandrasekaran M, Kim K, Selvakumar G, Sa T (2015) Halotolerant bacteria with ACC deaminase activity alleviate salt stress effect in canola seed germination. J Korean Soc Appl Biol Chem 58:237–241. https://doi.org/10.1007/s13765-015-0025-y
Snijder J, Rose RJ, Raijmakers R, Heck AJ (2011) Site-specific methionine oxidation in calmodulin affects structural integrity and interaction with Ca2+/calmodulin-dependent protein kinase II. J Struct Biol 174:187–195. https://doi.org/10.1016/j.jsb.2010.12.002
Stadtman ER (1993) Oxidation of free amino acids and amino acid residues in proteins by radiolysis and by metal-catalyzed reactions. Annu Rev Biochem 62:797–821. https://doi.org/10.1146/annurev.bi.62.070193.004053
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tang H, Bowers JE, Wang X, Ming R, Alam M, Paterson AH (2008) Synteny and collinearity in plant genomes. Science 320:486–488. https://doi.org/10.1126/science.1153917
Tarrago L, Kieffer-Jaquinod S, Lamant T, Marcellin MN, Garin JR, Rouhier N, Rey P (2012) Affinity chromatography: a valuable strategy to isolate substrates of methionine sulfoxide reductases? Antioxid Redox Signal 16:79–84. https://doi.org/10.1089/ars.2011.4153
Textor S, Bartram S, Kroymann J, Falk KL, Hick A, Pickett JA, Gershenzon J (2004) Biosynthesis of methionine-derived glucosinolates in Arabidopsis thaliana: recombinant expression and characterization of methylthioalkylmalate synthase, the condensing enzyme of the chain-elongation cycle. Planta 218:1026–1035. https://doi.org/10.1007/s00425-003-1184-3
Vogt W (1995) Oxidation of methionyl residues in proteins: tools, targets, and reversal. Free Radic Biol Med 18:93–105. https://doi.org/10.1016/0891-5849(94)00158-G
Wallbach M, Duque Escobar J, Babaeikelishomi R, Stahnke MJ, Blume R, Schroder S, Kruegel J, Maedler K, Kluth O, Kehlenbach RH, Miosge N, Oetjen E (2016) Distinct functions of the dual leucine zipper kinase depending on its subcellular localization. Cell Signal 28:272–283. https://doi.org/10.1016/j.cellsig.2016.01.002
Weissbach H, Resnick L, Brot N (2005) Methionine sulfoxide reductases: history and cellular role in protecting against oxidative damage. Biochim Biophys Acta 1703:203–212. https://doi.org/10.1016/j.bbapap.2004.10.004
Yang SF, Hoffman NE (1984) Ethylene biosynthesis and its regulation in higher plants. Annu Rev Plant Physiol 35:155–189. https://doi.org/10.1146/annurev.pp.35.060184.001103
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins 64:643–651. https://doi.org/10.1002/prot.21018
Zhang XH, Weissbach H (2008) Origin and evolution of the protein-repairing enzymes methionine sulphoxide reductases. Biol Rev Camb Philos Soc 83:249–257. https://doi.org/10.1111/j.1469-185X.2008.00042.x
Zhu J, Ding P, Li Q, Gao Y, Chen F, Xia G (2015) Molecular characterization and expression profile of methionine sulfoxide reductase gene family in maize (Zea mays) under abiotic stresses. Gene 562:159–168. https://doi.org/10.1016/j.gene.2015.02.066
Acknowledgments
This work was supported by the National Foundation for Science and Technology Development (NAFOSTED) under grant number 106-NN.02-2013.46 to DTL. We thank Vadim Gladyshev (Harvard Medical School, Boston, MA, USA) for wild-type and mutant yeast strains used in this study. Mark Dragich and Stephanie K. Dalquist are acknowledged for improving English usage in this manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no conflicts of interest.
Additional information
Handling Editor: Bhumi Nath Tripathi
Rights and permissions
About this article
Cite this article
Le, D.T., Nguyen, KL., Chu, H.D. et al. Function of the evolutionarily conserved plant methionine-S-sulfoxide reductase without the catalytic residue. Protoplasma 255, 1741–1750 (2018). https://doi.org/10.1007/s00709-018-1266-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-018-1266-5