Abstract
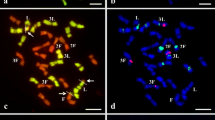
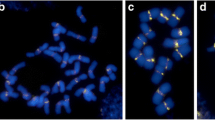
Fragile sites (FSs) in plants have been described for species like Lolium and other grasses. Whereas in humans FSs were shown to be involved in genome instabilities; the consequences of FSs expression in plants are not known yet. To evaluate whether FSs cause karyotype instabilities, we assessed the frequency of micronuclei and lagging chromosomes in meristematic cells, the stability of the DNA content, and the occurrence of neocentromeres in the presumed chromosomal fragments of Lolium perenne, Lolium multiflorum, Festuca arrundinacea, and two Festulolium hybrids. The cell cycle analysis along with flow cytometric genome size measurements showed high stability in all genomes evaluated. Neocentromeric activity was neither observed in the presumed fragments nor in any other chromosomal region, then this is not the mechanism responsible by the stability. However, Fluorescence in situ hybridization (FISH) with a 45S ribosomal DNA (rDNA) probe in combination with YOYO staining of metaphasic chromosomes showed that many extended nucleolus organizing region (NOR) form very thin YOYO-positive chromatin fibers connecting the acentric ‘fragment’ with the centromere-containing chromosome region. The obtained data indicate that the expression of FSs does not result in genome instabilities or neocentromere formation. The FS-containing 45S rDNA carrying chromatin fibers undergo a cell cycle and gene activity-dependent dynamic decondensation process.





Similar content being viewed by others
References
Aguilera A, Garcia-Muse T (2013) Causes of genome instability. Annu Rev Genet 47:1–32. doi:10.1146/annurev-genet-111212-133232
Barlow JH, Faryabi RB, Callen E, Wong N, Malhowski A, Chen HT, Gutierrez-Cruz G, Sun H, McKinnon P, Wright G, Casellas R, Robbiani DF, Staudt L, Fernandez-Capetillo O, Nussenzweig A (2013) Identification of early replicating fragile sites that contribute to genome instability. Cell 152:620–632. doi:10.1016/j.cell.2013.01.006
Brown JD, O’Neill RJ (2010) Chromosomes, conflict, and epigenetics: chromosomal speciation revisited. Annu Rev Genomics Hum Genet 11:291–316. doi:10.1146/annurev-genom-082509-141554
Burrack LS, Berman J (2012) Neocentromeres and epigenetically inherited features of centromeres. Chromosome Res 20:607–619. doi:10.1007/s10577-012-9296-x
Bustamante FO, Rocha LC, Torres GA, Davide LC, Mittelmann A, Techio VH (2014) Distribution of rDNA in diploid and polyploid Lolium multiflorum Lam. and fragile sites in 45S rDNA regions. Crop Sci 54:1–9. doi:10.2135/cropsci2013.05.0325
Butler DK (1992) Ribosomal DNA is a site of chromosome breakage in aneuploids trains of Neurospora. Genetics 131:581–592
Dolezel J, Greilhuber J, Suda J (2007) Estimation of nuclear DNA content in plants using flow cytometry. Nat Protoc 2:2233–2244. doi:10.1038/nprot.2007.310
Freudenreich CH, Kantrow SM, Zakian VA (1998) Expansion and length-dependent fragility of CTG repeats in yeast. Science 279:853–856. doi:10.1126/science.279.5352.853
Galbraith DW, Harkins KR, Maddox JM, Ayres NM, Sharma DP, Firoozabady E (1983) Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 220:1049–1051. doi:10.1126/science.220.4601.1049
Glover TW (2006) Common fragile sites. Cancer Lett 232:4–12. doi:10.1016/j.canlet.2005.08.032
Glover TW, Berger C, Coyle J, Echo B (1984) DNA polymerase a inhibition by aphidicolin induces gaps and breaks at common fragile sites in human chromosomes. Hum Genet 67:136–142
Grabowska-Joachimiak A, Kula A, Gernand-Kliefoth D, Joachimiak AJ (2015) Karyotype structure and chromosome fragility in the grass Phleum echinatum Host. Protoplasma 252:301–6. doi:10.1007/s00709-014-0681-5
Hall AE, Keith KC, Hall SE, Copenhaver GP, Preuss D (2004) The rapidly evolving field of plant centromeres. Curr Opin Plant Biol 7:108–114. doi:10.1016/j.pbi.2004.01.008
Houben A, Schroeder-Reiter E, Nagaki K, Nasuda S, Wanner G, Murata M et al (2007) CENH3 interacts with the centromeric retrotransposon cereba and GC-rich satellites and locates to centromeric substructures in barley. Chromosoma 116:275–283. doi:10.1007/s00412-007-0102-z
Huang J, Ma L, Yang F, S-z F, Li L (2008) 45S rDNA Regions are chromosome fragile sites expressed as gaps in vitro on metaphase chromosomes of root-tip meristematic cells in Lolium spp. PLoS ONE 3, e2167. doi:10.1371/journal.pone.0002167
Huang J, Ma L, Sundararajan S, Fei S-z, Li L (2009) Visualization by atomic force microscopy and FISH of the 45S rDNA gaps in mitotic chromosomes of Lolium perenne. Protoplasma 236:59–65. doi:10.1007/s00709-009-0051-x
Huang M, Li H, Zhang L, Gao F, Wang P, Hu Y, Yan S, Zhao L, Zhang Q, Tan J, Liu X, He S, Li L (2012) Plant 45S rDNA clusters are fragile sites and their instability is associated with epigenetic alterations. PLoS ONE 7, e35139. doi:10.1371/journal.pone.0035139
Longhese MP, Mantiero D, Clerici M (2006) The cellular response to chromosome breakage. Mol Microbiol 60:1099–1108. doi:10.1111/j.1365-2958.2006.05186.x
Marshall OJ, Chueh AC, Wong LH, Choo KA (2008) Neocentromeres: new insights into centromere structure, disease development, and karyotype evolution. Am J Hum Genet 82(2):261–282. doi:10.1016/j.ajhg.2007.11.009
Richards RI (2001) Fragile and unstable chromosomes in cancer: causes and consequences. Trends Genet 17:339–345. doi:10.1016/S0168-9525(01)02303-4
Rocha LC, de Oliveira BF, Silveira RA, Torres GA, Mittelmann A, Techio VH (2015) Functional repetitive sequences and fragile sites in chromosomes of Lolium perenne L. Protoplasma 252:451–460. doi:10.1007/s00709-014-0690-4
Ruiz-Herrera A, Robinson TJ (2007) Chromosomal instability in Afrotheria: fragile sites, evolutionary breakpoints and phylogenetic inference from genome sequence assemblies. BMC Evol Biol 7:199. doi:10.1186/1471-2148-7-199
Ruiz-Herrera A, Castresana J, Robinson TJ (2006) Is mammalian chromosomal evolution driven by regions of genome fragility? Genome Biol 7:R115. doi:10.1186/gb-2006-7-12-r115
Sanei M, Pickering R, Kumke K, Nasuda S, Houben A (2011) Loss of centromeric histone H3 (CENH3) from centromeres precedes uniparental chromosome elimination in interspecific barley hybrids. Proc Natl Acad Sci 108:E498–505
Singh M, Barman AS (2013) Chromosome breakages associated with 45S ribosomal DNA sequences in spotted snakehead fish Channa punctatus. Mol Biol Rep 40:723–729. doi:10.1007/s11033-012-2112-z
Šmarda P, Bureš P, Horová L, Foggi B, Rossi G (2008) Genome size and GC content evolution of Festuca: ancestral expansion and subsequent reduction. Ann Bot 101:421–433. doi:10.1093/aob/mcm307
Sobell HM (1985) Actinomycin and DNA transcription. Proc Natl Acad Sci 82:5328–5331
Song W, Dominska M, Greenwell PW, Petes TD (2014) Genome-wide high-resolution mapping of chromosome fragile sites in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 27 111(21):E2210–8. doi:10.1073/pnas.1406847111
Sutherland GR (1979) Heritable fragile sites on human chromosomes II. Distribution, phenotypic effects, and cytogenetics. Am J Hum Genet 31:136–148
Toledo F, Coquelle A, Svetlova E, Debatisse M (2000) Enhanced flexibility andaphidicolin-induced DNA breaks near mammalian replication origins: implications for replicon mapping and chromosome fragility. Nucleic Acids Res 28:4805–4813
Van Gent DC, Hoeijmakers JH, Kanaar R (2001) Chromosomal stability and the DNA double-stranded break connection. Nat Rev Genet 2:196–206. doi:10.1038/35056049
Waminal NE, Ryu KH, Choi SH, Kim HH (2013) Randomly detected genetically modified (GM) maize (Zea mays L.) near a transport route revealed a fragile 45S rDNA phenotype. PLoS ONE 8, e74060
Wang YH, Griffith J (1996) Methylation of expanded CCG triplet repeat DNA from fragile X syndrome patients enhances nucleosome exclusion. J Biol Chem 271:22937–22940
Wang Y, Bigelow CA, Jiang Y (2009) Ploidy level and DNA content of perennial ryegrass germplasm as determined by flow cytometry. HortSci 44:2049–2052
Yudkin D, Hayward BE, Aladjem MI, Kumari D, Usdin K (2014) Chromosome fragility and the abnormal replication of the FMR1 locus in fragile X syndrome. Hum Mol Genet. doi:10.1093/hmg/ddu006
Zhang N, Threadgill DW, Womack JE (1992) Synteny mapping in the bovine: genes from human chromosome 4. Genomics 14:131–136. doi:10.1016/S0888-7543(05)80295-2
Acknowledgments
The authors thank the Brazilian National Council for Scientific and Technological Development—CNPq, Foundation for Research Support of the State of Minas Gerais—FAPEMIG, Coordination for the Improvement of Higher Education Personnel—CAPES for the financial support to research and scholarship; and the Leibniz Institute of Plant Genetics and Crop Plant Research (IPK).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Handling Editor: Peter Nick
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 2215 kb)
Rights and permissions
About this article
Cite this article
Rocha, L.C., Jankowska, M., Fuchs, J. et al. Decondensation of chromosomal 45S rDNA sites in Lolium and Festuca genotypes does not result in karyotype instability. Protoplasma 254, 285–292 (2017). https://doi.org/10.1007/s00709-016-0942-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-016-0942-6