Abstract
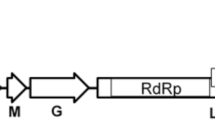
A novel negative-sense single-stranded RNA virus, tentatively named "rose-associated cytorhabdovirus" (RaCV), was identified by high-throughput sequencing. RaCV is 16,067 nucleotides in length and contains eight open reading frames (ORFs 1-8) encoding a nucleocapsid protein (N), a putative phosphoprotein (P), a putative P3 protein (P3), a putative P4 protein (P4), a putative matrix protein (M), a glycoprotein (G), a putative P7 protein (P7), and an RNA-dependent RNA polymerase (L), respectively. The coding genes are flanked by a 3′ leader sequence (228 nt) and a 5′ trailer sequence (251 nt) and are separated by conserved intergenic junctions (3′-AUUCUUUUUG(N)nCUN-5′). Phylogenetic analysis showed that RaCV clustered with yerba mate virus A (YmVA) within the cytorhabdovirus clade, and it exhibited low a degree of nt sequence similarity (<40% identity) to other rhabdoviruses. Amino acid sequence comparisons between the putative proteins of RaCV and the corresponding proteins of other cytorhabdoviruses showed that the sequence identity levels were far below the species demarcation cutoff of 80% for cytorhabdoviruses. These results suggest that RaCV should be classified as a new member of the genus Cytorhabdovirus.


Similar content being viewed by others
Data Availability
All relevant data are openly available in the National Center for Biotechnology Information, and within the manuscript and its additional files.
References
Wang Y, Wang G, Bai J, Zhang Y, Wang Y, Wen S, Li L, Yang Z, Hong N (2021) A novel Actinidia cytorhabdovirus characterized using genomic and viral protein interaction features. Mol Plant Pathol 22:1271–1287
Walker PJ, Freitas-Astúa J, Bejerman N, Blasdell KR, Breyta R, Dietzgen RG, Fooks AR, Kondo H, Kurath G, Kuzmin IV, Ramos-González PL, Shi M, Stone DM, Tesh RB, Tordo N, Vasilakis N, Whitfield AE, ICTV Report Consortium (2022) ICTV virus taxonomy profile: Rhabdoviridae 2022. J Gen Virol 103(6)
Zhang S, Geng J, Gi G, Wang L, Song C, Li S (2019) Research progress of plant rhabdovirus. J Hebei Agric Sci 23:72–75+78
Walker PJ, Firth C, Widen SG, Blasdell KR, Guzman H, Wood TG, Paradkar PN, Holmes EC, Tesh RB, Vasilakis N (2015) Evolution of genome size and complexity in the Rhabdoviridae. PLoS Pathog 11:e1004664
Bejerman N, Dietzgen RG, Debat H (2021) Illuminating the plant Rhabdovirus landscape through metatranscriptomics data. Viruses 13:1304
Qiu Y, Zhang S, Jin J, Xie J, Cao Y, Cao M (2020) Molecular characterization of a novel cytorhabdovirus associated with paper mulberry mosaic disease. Arch Virol 165:2703–2707
Dietzgen RG, Bejerman NE, Goodin MM, Higgins CM, Huot OB, Kondo H, Martin KM, Whitfield AE (2020) Diversity and epidemiology of plant rhabdoviruses. Virus Res 281:197942
Jackson AO, Dietzgen RG, Goodin MM, Bragg JN, Deng M (2005) Biology of plant rhabdoviruses. Annu Rev Phytopathol 43:623–660
Pinheiro-Lima B, Pereira-Carvalho RC, Alves-Freitas DMT, Kitajima EW, Vidal AH, Lacorte C, Godinho MT, Fontenele RS, Faria JC, Abreu EFZ, Varsani A, Ribeiro SG, Melo FL (2020) Transmission of the bean-associated cytorhabdovirus by the Whitefly Bemisia tabaci MEAM1. Viruses 12(9):1028
Whitfield AE, Huot OB, Martin KM, Kondo H, Dietzgen RG (2018) Plant rhabdoviruses-their origins and vector interactions. Curr Opin Virol 33:198–207
Gao Q, Xu WY, Yan T, Fang XD, Cao Q, Zhang ZJ, Ding ZH, Wang Y, Wang XB (2019) Rescue of a plant cytorhabdovirus as versatile expression platforms for planthopper and cereal genomic studies. New Phytol 223:2120–2133
Dietzgen RG, Kondo H, Goodin MM, Kurath G, Vasilakis N (2017) The family Rhabdoviridae: mono-and bipartite negative-sense RNA viruses with diverse genome organization and common evolutionary origins. Virus Res 227:158–170
Walker PJ, Dietzgen RG, Joubert DA, Blasdell KR (2011) Rhabdovirus accessory genes. Virus Res 162:110–125
Zhang X, Liu S, Yang Y, Li M, Hu A, Liu C, Zhang Z, Han C, Wang Y (2021) Virus identification and detection by high-throughput sequencing and RT-PCR from rose plants in Beijing. Acta Phytopathologica Sinica 51:525–535
Bolus S, Al Rwahnih M, Grinstead SC, Mollov D (2021) Rose virus R, a cytorhabdovirus infecting rose. Arch Virol 166:655–658
Blawid R, Silva JMF, Nagata T (2017) Discovering and sequencing new plant viral genomes by next-generation sequencing: description of a practical pipeline. Ann Appl Biol 170:301–314
Villamor DEV, Ho T, Al Rwahnih M, Martin RR, Tzanetakis IE (2019) High throughput sequencing for plant virus detection and discovery. Phytopathology 109:716–725
Bejerman N, Debat H, Dietzgen RG (2020) The plant negative-sense RNA virosphere: virus discovery through new eyes. Front Microbiol 11:588427
Zhao Y, Niu J (2008) Cloning of the terminal regions of apple stem pitting virus in pear trees using 3’ and 5’ RACE. Acta Horticulturae Sinica 35:493–500
Yan T, Zhu JR, Di D, Gao Q, Zhang Y, Zhang A, Yan C, Miao H, Wang XB (2015) Characterization of the complete genome of barley yellow striate mosaic virus reveals a nested gene encoding a small hydrophobic protein. Virology 478:112–122
Huang Y, Zhao H, Luo Z, Chen X, Fang RX (2003) Novel structure of the genome of rice yellow stunt virus: identification of the gene 6-encoded virion protein. J Gen Virol 84:2259–2264
Fránová J, Sarkisova T, Jakešová H, Koloniuk I (2019) Molecular and biological properties of two putative new cytorhabdoviruses infecting Trifolium pratense. Plant Pathol 68:1276–1286
Freitas-Astúa J, Dietzgen RG, Walker PJ, Blasdell KR, Breyta S (2019) Split the genus Nucleorhabdovirus, creating three new genera (Alphanucleorhabdovirus, Betanucleorhabdovirus and Gammanucleorhabdovirus) comprising sixteen species, including 23 six new species, in the Family Rhabdoviridae. International Committee on Taxonomy of Viruses
Acknowledgements
This research was supported by the National Natural Science Foundation of China (32072389), the Innovation Research 2035 Pilot Plan of Southwest University (SWU-XDPY22002), the National Key R&D Program of China (2019YFD1001800), the 111 project (B18044), the Chongqing Postgraduate Research and Innovation Project (CYS22259), the National Training Program of Innovation and Entrepreneurship for Undergraduates (202210635047), and the Municipal Training Program of Innovation and Entrepreneurship for Undergraduates (S202210635179).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Ralf Georg Dietzgen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Y., Yang, M., Yang, H. et al. Identification and molecular characterization of a novel cytorhabdovirus from rose plants (Rosa chinensis Jacq.). Arch Virol 168, 118 (2023). https://doi.org/10.1007/s00705-023-05742-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05742-5