Abstract
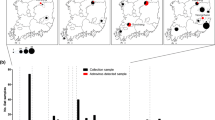
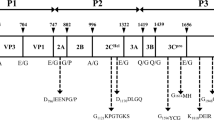
The family Picornaviridae includes important human and animal pathogens that are associated with a wide range of diseases and, in some cases, have zoonotic potential. During epidemiological surveillance of bats, we identified, by next-generation sequencing (NGS) techniques, the presence of picornavirus RNA in a common pipistrelle bat (Pipistrellus pipistrellus). By coupling NGS, primer-walking strategies, and sequence-independent protocols to obtain the sequences of the 5′ and 3′ termini, we reconstructed the genome sequence of picornavirus strain ITA/2017/189/18-155. The genome of the bat picornavirus is 8.2 kb in length and encodes a polyprotein of 2462 amino acids. A comparison of polyprotein sequences revealed that this virus is distantly related (65.1% and 70.9% sequence identity at the nucleotide and amino acid level, respectively) to a bat aichivirus identified in 2010. Phylogenetic analysis showed that this picornavirus clustered closely with members of the genus Kobuvirus, which also includes human and animal aichiviruses. The identification of aichiviruses in several animal hosts is providing hints that will lead to an understanding of their origin and evolutionary patterns.


Similar content being viewed by others
References
Tracy S, Chapman NM, Drescher KM et al (2006) Evolution of virulence in picornaviruses. Curr Top Microbiol Immunol. https://doi.org/10.1007/3-540-26397-7_7
Zell R (2018) Picornaviridae—the ever-growing virus family. Arch Virol. https://doi.org/10.1007/s00705-017-3614-8
International Committee on Taxonomy of Viruses (ICTV) Virus taxonomy: 2018b release. https://talk.ictvonline.org/taxonomy/. Accessed Nov 2019
Calisher CH, Childs JE, Field HE et al (2006) Bats: important reservoir hosts of emerging viruses. Clin Microbiol Rev. https://doi.org/10.1128/CMR.00017-06
Diakoudi G, Lanave G, Moreno A et al (2019) Surveillance for adenoviruses in bats in Italy. Viruses. https://doi.org/10.3390/v11060523
Jamnikar-Ciglenecki U, Toplak I, Kuhar U (2017) Complete genome of Chronic bee paralysis virus strain SLO/M92/2010, detected from Apis mellifera carnica. Genome Announc. https://doi.org/10.1128/genomeA.00602-17
Nurk S, Bankevich A, Antipov D et al (2013) Assembling single-cell genomes and mini-metagenomes from chimeric MDA products. J Comput Biol. https://doi.org/10.1089/cmb.2013.0084
Scotto-Lavino E, Du G, Frohman MA (2007) 5′ end cDNA amplification using classic RACE. Nat Protoc. https://doi.org/10.1038/nprot.2006.480
Scotto-Lavino E, Du G, Frohman MA (2007) 3′ end cDNA amplification using classic RACE. Nat Protoc. https://doi.org/10.1038/nprot.2006.481
Tcherepanov V, Ehlers A, Upton C (2006) Genome annotation transfer utility (GATU): rapid annotation of viral genomes using a closely related reference genome. BMC Genomics. https://doi.org/10.1186/1471-2164-7-150
Larkin MA, Blackshields G, Brown NP et al (2007) Clustal W and Clustal X version 20. Bioinformatics https://doi.org/10.1093/bioinformatics/btm404
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. https://doi.org/10.1093/molbev/msy096
Kemenesi G, Zhang D, Marton S et al (2015) Genetic characterization of a novel picornavirus detected in miniopterus schreibersii bats. J Gen Virol. https://doi.org/10.1099/jgv.0.000028
Wu Z, Yang L, Ren X et al (2016) Deciphering the bat virome catalog to better understand the ecological diversity of bat viruses and the bat origin of emerging infectious diseases. ISME J. https://doi.org/10.1038/ismej.2015.138
Yamashita T, Kobayashi S, Sakae K et al (1991) Isolation of cytopathic small round viruses with BS-C-1 cells from patients with gastroenteritis. J Infect Dis. https://doi.org/10.1093/infdis/164.5.954
Olarte-Castillo XA, Heeger F, Mazzoni CJ et al (2015) Molecular characterization of canine kobuvirus in wild carnivores and the domestic dog in Africa. Virology. https://doi.org/10.1016/j.virol.2015.01.010
Funding
This work was supported by the Slovenian Research Agency, Program Group Animal Health, Environment and Food Safety (P4-0092).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The Authors declare that they do not have conflict of interest.
Ethical approval
The research did not involve human partecipants and/or experiments/procedures on live animals.
Accession number
The sequence reported here was deposited in the GenBank database under the accession number MN602325.
Additional information
Handling Editor: Ana Cristina Bratanich.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Diakoudi, G., Jamnikar-Ciglenečki, U., Lanave, G. et al. Genome sequence of an aichivirus detected in a common pipistrelle bat (Pipistrellus pipistrellus). Arch Virol 165, 1019–1022 (2020). https://doi.org/10.1007/s00705-020-04548-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-020-04548-z