Abstract
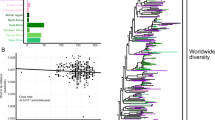
Kaposi’s sarcoma-associated herpesvirus (KSHV) has become widely dispersed worldwide since it was first reported in 1994, but the seroprevalence of KSHV varies geographically. KSHV is relatively ubiquitous in Mediterranean areas and the Xinjiang Uygur Autonomous Region, China. The origin of KSHV has long been puzzling. In the present study, we collected and analysed 154 KSHV ORF-K1 sequences obtained from samples originating from Xinjiang, Italy, Greece, Iran and southern Siberia using Bayesian evolutionary analysis in BEAST to test the hypothesis that KSHV was introduced into Xinjiang via the ancient Silk Road. According to the phylogenetic analysis, 72 sequences were subtype A and 82 subtype C, with C2 (n = 56) being the predominant subtype. The times to the most recent common ancestors (tMRCAs) of KSHV were 29,872 years (95% highest probability density [HPD], 26,851-32,760 years) for all analysed sequences and 2037 years (95% HPD, 1843-2229 years) for Xinjiang sequences in particular. The tMRCA of Xinjiang KSHV was exactly matched with the time period of the ancient Silk Road approximately two thousand years ago. This route began in Chang’an, the capital of the Han dynasty of China, and crossed Central Asia, ending in the Roman Empire. The evolution rate of KSHV was slow, with 3.44 × 10−6 substitutions per site per year (95% HPD, 2.26 × 10−6 to 4.71 × 10−6), although 11 codons were discovered to be under positive selection pressure. The geographic distances from Italy to Iran and Xinjiang are more than 4000 and 7000 kilometres, respectively, but no explicit relationship between genetic distance and geographic distance was detected.



Similar content being viewed by others
References
Centers for Disease Control (1982) Current trends update on acquired immune deficiency syndrome (AIDS)—United States. MMWR 31:277–279
Antman K, Chang Y (2000) Kaposi’s sarcoma. N Engl J Med 342:1027–1038
Barre-Sinoussi F, Chermann JC, Rey F, Nugeyre MT, Chamaret S, Gruest J, Dauguet C, Axler-Blin C, Vezinet-Brun F, Rouzioux C, Rozenbaum W, Montagnier L (1983) Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS). Science (New York, NY) 220:868–871
Biggar RJ, Whitby D, Marshall V, Linhares AC, Black F (2000) Human herpesvirus 8 in Brazilian Amerindians: a hyperendemic population with a new subtype. J Infect Dis 181:1562–1568
Blauvelt A, Sei S, Cook PM, Schulz TF, Jeang KT (1997) Human herpesvirus 8 infection occurs following adolescence in the United States. J Infect Dis 176:771–774
Cao Y, Minhas V, Tan X, Huang J, Wang B, Zhu M, Gao Y, Zhao T, Yang L, Wood C (2014) High prevalence of early childhood infection by Kaposi’s sarcoma-associated herpesvirus in a minority population in China. Clin Microbiol Infect: Off Publ Eur Soc Clin Microbiol Infect Dis 20:475–481
Cassar O, Bassot S, Plancoulaine S, Quintana-Murci L, Harmant C, Gurtsevitch V, Senyuta NB, Yakovleva LS, de The G, Gessain A (2010) Human herpesvirus 8, Southern Siberia. Emerg Infect Dis 16:580–582
Cesarman E, Chang Y, Moore PS, Said JW, Knowles DM (1995) Kaposi’s sarcoma-associated herpesvirus-like DNA sequences in AIDS-related body-cavity-based lymphomas. N Engl J Med 332:1186–1191
Cook PM, Whitby D, Calabro ML, Luppi M, Kakoola DN, Hjalgrim H, Ariyoshi K, Ensoli B, Davison AJ, Schulz TF (1999) Variability and evolution of Kaposi’s sarcoma-associated herpesvirus in Europe and Africa. International Collaborative Group. AIDS (London, England) 13:1165–1176
Cordiali-Fei P, Trento E, Giovanetti M, Lo Presti A, Latini A, Giuliani M, D’Agosto G, Bordignon V, Cella E, Farchi F, Ferraro C, Lesnoni La Parola I, Cota C, Sperduti I, Vento A, Cristaudo A, Ciccozzi M, Ensoli F (2015) Analysis of the ORFK1 hypervariable regions reveal distinct HHV-8 clustering in Kaposi’s sarcoma and non-Kaposi’s cases. J Exp Clin Cancer Res: CR 34:1
Davidovici B, Karakis I, Bourboulia D, Ariad S, Zong J, Benharroch D, Dupin N, Weiss R, Hayward G, Sarov B, Boshoff C (2001) Seroepidemiology and molecular epidemiology of Kaposi’s sarcoma-associated herpesvirus among Jewish population groups in Israel. J Natl Cancer Inst 93:194–202
Dedicoat M, Newton R (2003) Review of the distribution of Kaposi’s sarcoma-associated herpesvirus (KSHV) in Africa in relation to the incidence of Kaposi’s sarcoma. Br J Cancer 88:1–3
deMenocal PB, Stringer C (2016) Human migration: climate and the peopling of the world. Nature 538:49–50
Dilnur P, Katano H, Wang ZH, Osakabe Y, Kudo M, Sata T, Ebihara Y (2001) Classic type of Kaposi’s sarcoma and human herpesvirus 8 infection in Xinjiang, China. Pathol Int 51:845–852
Drummond AJ, Ho SY, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4:e88
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29:1969–1973
Duprez R, Hbid O, Afonso P, Quach H, Belloul L, Fajali N, Ismaili N, Benomar H, Hassane Tahri E, Huerre M, Quintana-Murci L, Gessain A (2006) Molecular epidemiology of the HHV-8 K1 gene from Moroccan patients with Kaposi’s sarcoma. Virology 353:121–132
Elisseeff V (2001) The Silk Road: highways of culture and commerce. UNESCO Publishing/Berghahn Books, New York
Engels EA, Atkinson JO, Graubard BI, McQuillan GM, Gamache C, Mbisa G, Cohn S, Whitby D, Goedert JJ (2007) Risk factors for human herpesvirus 8 infection among adults in the United States and evidence for sexual transmission. J Infect Dis 196:199–207
Gao SJ, Kingsley L, Li M, Zheng W, Parravicini C, Ziegler J, Newton R, Rinaldo CR, Saah A, Phair J, Detels R, Chang Y, Moore PS (1996) KSHV antibodies among Americans, Italians and Ugandans with and without Kaposi’s sarcoma. Nat Med 2:925–928
Goebel T, Waters MR, O’Rourke DH (2008) The late Pleistocene dispersal of modern humans in the Americas. Science (New York, NY) 319:1497–1502
Goedert JJ, Martin MP, Vitale F, Lauria C, Whitby D, Qi Y, Gao X, Carrington M (2016) Risk of classic Kaposi sarcoma with combinations of killer immunoglobulin-like receptor and human leukocyte antigen loci: a population-based case–control study. J Infect Dis 213:432–438
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Iscovich J, Boffetta P, Franceschi S, Azizi E, Sarid R (2000) Classic kaposi sarcoma: epidemiology and risk factors. Cancer 88:500–517
Jalilvand S, Shoja Z, Mokhtari-Azad T, Nategh R, Gharehbaghian A (2011) Seroprevalence of human herpesvirus 8 (HHV-8) and incidence of Kaposi’s sarcoma in Iran. Infect Agents Cancer 6:5
Liu W, Martinon-Torres M, Cai YJ, Xing S, Tong HW, Pei SW, Sier MJ, Wu XH, Edwards RL, Cheng H, Li YY, Yang XX, de Castro JM, Wu XJ (2015) The earliest unequivocally modern humans in southern China. Nature 526:696–699
Mancuso R, Biffi R, Valli M, Bellinvia M, Tourlaki A, Ferrucci S, Brambilla L, Delbue S, Ferrante P, Tinelli C, Clerici M (2008) HHV8 a subtype is associated with rapidly evolving classic Kaposi’s sarcoma. J Med Virol 80:2153–2160
Ouyang X, Zeng Y, Fu B, Wang X, Chen W, Fang Y, Luo M, Wang L (2014) Genotypic analysis of Kaposi’s sarcoma-associated herpesvirus from patients with Kaposi’s sarcoma in Xinjiang, China. Viruses 6:4800–4810
Pond SL, Frost SD (2005) Datamonkey: rapid detection of selective pressure on individual sites of codon alignments. Bioinformatics (Oxford, England) 21:2531–2533
Pond SL, Frost SD, Muse SV (2005) HyPhy: hypothesis testing using phylogenies. Bioinformatics (Oxford, England) 21:676–679
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics (Oxford, England) 14:817–818
Regamey N, Cathomas G, Schwager M, Wernli M, Harr T, Erb P (1998) High human herpesvirus 8 seroprevalence in the homosexual population in Switzerland. J Clin Microbiol 36:1784–1786
Rohner E, Wyss N, Heg Z, Faralli Z, Mbulaiteye SM, Novak U, Zwahlen M, Egger M, Bohlius J (2016) HIV and human herpesvirus 8 co-infection across the globe: systematic review and meta-analysis. Int J Cancer 138:45–54
Russo JJ, Bohenzky RA, Chien MC, Chen J, Yan M, Maddalena D, Parry JP, Peruzzi D, Edelman IS, Chang Y, Moore PS (1996) Nucleotide sequence of the Kaposi sarcoma-associated herpesvirus (HHV8). Proc Natl Acad Sci USA 93:14862–14867
Soulier J, Grollet L, Oksenhendler E, Cacoub P, Cazals-Hatem D, Babinet P, d’Agay MF, Clauvel JP, Raphael M, Degos L et al (1995) Kaposi’s sarcoma-associated herpesvirus-like DNA sequences in multicentric Castleman’s disease. Blood 86:1276–1280
Stolka K, Ndom P, Hemingway-Foday J, Iriondo-Perez J, Miley W, Labo N, Stella J, Abassora M, Woelk G, Ryder R, Whitby D, Smith JS (2014) Risk factors for Kaposi’s sarcoma among HIV-positive individuals in a case control study in Cameroon. Cancer Epidemiol 38:137–143
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tornesello ML, Biryahwaho B, Downing R, Hatzakis A, Alessi E, Cusini M, Ruocco V, Katongole-Mbidde E, Loquercio G, Buonaguro L, Buonaguro FM (2010) Human herpesvirus type 8 variants circulating in Europe, Africa and North America in classic, endemic and epidemic Kaposi’s sarcoma lesions during pre-AIDS and AIDS era. Virology 398:280–289
Touloumi G, Kaklamanis L, Potouridou I, Katsika-Hatziolou E, Stratigos J, Mueller N, Hatzakis A (1997) The epidemiologic profile of Kaposi’s sarcoma in Greece prior to and during the AIDS era. Int J Cancer 70:538–541
Wang X, Wang H, He B, Hui Y, Lv G, Li L, Wen H (2012) Virological and molecular characterization of Kaposi’s sarcoma-associated herpesvirus strains from Xinjiang, China. Eur J Clin Microbiol Infect Dis 31:53–59
White TD, Asfaw B, DeGusta D, Gilbert H, Richards GD, Suwa G, Howell FC (2003) Pleistocene Homo sapiens from Middle Awash, Ethiopia. Nature 423:742–747
Wu X, Wan XF, Wu G, Xu D, Lin G (2006) Phylogenetic analysis using complete signature information of whole genomes and clustered Neighbour-Joining method. Int J Bioinform Res Appl 2:219–248
Zhang D, Pu X, Wu W, Jin Y, Juhear M, Wu X (2008) Genotypic analysis on the ORF-K1 gene of human herpesvirus 8 from patients with Kaposi’s sarcoma in Xinjiang, China. J Genet Genomics = Yi chuan xue bao 35:657–663
Zhang T, Shao X, Chen Y, Zhang T, Minhas V, Wood C, He N (2012) Human herpesvirus 8 seroprevalence, China. Emerg Infect Dis 18:150–152
Zong JC, Ciufo DM, Alcendor DJ, Wan X, Nicholas J, Browning PJ, Rady PL, Tyring SK, Orenstein JM, Rabkin CS, Su IJ, Powell KF, Croxson M, Foreman KE, Nickoloff BJ, Alkan S, Hayward GS (1999) High-level variability in the ORF-K1 membrane protein gene at the left end of the Kaposi’s sarcoma-associated herpesvirus genome defines four major virus subtypes and multiple variants or clades in different human populations. J Virol 73:4156–4170
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, Z., Fang, Q., Zuo, J. et al. Was Kaposi’s sarcoma-associated herpesvirus introduced into China via the ancient Silk Road? An evolutionary perspective. Arch Virol 162, 3061–3068 (2017). https://doi.org/10.1007/s00705-017-3467-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3467-1