Abstract
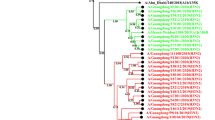
In this study, we investigated the molecular epidemiology and evolution of influenza viruses from patients infected during the 2013–2014 influenza season in Beijing. A phylogenetic analysis of the hemagglutinin (HA) and neuraminidase (NA) sequences of influenza A and B viruses from 18 patients (6 A(H1N1)pdm09, 4 H3N2, and 8 influenza B virus) was performed. Among the influenza A viruses, A(H1N1)pdm09 was the dominant subtype, whereas the B/Yamagata lineage was predominant for influenza B. The influenza B HA and NA strains in Beijing were dominated by reassortants derived from the Yamagata lineage and the Victoria lineage, respectively. All six A(H1N1)pdm09 strains fell into the 6B genetic group with amino acid substitutions D97N, S185T, K163Q, and A256T; the four H3N2 strains fell into genetic group 3C.3 with substitutions T128A, R142G, N145S, and V186G, and the eight influenza B strains were categorized into subgroup 3.1 and harbored an N217S mutation. Two new mutations (K180Q and G187E at the Sa and Ca antigenic sites of the H1 segment, respectively), which were not detected during the preceding influenza season, were identified. Mutations N131K, S165I, N181Y, and D212N in HA of influenza B mapped to the 120-loop, 150-loop, 160-loop, and 190-helix, respectively. Our results reveal the molecular epidemiology and phylogenetic characteristics of influenza viruses within a single geographic location and can have implications for vaccination selection in northern China.





Similar content being viewed by others
References
Air GM (2012) Influenza neuraminidase. Influenza Other Respir Viruses 6:245–256
Ali G, Amer HM, Almajhdi FN (2013) Hemagglutinin and neuraminidase genes of influenza B viruses circulating in Riyadh, Saudi Arabia during 2010–2011: Evolution and sequence analysis. J Med Virol 86:1003–1016
Berton MT, Webster RG (1985) The antigenic structure of the influenza B virus hemagglutinin: operational and topological mapping with monoclonal antibodies. Virology 143:583–594
Chen G-W, Shih S-R, Hsiao M-R, Chang S-C, Lin S-H, Sun C-F, Tsao K-C (2007) Multiple genotypes of influenza B viruses cocirculated in Taiwan in 2004 and 2005. J Clin Microbiol 45:1515–1522
Chen W, Zhong Y, Qin Y, Sun S, Li Z (2012) The evolutionary pattern of glycosylation sites in influenza virus (H5N1) hemagglutinin and neuraminidase. PLoS One 7:e49224
Cherry JL, Lipman DJ, Nikolskaya A, Wolf YI (2009) Evolutionary dynamics of N-glycosylation sites of influenza virus hemagglutinin. PLoS Curr 1:RRN1001
Chi XS, Bolar TV, Zhao P, Rappaport R, Cheng S-M (2003) Cocirculation and evolution of two lineages of influenza B viruses in Europe and Israel in the 2001-2002 season. J Clin Microbiol 41:5770–5773
CNIC NIC (2013-2014) Weekly Influenza Surveillance
Dangi T, Jain B, Singh AK, Singh J, Kumar R, Dwivedi M, Verma AK, Chadha MS, Jain A (2014) Molecular characterization of circulating pandemic strains of influenza A virus during 2012 to 2013 in Lucknow (India). J Med Virol 86:2134–2141
Fang Q, Gao Y, Chen M, Guo X, Yang X, Yang X, Wei L (2014) Molecular epidemiology and evolution of A (H1N1) pdm09 and H3N2 virus during winter 2012–2013 in Beijing China. Infect Genet Evol 26:228–240
Foster PL (2000) Adaptive mutation: implications for evolution. BioEssays 22:1067–1074
Ghedin E, Sengamalay NA, Shumway M, Zaborsky J, Feldblyum T, Subbu V, Spiro DJ, Sitz J, Koo H, Bolotov P (2005) Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution. Nature 437:1162–1166
Graham M, Liang B, Van Domselaar G, Bastien N, Beaudoin C, Tyler S, Kaplen B, Landry E, Li Y (2011) Nationwide molecular surveillance of pandemic H1N1 influenza A virus genomes: Canada, 2009. PLoS One 6:e16087
Hatakeyama S, Sugaya N, Ito M, Yamazaki M, Ichikawa M, Kimura K, Kiso M, Shimizu H, Kawakami C, Koike K (2007) Emergence of influenza B viruses with reduced sensitivity to neuraminidase inhibitors. Jama 297:1435–1442
Hite LK, Glezen WP, Demmler GJ, Munoz FM (2007) Medically attended pediatric influenza during the resurgence of the Victoria lineage of influenza B virus. Int J Infect Dis 11:40–47
Hu W (2009) Analysis of correlated mutations, stalk motifs, and phylogenetic relationship of the 2009 influenza A virus neuraminidase sequences. J Biomed Sci Eng 2:550–558
Kanegae Y, Sugita S, Endo A, Ishida M, Senya S, Osako K, Nerome K, Oya A (1990) Evolutionary pattern of the hemagglutinin gene of influenza B viruses isolated in Japan: cocirculating lineages in the same epidemic season. J Virol 64:2860–2865
Kiso M, Mitamura K, Sakai-Tagawa Y, Shiraishi K, Kawakami C, Kimura K, Hayden FG, Sugaya N, Kawaoka Y (2004) Resistant influenza A viruses in children treated with oseltamivir: descriptive study. Lancet 364:759–765
Langenhorst D, Gogishvili T, Ribechini E, Kneitz S, McPherson K, Lutz MB, Hünig T (2012) Molecular Characterization of the Predominant Influenza A (H1N1) pdm09 Virus in Mexico, December 2011-February 2012. PLoS One 7:e50116
Li W-C, Shih S-R, Huang Y-C, Chen G-W, Chang S-C, Hsiao M-J, Tsao K-C, Lin T-Y (2008) Clinical and genetic characterization of severe influenza B-associated diseases during an outbreak in Taiwan. J Clin Virol 42:45–51
Mao HY, Lu YY, Yan JY (2008) Molecular and antigenic characteristics of influenza B virus isolated in Zhejiang province in 2006. Zhonghua Liu Xing Bing Xue Za Zhi 29:413–414
Maurer-Stroh S, Ma J, Lee RT, Sirota FL, Eisenhaber F (2009) Mapping the sequence mutations of the 2009 H1N1 influenza A virus neuraminidase relative to drug and antibody binding sites. Biol Direct 4:18 (discussion 18)
McCullers JA, Wang GC, He S, Webster RG (1999) Reassortment and insertion-deletion are strategies for the evolution of influenza B viruses in nature. J Virol 73:7343–7348
Nerome R, Hiromoto Y, Sugita S, Tanabe N, Ishida M, Matsumoto M, Lindstrom SE, Takahashi T, Nerome K (1998) Evolutionary characteristics of influenza B virus since its first isolation in 1940: dynamic circulation of deletion and insertion mechanism. Arch Virol 143:1569–1583
Neumann G, Noda T, Kawaoka Y (2009) Emergence and pandemic potential of swine-origin H1N1 influenza virus. Nature 459:931–939
Padilla C, Condori F, Huaringa M, Marcos P, Rojas N, Gutierrez V, Cáceres O (2014) Full genome analysis of influenza A (H1N1) pdm09 virus isolated from Peru, 2013. Genome Announc 2:e00114–e00191
Rota PA, Wallis TR, Harmon MW, Rota JS, Kendal AP, Nerome K (1990) Cocirculation of two distinct evolutionary lineages of influenza type B virus since 1983. Virology 175:59–68
Roy T, Agrawal AS, Mukherjee A, Mishra AC, Chadha MS, Kaur H, Chawla-Sarkar M (2011) Surveillance and molecular characterization of human influenza B viruses during 2006-2010 revealed co-circulation of Yamagata-like and Victoria-like strains in eastern India. Infect Genet Evol 11:1595–1601
Shen J, Kirk BD, Ma J, Wang Q (2009) Diversifying selective pressure on influenza B virus hemagglutinin. J Med Virol 81:114–124
Shibata S, Yamamoto-Goshima F, Maeno K, Hanaichi T, Fujita Y, Nakajima K, Imai M, Komatsu T, Sugiura S (1993) Characterization of a temperature-sensitive influenza B virus mutant defective in neuraminidase. J Virol 67:3264–3273
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tan Y, Guan W, Lam TT, Pan S, Wu S, Zhan Y, Viboud C, Holmes EC, Yang Z (2013) Differing epidemiological dynamics of influenza B virus lineages in Guangzhou, southern China, 2009-2010. J Virol 87:12447–12456
Tsai H-P, Wang H-C, Kiang D, Huang S-W, Kuo P-H, Liu C-C, Su I-J, Wang J-R (2006) Increasing appearance of reassortant influenza B virus in Taiwan from 2002 to 2005. J Clin Microbiol 44:2705–2713
Verhoeyen M, Van Rompuy L, Jou WM, Huylebroeck D, Fiers W (1983) Complete nucleotide sequence of the influenza B/Singapore/222/79 virus hemagglutinin gene and comparison with the B/Lee/40 hemagglutinin. Nucleic Acids Res 11:4703–4712
Wang Q, Cheng F, Lu M, Tian X, Ma J (2008) Crystal structure of unliganded influenza B virus hemagglutinin. J Virol 82:3011–3020
WHO (2009) WHO recommended surveillance standards, Second edition [cited 2009 Aug 19]
WHO (2012) Influenza Centre London. February 2012 interim report. Report prepared for the WHO annual consultation on the composition of influenza vaccine for the Northern Hemisphere. 20th–22nd February 2012
WHO (2014) Recommended composition of influenza virus vaccines for use in the 2014-2015 northern hemisphere influenza season
WHO (2014) Report prepared for the WHO annual consultation on the composition of influenza vaccine for the Northern Hemisphere 2014/15
WHO (2014) Northern Hemisphere Vaccine Recommendation Meeting for 2014-15 influenza season
Acknowledgements
We thank the doctors (Meng Xi, Chunling Sun, Jianying Zhu, Ying Zuo, Yanmin Zhang, and Yi Zhang) in the Department of Infectious Disease of Peking University People’s Hospital for collecting cases and throat/nasal specimens. We also thank the laboratory staff (Xu Cong, Xiaoben Pan, Jinchao Han, and Huan Mai) in Peking University Hepatology Institute of Peking University People’s Hospital for their help with the experiments. This work was supported by the research and development funds of Peking University People's Hospital (RDC2014-06 No. 2118000586).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fang, Q., Gao, Y., Chen, M. et al. Molecular epidemiology and evolution of influenza A and B viruses during winter 2013-2014 in Beijing, China. Arch Virol 160, 1083–1095 (2015). https://doi.org/10.1007/s00705-015-2362-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2362-x