Abstract
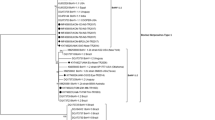
Bovine coronaviruses (BCoVs) isolated in Japan consist of four genetic groups, as determined by phylogenetic analysis using the polymorphic region (aa 456–592) of the S glycoprotein gene. Japanese field isolates of BCoV, reference Kakegawa strain, and vaccine strain 66/H were analyzed for their antigenic properties by indirect immunofluorescence and neutralization testing. There were no significant differences observed among these BCoVs in direct immunofluorescence tests. However, antigenic differences were observed between BCoVs in the neutralization tests, although there was no clear indication of a distinct serotype. A monoclonal antibody, 4H4, against the Kakegawa strain belonging to group 1 lacked significant neutralizing activity for viruses of groups 2, 3, and 4. Therefore, we speculate that the genetic differences between these groups may have altered their antigenicity. Analysis of mutant viruses resistant to neutralization by 4H4 revealed that the antigenic site of the Kakegawa strain maps to amino acid position 284 of the S glycoprotein. This site is not homologous to a known antigenic site (aa 528) of the Quebec strain belonging to group 1, and it is not located in the conformational domain comprising domain I (aa 351–403) and domain II (aa 517–621). This amino acid constitutes a neutralization epitope of BCoV, which is distinct from aa 528 of the Quebec strain. These results indicate antigenic evolution of BCoV between the genetic groups circulating in Japan.
Similar content being viewed by others
Introduction
Bovine coronavirus (BCoV) infects the respiratory and intestinal tracts of cattle. Three types of clinical symptoms occur in cattle infected with BCoV: (1) calf diarrhea (CD) [22]; (2) diarrhea in adult cattle, called winter dysentery (WD) [27]; and (3) respiratory symptoms, including shipping fever in feedlot cattle [31].
BCoV is a member of the order Nidovirales, family Coronaviridae, subfamily Coronavirinae, genus Betacoronavirus, species Betacoronavirus 1, along with human coronavirus OC43 (HCoV-OC43), porcine hemagglutinating encephalomyelitis virus (PHEV), and equine coronavirus (ECoV) [9, 24]. BCoV possesses a single-stranded, positive-sense, non-segmented RNA genome that is 31 kb in length. The virion contains five structural proteins: nucleocapsid (N) protein, transmembrane (M) protein, spike (S) protein, small envelope (E) protein, and hemagglutinin-esterase (HE) protein, which is specific to members of the genus Betacoronavirus [20].
Among these proteins, the S glycoprotein of the coronavirus forms large, petal-shaped spikes on the surface of the virion and is responsible for viral binding to host-cell receptors [8, 18], induction of neutralizing antibody [32, 35], and hemagglutinating activity [30]. Sequences of S glycoproteins are variable, and mutations in this region have been associated with altered antigenicity and viral pathogenicity [2, 11].
BCoV infection occurs in many countries, with a high prevalence in countries where cows are kept as livestock. However, to our knowledge, molecular epidemiological analysis among epidemic BCoVs has not been reported before, except in a report from Canada [26]. Genetic analysis among domestic epidemic BCoVs has been reported recently in the United States [10], Korea [13, 17, 25], Denmark [21], and Brazil [4]. There has been no report of nationwide genetic or antigenic analysis of BCoVs in Japan. Recently, we attempted to clarify the correlation among epidemic viruses in Japan by isolating viruses from feces or nasal discharge of cattle with BCoV infections and analyzing their genes. For comparison, we used a polymorphic region (aa 456–592) in the S gene [26]. We compared (1) isolates of causal viruses in 13 prefectures (148 isolates) in the period from 1999 to 2008, (2) the strain isolated from the first infection case that occurred in 1976 (Kakegawa strain) and the vaccine strain (66/H), and (3) foreign strains registered in the database. Phylogenetic analysis of the polymorphic region of the S gene grouped these strains into four clusters, which we named groups 1 to 4 [14, 15]. Group 1 included the Kakegawa and vaccine strains and foreign strains identified from the 1960s to 1970s. In 1999, most epidemic viruses in Japan belonged to group 2, but the predominant group changed to group 3 from 2002 to 2004, and viruses of group 4 have been isolated since 2004. There was no regional difference in years of incidence and groups, but it was revealed that since 2005, group 4 is the predominant lineage and the most widespread throughout Japan. Although BCoV is thought to have one serotype, its antigenicity and pathogenicity may vary among clusters; therefore, a comparative analysis of antigenicity and pathogenicity among clusters should be conducted for undertaking preventive measures against this disease. Herein, we report the results of a comparative analysis of antigenicity among bovine coronaviruses that belong to distinct groups as determined by phylogenetic analysis. Furthermore, we identified the location of a neutralization antigenic site of BCoV using a neutralizing monoclonal antibody (mAb) and neutralization-resistant mutants.
Materials and methods
Viruses and antisera
Hyperimmune antisera against Kakegawa (group 1), vaccine strain 66/H (1), Ishikawa/2/99 (2), Hokkaido/12/03 (3), Kumamoto/1/07 (4), Hokkaido/25/05 (4), Hokkaido/40/08 (4), and Hokkaido/45/08 (4) strains [14, 15] were prepared in rabbits (Japanese white rabbit). All viruses except for Kakegawa and 66/H were used at low passage number (up to passage 2 or 3), because increasing viral passage cycles in cell culture may affect the antigenic composition of the virus [12]. Each of the eight virus strains was propagated in HRT-18G human rectal tumor cells and concentrated by ultracentrifugation. Rabbits were inoculated subcutaneously at 2-week intervals for 6 weeks with each concentrated, purified virus strain emulsified in a 1:1 mixture with Freund’s complete adjuvant or incomplete adjuvant. The rabbits were bled for serum collection at 2 weeks after the final inoculation.
Indirect immunofluorescence test
The antigenicity of each group of BCoVs was examined by indirect immunofluorescence testing. HRT-18G cell monolayers grown in an 8-well Lab-Tek II chamber slide (Thermo Fisher Scientific K.K., Yokohama, Japan) were fixed in acetone 3 days after virus inoculation. Rabbit sera against each virus were diluted from 1000 to 2000 and incubated with the HRT-18G cells. For the detection phase, fluorescein isothiocyanate (FITC)-conjugated goat anti-rabbit IgA, IgG, and IgM (H + L) serum (Cappel, Cooper Biomedical, Inc., Malvern, PA, USA) was used.
Cross-neutralization test
The neutralizing antigenicity of viruses of each group was examined by using virus neutralization (VN) tests. Briefly, twofold serially diluted rabbit sera against each virus were reacted in duplicate with one hundred 50 % tissue culture infective doses (TCID50) of each virus, followed by incubation for 1 h at 37 °C. After incubation, the serum-virus mixture was transferred onto the monolayered HRT-18G cells cultured in microplates and incubated for 5 days at 37 °C. The neutralization antibody titers were expressed as the reciprocal of the highest serum dilution that completely inhibited the cytopathic effect (CPE). The antigenic relationship (R) between the strains was calculated using the following formula [1, 6]:
where r1 = heterologous titer (strain 2)/homologous titer (strain 1); and r2 = heterologous titer (strain 1)/homologous titer (strain 2).
Neutralizing antibody titers of bovine sera against viruses of each group
To elucidate the antigenicity of the most recent predominant group in Japan, VN tests were conducted on bovine sera obtained by experimental infection using Kumamoto/1/07 virus as an inoculum. In brief, five-month-old cattle were inoculated orally with 106 TICD50/ml of the virus in 50 ml of tissue culture fluid. Blood was collected after inoculation, and neutralizing antibody titers against viruses of each group were examined.
Neutralization test using mAb 4H4 against viruses of each group
The neutralizing mAb 4H4 against Kakegawa has been described by Sato et al. [29]. Neutralizing antibody titers of 4H4 against viruses of each group in supernatants of hybridoma cultures and ascites of mice were examined by VN tests, as described above.
SDS-PAGE and western blotting
The BCoV Kakegawa strain was purified by ultracentrifugation and resuspended in 2× sample buffer [19] with or without 2-mercaptoethanol (2-ME). After boiling for 5 min, viral proteins were fractionated by 10 % SDS-PAGE. Proteins were then transferred to Immun-Blot PVDF membrane (Bio-Rad Laboratories, Tokyo, Japan) by electroblotting at 15 V for 30 min in Tris-glycine/methanol buffer as described by Bjerrum and Shafer-Nielsen [3]. The blocking reaction was performed with 5 % skim milk in PBS containing 0.1 % Tween-20 (PBST) at room temperature overnight. The membrane was incubated with a 1:2500 dilution of ascitic fluid from 4H4 in PBST for 1 h at room temperature. Next, the membrane was washed four times with PBST and then incubated with a 1:20,000 dilution of ECL anti-mouse IgG HRPO (GE Healthcare, Tokyo, Japan) for 1 h at room temperature. After washing as described above, the membranes were developed by reaction with substrate in ECL Plus Western Blotting Detection System (GE Healthcare) and visualized by autoradiography.
Generation of neutralization-resistant mutants
Neutralization-resistant mutants were generated using hybridoma supernatants of 4H4. Tenfold serial dilutions of Kakegawa strain were made in 96-well tissue culture plates. These samples were then mixed with an equal volume of hybridoma supernatant dilution containing more than a 100-fold excess of antibody above endpoint titers, as determined by 50 % plaque reduction in neutralization tests. After incubation for 1 h at 37 °C, HRT-18G cells were added, and the plates were maintained for 5 days at 37 °C. Wells were selected from the highest dilutions resulting in viral growth, and the mutant virus was plaque-purified twice.
RT-PCR and cDNA sequencing of escape mutants
Reverse transcription polymerase chain reaction (RT-PCR) was performed to amplify the full-length cDNA of the S gene of each of the escape mutants. Next, RNA was extracted from the virus culture using a High Pure Viral RNA Kit (Roche Diagnostics K.K., Tokyo, Japan). The oligonucleotide primers used for RT-PCR were designed from the nucleotide sequence of the Kakegawa strain (GenBank accession no. AB354579). The primers were BCoV-SF1, 5′-GTTATATTTTATGGTGGATAATG-3′ (nt 23581–23603, sense primer, start at position −60 of the S gene), and BCoV-SR1, 5′-TGGGTCTTATAATTAGCTTCACA-3′ (nt 27784–27806, antisense primer, start at position +74 of the S gene). RT-PCR was performed using a Titan One Tube RT-PCR Kit (Roche Diagnostics K.K.), followed by purification of the amplified DNA fragments using a QIAquick PCR purification kit (QIAGEN, Tokyo, Japan). The amplification fragments (4.2 kb in length) were subsequently used for sequencing.
The sequencing reaction was performed using a BigDye Terminator v 3.1 Cycle Sequencing Kit (Life Technologies, Tokyo, Japan). The sequencing primers were designed based on the sequence of the Kakegawa strain, in addition to the sequences of BCoV-SF1 and BCoV-SR1, as detailed in Table 1. Sequencing was performed by using an ABI3130 Genetic Analyzer (Life Technologies).
Ethics
The Animal Care and Use Committee of the National Institute of Animal Health approved all animal procedures prior to initiation of this study.
Results
Indirect immunofluorescence test
The hyperimmune rabbit sera against Kakegawa, 66/H, Ishikawa/2/99, Hokkaido/12/03, Kumamoto/1/07, Hokkaido/25/05, Hokkaido/40/08, and Hokkaido/45/08 strains reacted with all the viruses in the indirect immunofluorescence test (data not shown).
Cross-neutralization test
The virus strains Kakegawa, 66/H, Ishikawa/2/99, Hokkaido/12/03, Kumamoto/1/07, Hokkaido/25/05, Hokkaido/40/08, and Hokkaido/45/08 were compared for neutralizing antigenicity by cross-neutralization studies using antisera to these viruses (Table 2). All of the antisera neutralized the heterologous strains, whereas a certain antigenic difference was observed among groups. In particular, the antisera against the classical strains, such as Kakegawa and 66/H, belonging to group 1, showed lower R% values when compared to viruses belonging to the most recent predominant group (group 4).
Neutralizing antibody titers of bovine sera against viruses of each group
The bovine inoculated with the Kumamoto/1/07 strain showed seroconversion against all four groups of viruses (Table 3). However, when compared to antisera against the homologous virus, the neutralizing antibody titers were more than fourfold lower against group 1 and 2 viruses at 24 and 29 days after inoculation, respectively.
Neutralizing antigenicity of mAb 4H4 against viruses of each group
The neutralizing antibody titer of 4H4 in supernatants of hybridoma culture and ascites of mice against viruses of each group was examined by VN tests. The mAb 4H4 neutralized the homologous strain Kakegawa and 66/H strains belonging to group 1. The antibody titers to the other groups were significantly reduced (Table 4).
SDS-PAGE and western blotting
To identify the viral protein recognized by mAb 4H4, the binding of 4H4 in western blotting assays was examined. The assays showed that in the absence of 2-ME, the mAb 4H4 bound to a 105-kDa protein, which was identified as the S1 subunit of the S protein [34]. In the presence of 2-ME, the mAb 4H4 did not bind to any antigen (Fig. 1).
Generation and sequencing of neutralization-resistant mutants
The harvested viruses were plaque-purified, and 15 mutants were obtained. Sequencing of the S gene showed that all mutants possessed a single nucleotide substitution from T to C at position 851 of the S gene when compared to the Kakegawa strain. This substitution resulted in the change of a single codon from valine to alanine at amino acid position 284 of the S protein.
Discussion
The results of our phylogenetic analysis using a polymorphic region (aa 456–592) in the S gene of BCoV, Japanese BCoV isolates (1999–2008), and reference strains such as Mebus, Quebec, and Kakegawa demonstrated that these viruses clustered into four groups [14, 15]. Our previous study revealed that group 4 is the predominant lineage and has been widespread throughout Japan since 2005. Although these recent isolates (2009–2012) were still clustered into group 4 (data not shown), a novel group is certain to arise in the near future due to the progression of genetic divergence in the polymorphic region of the S gene. Although BCoV is thought to have only one serotype, its antigenicity may vary among several genetic groups; thus, we conducted a comparative analysis of antigenicity among isolates belonging to each group.
No substantial differences were observed among isolates in indirect immunofluorescence testing. However, cross-VN testing uncovered antigenic differences among the groups, although there was no clear indication of a distinct serotype. In particular, there was a marked difference between groups 1 and 4, suggesting that the genetic differences between these groups may have altered their antigenicity. Minor antigenic differences in VN test results have been observed among BCoV isolates in the United States [33] and Japan [7]. Furthermore, Canadian isolates collected between 1987 and 1990 have been classified into three antigenic subgroups on the basis of their reactivity with a panel of mAbs [23]. Hemagglutination-inhibition (HI) activity of sera against Mebus also revealed antigenic differences in field isolates in the United States, which were classified into three subgroups [16]. For the serological diagnosis of BCoV, reference strains such as Mebus or Kakegawa have been used in the hemagglutination inhibition (HI) test and VN test. However, considering that the antigenicity will continuously change, it may not be appropriate to use these strains. Indeed, when compared to antisera against the homologous virus, bovine sera inoculated with Kumamoto/1/07 virus showed more than a fourfold reduction in VN antibody titers against Kakegawa and Ishikawa/2/99 at 24 and 29 days after inoculation, respectively (Table 3). Furthermore, antigenic differences exist, even within group 4 strains (Table 2). In brief, the antigenic relationship (R) among the four strains allowed them to be divided into two groups: (1) Kumamoto/1/07 and Hokkaido/25/05 and (2) Hokkaido/40/08 and Hokkaido/45/08. This result suggested that antigenicity may vary, even within the same genetic group, and that neutralization antigenic sites may exist in a region other than the polymorphic region used in our phylogenetic study.
In our immunoblotting study, mAb 4H4 bound to the S1 subunit of the S protein in the absence of 2-ME, thereby revealing that 4H4 recognizes a conformational epitope of the virus (Fig. 1). All 15 of the neutralization-resistant mutants generated by co-culture with 4H4 possessed only a single nucleotide substitution at position 851 of the S gene when compared to the sequence of the Kakegawa strain (GenBank accession no. AB354579), resulting in the alteration of the S protein at amino acid position 284 from valine to alanine. Yoo et al. reported that mAbs against the Quebec strain belonging to group 1 recognized the conformational domain consisting of domain I (aa 351–403) and domain II (aa 517–621); further analysis using neutralization-resistant mutants revealed that the neutralization antigenic site was at aa 528 of the S protein in the polymorphic region [35]. The antigenic site at aa 284 in our study is not located in the polymorphic region (aa 456–592) nor in domain I or II; therefore, it has been suggested that this amino acid constitutes an alternate neutralization epitope compared with that of aa 528 of the Quebec strain. However, the Ishikawa/2/99, Hokkaido/12/03, and Kumamoto/1/07 strains, which were resistant to 4H4, possess a valine at aa 284, equivalent to the Kakegawa strain. This observation suggested that these viruses have amino acid alterations at the alternate site, which consists of the same neutralization epitope with aa 284, an alteration that might affect reactivity against 4H4.
In Japan, monovalent inactivated vaccine for adult cattle or calves (administered twice at one-month intervals), and a combined inactivated vaccine (including three types of rotavirus and Escherichia coli) for adult cattle (administered once at a half month before delivery) are used to prevent the development of calf diseases through colostrum. In the United States, a modified live vaccine combined with rotavirus is available in addition to inactivated vaccine. However, the preventive effects of these vaccines are uncertain [5]. Since BCoV infects and replicates in the intestinal and respiratory mucosa, it is vital to induce mucosal immunity for optimal protection. The aforementioned vaccine may be expected to induce antibodies in the blood, although the quantity that circulates to the intestinal tract is, at times, inadequate to prevent infection and replication of the virus. Therefore, the aim of vaccination for BCoV should be to prevent a severe disease requiring treatment and reduction in weight gain in infected animals [28]. These objectives may be accomplished by the selection of the optimal viral strain, whose antigenicity is close to that of the epidemic strain, as well as a vaccine program. Furthermore, BCoV causes three types of clinical symptoms, including CD, WD, and respiratory symptoms, although it is not yet apparent whether the existence of certain genetic markers correlates with the presence of different clinical symptoms. In the future, it will be important to use a viral strain for a vaccine that is protective against distinct clinical symptoms. Consequently, it is a significant task to conduct comparative analysis of antigenicity of field isolates in each country, as well as a phylogenetic study that provides us with all of the necessary epidemiological information about BCoV.
References
Archetti I, Horsfall FL Jr (1950) Persistent antigenic variation of influenza A viruses after incomplete neutralization in ovo with heterologous immune serum. J Exp Med 92:441–462
Ballesteros ML, Sanchez CM, Enjuanes L (1997) Two amino acid changes at the N-terminus of transmissible gastroenteritis coronavirus spike protein result in the loss of enteric tropism. Virology 227:378–388
Bjerrum OJ, Schafer-Nielsen C (1986) Buffer systems and transfer parameters for semidry electroblotting with a horizontal apparatus. In Electrophoresis ‘86: Proceeding of the fifth meeting of the international electrophoresis society, pp 315–327
Brandao PE, Gregori F, Richtzenhain LJ, Rosales CA, Villarreal LY, Jerez JA (2006) Molecular analysis of Brazilian strains of bovine coronavirus (BCoV) reveals a deletion within the hypervariable region of the S1 subunit of the spike glycoprotein also found in human coronavirus OC43. Arch Virol 151:1735–1748
Collins JK, Riegel CA, Olson JD, Fountain A (1987) Shedding of enteric coronavirus in adult cattle. Am J Vet Res 48:361–365
el-Ghorr AA, Snodgrass DR, Scott FM, Campbell I (1989) A serological comparison of bovine coronavirus strains. Arch Virol 104:241–248
Fukutomi T, Tsunemitsu H, Akashi H (1999) Detection of bovine coronaviruses from adult cows with epizootic diarrhea and their antigenic and biological diversities. Arch Virol 144:997–1006
Godet M, Grosclaude J, Delmas B, Laude H (1994) Major receptor-binding and neutralization determinants are located within the same domain of the transmissible gastroenteritis virus (coronavirus) spike protein. J Virol 68:8008–8016
Guy JS, Breslin JJ, Breuhaus B, Vivrette S, Smith LG (2000) Characterization of a coronavirus isolated from a diarrheic foal. J Clin Microbiol 38:4523–4526
Hasoksuz M, Sreevatsan S, Cho KO, Hoet AE, Saif LJ (2002) Molecular analysis of the S1 subunit of the spike glycoprotein of respiratory and enteric bovine coronavirus isolates. Virus Res 84:101–109
Hingley ST, Gombold JL, Lavi E, Weiss SR (1994) MHV-A59 fusion mutants are attenuated and display altered hepatotropism. Virology 200:1–10
Hussain KA, Storz J, Kousoulas KG (1991) Comparison of bovine coronavirus (BCV) antigens: monoclonal antibodies to the spike glycoprotein distinguish between vaccine and wild-type strains. Virology 183:442–445
Jeong JH, Kim GY, Yoon SS, Park SJ, Kim YJ, Sung CM, Shin SS, Lee BJ, Kang MI, Park NY, Koh HB, Cho KO (2005) Molecular analysis of S gene of spike glycoprotein of winter dysentery bovine coronavirus circulated in Korea during 2002–2003. Virus Res 108:207–212
Kanno T, Hatama S, Ishihara R, Uchida I (2007) Molecular analysis of the S glycoprotein gene of bovine coronaviruses isolated in Japan from 1999 to 2006. J Gen Virol 88:1218–1224
Kanno T, Kamiyoshi T, Ishihara R, Hatama S, Uchida I (2009) Phylogenetic studies of bovine coronaviruses isolated in Japan. J Vet Med Sci 71:83–86
Kapil S, Richardson KL, Maag TR, Goyal SM (1999) Characterization of bovine coronavirus isolates/from eight different states in the USA. Vet Microbiol 67:221–230
Ko CK, Kang MI, Lim GK, Kim GY, Yoon SS, Park JT, Jeong C, Park SH, Park SJ, Kim YJ, Jeong JH, Kim SK, Park SI, Kim HH, Kim KY, Cho KO (2006) Molecular characterization of HE, M, and E genes of winter dysentery bovine coronavirus circulated in Korea during 2002–2003. Virus Genes 32:129–136
Kubo H, Yamada YK, Taguchi F (1994) Localization of neutralizing epitopes and the receptor-binding site within the amino-terminal 330 amino acids of the murine coronavirus spike protein. J Virol 68:5403–5410
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lai MM, Cavanagh D (1997) The molecular biology of coronaviruses. Adv Virus Res 48:1–100
Liu L, Hagglund S, Hakhverdyan M, Alenius S, Larsen LE, Belak S (2006) Molecular epidemiology of bovine coronavirus on the basis of comparative analyses of the S gene. J Clin Microbiol 44:957–960
Mebus CA, Stair EL, Rhodes MB, Twiehaus MJ (1973) Neonatal calf diarrhea: propagation, attenuation, and characteristics of a coronavirus-like agent. Am J Vet Res 34:145–150
Michaud L, Dea S (1993) Characterization of monoclonal antibodies to bovine enteric coronavirus and antigenic variability among Quebec isolates. Arch Virol 131:455–465
Oue Y, Ishihara R, Edamatsu H, Morita Y, Yoshida M, Yoshima M, Hatama S, Murakami K, Kanno T (2011) Isolation of an equine coronavirus from adult horses with pyrogenic and enteric disease and its antigenic and genomic characterization in comparison with the NC99 strain. Vet Microbiol 150:41–48
Park SJ, Lim GK, Park SI, Kim HH, Koh HB, Cho KO (2007) Detection and molecular characterization of calf diarrhoea bovine coronaviruses circulating in South Korea during 2004–2005. Zoonoses Public Health 54:223–230
Rekik MR, Dea S (1994) Comparative sequence analysis of a polymorphic region of the spike glycoprotein S1 subunit of enteric bovine coronavirus isolates. Arch Virol 135:319–331
Saif LJ, Brock KV, Redman DR, Kohler EM (1991) Winter dysentery in dairy herds: electron microscopic and serological evidence for an association with coronavirus infection. Vet Rec 128:447–449
Saif LJ (2010) Bovine respiratory coronavirus. Vet Clin N Am Food Anim Pract 26:349–364
Sato M, Akashi H, Hirai S, Kimura Y, Takahashi M (1991) Production of monoclonal antibodies against the Kakegawa strain of bovine coronavirus and their characterization. J Vet Med Sci 53:147–148
Schultze B, Gross HJ, Brossmer R, Herrler G (1991) The S protein of bovine coronavirus is a hemagglutinin recognizing 9-O-acetylated sialic acid as a receptor determinant. J Virol 65:6232–6237
Storz J, Stine L, Liem A, Anderson GA (1996) Coronavirus isolation from nasal swab samples in cattle with signs of respiratory tract disease after shipping. J Am Vet Med Assoc 208:1452–1455
Takase-Yoden S, Kikuchi T, Siddell SG, Taguchi F (1991) Localization of major neutralizing epitopes on the S1 polypeptide of the murine coronavirus peplomer glycoprotein. Virus Res 18:99–107
Tsunemitsu H, el-Kanawati ZR, Smith DR, Saif LJ, Reed HH (1995) Isolation of coronaviruses antigenically indistinguishable from bovine coronavirus from wild ruminants with diarrhea. J Clin Microbiol 33:3264–3269
Vautherot JF, Laporte J, Boireau P (1992) Bovine coronavirus spike glycoprotein: localization of an immunodominant region at the amino-terminal end of S2. J Gen Virol 73(Pt 12):3289–3294
Yoo D, Deregt D (2001) A single amino acid change within antigenic domain II of the spike protein of bovine coronavirus confers resistance to virus neutralization. Clin Diagn Lab Immunol 8:297–302
Acknowledgments
This study was supported by a Grant-in-Aid for Specially Weighted Research, National Institute of Animal Health, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kanno, T., Ishihara, R., Hatama, S. et al. Antigenic variation among recent Japanese isolates of bovine coronaviruses belonging to phylogenetically distinct genetic groups. Arch Virol 158, 1047–1053 (2013). https://doi.org/10.1007/s00705-012-1587-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-012-1587-1