Abstract
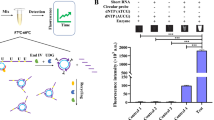
MicroRNAs (miRNAs), found in blood and body fluids, have emerged as potential non-invasive biomarkers for disease and injury. miRNAs are quantitatively evaluated using typical RNA analysis methods such as the quantitative reverse transcription polymerase chain reaction, microarrays, and Northern blot, all of which require complex procedures and expensive reagents. To utilize miRNAs as practical biomarkers, it will be helpful to develop simple and user-friendly sensors. In this study, a paper-based miRNA sensor was developed by combining two methods: (1) target-recycled DNAzyme (Dz) amplification and (2) graphene oxide-assisted Dz blotting on paper. The Dz spots on paper caused a miRNA-dependent color change in presence of colorimetric reagents and facilitated the quantification of absolute amount of the target miRNA, irrespective of the volume, with high reproducibility. This approach is technologically straightforward and enables quantification of as low as 7.75 fmol miRNA using a portable smartphone.
Graphical abstract







Similar content being viewed by others
Data availability
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
Abbreviations
- ABTS :
-
2,2-Azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) diammonium salt
- Dz :
-
DNAzyme
- FAM-Dz :
-
Fluorescein-labeled Dz strand
- FRET :
-
Fluorescence resonance energy transfer
- G :
-
Guanine
- GO :
-
Graphene oxide
- LOD :
-
Limit of detection
- miR-122 :
-
MiRNA-122
- miRNA :
-
MicroRNA
- S :
-
Slope
- SEM :
-
Standard error of the mean
- SD :
-
Standard deviation
- ssDNA :
-
Single-stranded DNA
- Sup :
-
Supernatant
- TMSD :
-
Toehold-mediated strand displacement
References
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114. https://doi.org/10.1038/nrg2290
Hwang HW, Mendell JT (2006) MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer 94:776–780. https://doi.org/10.1038/sj.bjc.6603023
Jovanovic M, Hengartner MO (2006) miRNAs and apoptosis: RNAs to die for. Oncogene 25:6176–6187. https://doi.org/10.1038/sj.onc.1209912
Inui M, Martello G, Piccolo S (2010) MicroRNA control of signal transduction. Nat Rev Mol Cell Biol 11:252–263. https://doi.org/10.1038/nrm2868
Sayed D, Abdellatif M (2011) MicroRNAs in development and disease. Physiol Rev 91:827–887. https://doi.org/10.1152/physrev.00006.2010
Hayes J, Peruzzi PP, Lawler S (2014) MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med 20:460–469. https://doi.org/10.1016/j.molmed.2014.06.005
Reddy KB (2015) MicroRNA (miRNA) in cancer. Cancer Cell Int 15:38. https://doi.org/10.1186/s12935-015-0185-1
Skalsky RL, Cullen BR (2010) Viruses, microRNAs, and host interactions. Annu Rev Microbiol 64:123–141. https://doi.org/10.1146/annurev.micro.112408.134243
Jopling C (2012) Liver-specific microRNA-122: biogenesis and function. RNA Biol 9:137–142. https://doi.org/10.4161/rna.18827
Kosaka N, Iguchi H, Ochiya T (2010) Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci 101:2087–2092. https://doi.org/10.1111/j.1349-7006.2010.01650.x
Bala S, Petrasek J, Mundkur S, Catalano D, Levin I, Ward J, Alao H, Kodys K, Szabo G (2012) Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology 56:1946–1957. https://doi.org/10.1002/hep.25873
Sharapova T, Devanarayan V, LeRoy B, Liguori MJ, Blomme E, Buck W, Maher J (2016) Evaluation of miR-122 as a serum biomarker for hepatotoxicity in investigative rat toxicology studies. Vet Pathol 53:211–221. https://doi.org/10.1177/0300985815591076
Schmittgen TD, Lee EJ, Jiang J, Sarkar A, Yang L, Elton TS, Chen C (2008) Real-time PCR quantification of precursor and mature microRNA. Methods 44:31–38. https://doi.org/10.1016/j.ymeth.2007.09.006
Várallyay E, Burgyán J, Havelda Z (2008) MicroRNA detection by Northern blotting using locked nucleic acid probes. Nat Protoc 3:190–196. https://doi.org/10.1038/nprot.2007.528
Degliangeli F, Pompa PP, Fiammengo R (2014) Nanotechnology-based strategies for the detection and quantification of microRNA. Chemistry 20:9476–9492. https://doi.org/10.1002/chem.201402649
Chandrasekaran AR, Punnoose JA, Zhou L, Dey P, Dey BK, Halvorsen K (2019) DNA Nanotechnology approaches for microRNA detection and diagnosis. Nucleic Acids Res 47:10489–10505. https://doi.org/10.1093/nar/gkz580
Yang L, Liu C, Ren W, Li Z (2012) Graphene surface-anchored fluorescence sensor for sensitive detection of microRNA coupled with enzyme-free signal amplification of hybridization chain reaction. ACS Appl Mater Interfaces 4:6450–6453. https://doi.org/10.1021/am302268t
Hakimian F, Ghourchian H, Hashemi AS, Arastoo MR, Behnam Rad M (2018) Ultrasensitive optical biosensor for detection of miRNA-155 using positively charged Au nanoparticles. Sci Rep 8:2943. https://doi.org/10.1038/s41598-018-20229-z
Jebelli A, Oroojalian F, Fathi F, Mokhtarzadeh A, Guardia M (2020) Recent advances in surface plasmon resonance biosensors for microRNAs detection. Biosens Bioelectron 169:112599. https://doi.org/10.1016/j.bios.2020.112599
Shabaninejad Z, Yousefi F, Movahedpour A, Ghasemi Y, Dokanehiifard S, Rezaei S, Aryan R, Savardashtaki A, Mirzaei H (2019) Electrochemical-based biosensors for microRNA detection: nanotechnology comes into view. Anal Biochem 581:113349. https://doi.org/10.1016/j.ab.2019.113349
Zhou X, Cao P, Zhu Y, Lu W, Gu N, Mao C (2015) Phage-mediated counting by the naked eye of miRNA molecules at attomolar concentrations in a Petri dish. Nat Mater 14:1058–1064. https://doi.org/10.1038/nmat4377
Persano S, Guevara ML, Wolfram J, Blanco E, Shen H, Ferrari M, Pompa PP (2016) Label-free isothermal amplification assay for specific and highly sensitive colorimetric miRNA detection. ACS Omega 1:448–455. https://doi.org/10.1021/acsomega.6b00109
Mundinamani S (2020) Large area, multilayer graphene films as a flexible electronic material. ACS Omega 5:17479–17485. https://doi.org/10.1021/acsomega.0c01982
Perreault F, Fonseca de Faria A, Elimelech M (2015) Environmental applications of graphene-based nanomaterials. Chem Soc Rev 44:5861–5896. https://doi.org/10.1039/c5cs00021a
Li X, Zhi L (2018) Graphene hybridization for energy storage applications. Chem Soc Rev 47:3189–3216. https://doi.org/10.1039/c7cs00871f
Reina G, González-Domínguez JM, Criado A, Vázquez E, Bianco A, Prato M (2017) Promises, facts and challenges for graphene in biomedical applications. Chem Soc Rev 4400(46):4400–4416. https://doi.org/10.1039/c7cs00363c
Chen D, Feng H, Li J (2012) Graphene oxide: preparation, functionalization, and electrochemical applications. Chem Rev 112:6027–6053. https://doi.org/10.1021/cr300115g
Mouhat F, Coudert FX, Bocquet ML (2020) Structure and chemistry of graphene oxide in liquid water from first principles. Nat Commun 11:1566. https://doi.org/10.1038/s41467-020-15381-y
Paul T, Bera SC, Agnihotri N, Mishra PP (2016) Single-molecule FRET studies of the hybridization mechanism during noncovalent adsorption and desorption of DNA on graphene oxide. J Phys Chem B 120:11628–11636. https://doi.org/10.1021/acs.jpcb.6b06017
Li F, Pei H, Wang L, Lu J, Gao J, Jiang B, Zhao X, Fan C (2013) Nanomaterial-based fluorescent DNA analysis: a comparative study of the quenching effects of graphene oxide, carbon nanotubes, and gold nanoparticles. Adv Func Mat 23:4140. https://doi.org/10.1002/adfm.201203816
Georgakilas V, Perman JA, Tucek J, Zboril R (2015) Broad family of carbon nanoallotropes: classification, chemistry, and applications of fullerenes, carbon dots, nanotubes, graphene, nanodiamonds, and combined superstructures. Chem Rev 115:4744. https://doi.org/10.1021/cr500304f
Hwang HS, Jeong JW, Kim YA, Chang M (2020) Carbon nanomaterials as versatile platforms for biosensing applications. Micromachines 11:814. https://doi.org/10.3390/mi11090814
Lee J, Kim Y-k, Lee S, Yoon S, Kim W-k (2019) Graphene oxide-based NET strategy for enhanced colorimetric sensing of miRNA. Sens Actuators B 282:861–867. https://doi.org/10.1016/j.snb.2018.11.149
Li W, Li Y, Liu Z, Lin B, Yi H, Xu F, Nie Z, Yao S (2016) Insight into G-quadruplex-hemin DNAzyme/RNAzyme: adjacent adenine as the intramolecular species for remarkable enhancement of enzymatic activity. Nucleic Acids Res 44:7373–7384. https://doi.org/10.1093/nar/gkw634
Zhang DY, Winfree E (2009) Control of DNA strand displacement kinetics using toehold exchange. J Am Chem Soc 131:17303–17314. https://doi.org/10.1021/ja906987s
Kim J, Cote LJ, Kim F, Huang J (2010) Visualizing graphene based sheets by fluorescence quenching microscopy. J Am Chem Soc 132:260–267. https://doi.org/10.1021/ja906730d
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2020R1F1A1072921) and the Korea Institute of Toxicology (Grant No. KK-2101–01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J., Na, HK., Lee, S. et al. Advanced graphene oxide-based paper sensor for colorimetric detection of miRNA. Microchim Acta 189, 35 (2022). https://doi.org/10.1007/s00604-021-05140-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-021-05140-1