Abstract
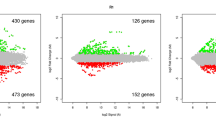
The shoots of young conifer trees represent an interesting model to study the development and growth of conifers from meristematic cells in the shoot apex to differentiated tissues at the shoot base. In this work, microarray analysis was used to monitor contrasting patterns of gene expression between the apex and the base of maritime pine shoots. A group of differentially expressed genes were selected and validated by examining their relative expression levels in different sections along the stem, from the top to the bottom. After validation of the microarray data, additional gene expression analyses were also performed in the shoots of young maritime pine trees exposed to different levels of ammonium nutrition. Our results show that the apex of maritime pine trees is extremely sensitive to conditions of ammonium excess or deficiency, as revealed by the observed changes in the expression of stress-responsive genes. This new knowledge may be used to precocious detection of early symptoms of nitrogen nutritional stresses, thereby increasing survival and growth rates of young trees in managed forests.





Similar content being viewed by others
References
Alonso P, Cortizo M, Cantón FR, Fernández B, Rodríguez A, Cánovas FM, Ordás R (2007) Identification of genes differentially expressed during adventitious shoot induction in Pinus pinea L. cotyledons by subtractive PCR. Tree Physiol 27:1721–1730
Andersen CL, Jensen JL, Orntof TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64:5245–5250
Aranda I, Alía R, Ortega U, Dantas ÂK, Majada J (2010) Intra-specific variability in biomass partitioning and carbon isotopic discrimination under moderate drought stress in seedlings from four Pinus pinaster populations. Tree Genet Genomes 6:169–178
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate—a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300
Canales J, Flores-Monterrosso A, Rueda-López M, Avila C, Cánovas FM (2010) Identification of genes regulated by ammonium availability in the roots of maritime pine trees. Amino Acids 39:991–1001
Canales J, Avila C, Cánovas FM (2011) A maritime pine antimicrobial peptide involved in ammonium nutrition. Plant Cell Environ 34:1443–1453
Cañas RA, de la Torre F, Cánovas FM, Cantón FR (2006) High levels of asparagine synthetase in hypocotyls of pine seedlings reveal an essential role of the enzyme in re-allocation of seed-stored nitrogen. Planta 224:83–95
Cánovas FM, Avila C, Cantón FR, Cañas R, de la Torre F (2007) Ammonium assimilation and amino acid metabolism in conifers. J Exp Bot 58:2307–2318 (Special Issue Nitrogen Nutrition)
Cánovas FM, Canales J, Flores-Monterrosso A, Rueda M, Avila C (2009) Genomics approaches to study ammonium nutrition and amino acid biosynthesis in conifers. Amino Acids 37(Suppl 1):45–46
Cantón FR, Suárez MF, Josè-Estanyol M, Cánovas FM (1999) Expression analysis of a cytosolic glutamine synthetase gene in cotyledons of Scots pine seedlings: developmental, light regulation and spatial distribution of specific transcripts. Plant Mol Biol 40:623–634
Cantón FR, Le Provost G, García V, Barré A, Frigerio J-M, Fevereiro P, Avila C, Mouret J-F, de Daruvar A, Cánovas FM, Plomion C (2003) Transcriptome analysis of wood formation in maritime pine. In: Ritter E, Espinel S, Barredo Y et al (eds) Sustainable forestry. Woods products and biotecnology. DFA-AFA Press, Vitoria-Gasteiz, pp 333–348
Cato S, McMillan L, Donaldson L, Richardson T, Echt C, Richard Gardner R (2006) Wood formation from the base to the crown in Pinus radiata: gradients of tracheid wall thickness, wood density, radial growth rate and gene expression. Plant Mol Biol 60:565–581
Driscoll CT, Whitall D, Aber J, Boyer E, Castro M, Cronan C, Goodale CL, Groffman P, Charles Hopkinson C, Lambert K, Gregory Lawrence G, Ollinger S (2003) Nitrogen pollution in the Northeastern United States: sources, effects, and management options. BioScience 53:357–374
Farjon A (2010) A handbook of the World’s conifers. E.J. Brill, Leiden/Boston
Friedmann M, Ralph SG, Aeschliman D, Zhuang J, Ritland K, Ellis BE, Bohlmann J, Douglas CJ (2007) Microarray gene expression profiling of developmental transitions in Sitka spruce (Picea sitchensis) apical shoots. J Exp Bot 58:593–614
Galiano L, Martínez-Vilalta J, Lloret F (2010) Drought-induced multifactor decline of Scots pine in the pyrenees and potential vegetation change by the expansion of co-occurring oak species. Ecosystems 13:978–991
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JY, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Hornung M, Langan SJ (1999) Nitrogen deposition: sources, impacts and responses in natural and semi-natural ecosystems. In: Langan SJ (ed) The impact of nitrogen deposition on natural, semi-natural ecosystems. Kluwer AcademicPublishers, Dordrecht, pp 1–13
Kronzucker HJ, Siddiqi MY, Glass ADM (1997) Conifer root discrimination against soil nitrate and the ecology of forest succession. Nature 385:59–61
Mohr H (1986) Die Erforschung der neuartigen Waldschäden. BIUZ 16:83–89
Nihlgard B (1985) The ammonium hypothesis: an additional explanation for the forest dieback in Europe. Ambio 14:2–8
Lea PJ, Morot-Gaudry JF (2001) Plant nitrogen. Springer, Berlin
Lea PJ, Sodek L, Parry MAJ, Shewry PR, Halford NG (2007) Asparagine in plants. Ann Appl Biol 150:1–26
Liao Z, Chen M, Guo L, Gong Y, Tang F, Sun X, Tang K (2004) Rapid isolation of high-quality total RNA from Taxus and Ginkgo. Prep Biochem Biotechnol 34:209–214
Martin F, Kohler A, Duplessis S (2007) Living in harmony in the wood underground: ectomycorrhizal genomics. Curr Opin Plant Biol 10:204–210
Ohlünd J, Näsholm T (2004) Regulation of organic and inorganic nitrogen uptake in Scots pine (Pinus sylvestris) seedlings. Tree Physiol 24:1397–1402
Paiva JAP, Garcés M, Alves A, Garnier-Géré P, Rodrigues JC, Lalanne C, Porcon S, Le Provost G, da Silva Perez D, Brach J, Frigerio J-M, Claverol S, Barré A, Fevereiro P, Plomion C (2008) Molecular and phenotypic profiling from the base to the crown in maritime pine wood-forming tissues. New Phytol 178:283–301
Pavy N, Boyle B, Nelson C, Paule C, Giguere I, Caron S, Parsons LS, Dallaire N, Bedon F, Berube H, Cooke J, Mackay J (2008) Identification of conserved core xylem gene sets: conifer cDNA microarray development, transcript profiling and computational analyses. New Phytol 180:766–786
Ralph SG, Yueh H, Friedmann M, Aeschliman D, Zeznik JA, Nelson CC, Butterfield YSN, Kirkpatrick R, Liu J, Jones SJM, Marra MA, Douglas CJ, Ritland K, Bohlmann J (2006) Conifer defence against insects: microarray gene expression profiling of Sitka spruce (Picea sitchensis) induced by mechanical wounding or feeding by spruce budworms (Choristoneura occidentalis) or white pine weevils (Pissodes strobi) reveals large-scale changes of the host transcriptome. Plant Cell Environ 29:1545–1570
Regan S, Bourquin V, Tuominen H, Sundberg B (1999) Accurate and high resolution in situ hybridization analysis of gene expression in secondary stem tissues. Plant J 19:363–369
Ritchie ME, Silver J, Oshlack A, Holmes M, Diyagama D, Holloway A, Smyth GK (2007) A comparison of background correction methods for two-colour microarrays. Bioinformatics 23:2700–2707
Ruijter JM, Ramakers C, Hoogaars WM, Karlen Y, Bakker O, van den Hoff MJ, Moorman AF (2009) Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucl Acids Res 37:e45
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:Article3
Smyth GK (2005) Limma: linear models for microarray data. In: Gentleman R, Carey V, Irizarry R, Huber W, Dudoit S (eds) Bioinformatics and computational biology solutions using R and bioconductor. Springer, New York, pp 397–420
Suárez MF, Avila C, Gallardo F, Cantón FR, García-Gutiérrez A, Claros MG, Cánovas FM (2002) Molecular and enzymatic analysis of ammonium assimilation in woody plants. J Exp Bot 53:891–904
Untergasser A, Nijveen H, Rao X, Bisseling T, Geurts R, Leunissen JA (2007) Primer3Plus, an enhanced web interface to Primer3. Nucl Acids Res 35 (Web Server issue):W71–W74
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:34.1–34.11
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J, Speed TP (2002) Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucl Acids Res 30:e15
Yang SH, van Zyl L, No EG, Loopstra CA (2004) Microarray analysis of genes preferentially expressed in differentiating xylem of loblolly pine (Pinus taeda). Plant Sci 166:1185–1195
Acknowledgments
This research was supported by grants from the Junta de Andalucía (P05-AGR663) and the Ministerio de Ciencia e Innovación (BIO2009-07490) to F.M.C and the research group BIO-114. We would like to thank Noé Fernández-Pozo for technical assistance. We appreciate the efforts of the two anonymous reviewers and their valuable comments and suggestions for improving the manuscript. This work is part of the activities of the Andalusian Platform for Genomics, Proteomics and Bioinformatics.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Klimaszewska.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Canales, J., Ávila, C., Cantón, F.R. et al. Gene expression profiling in the stem of young maritime pine trees: detection of ammonium stress-responsive genes in the apex. Trees 26, 609–619 (2012). https://doi.org/10.1007/s00468-011-0625-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-011-0625-z