Abstract
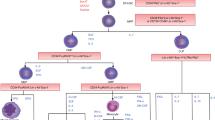
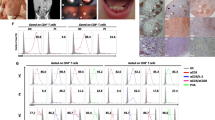
Osteoclasts (OCLs) are multinucleated giant cells and are formed by the fusion of mononuclear progenitors of monocyte/macrophage lineage. It is known that macrophages derived from different genetic backgrounds exhibit quite distinct characteristics of immune responses. However, it is unknown whether OCLs from different genetic backgrounds show distinct characteristics. In this study, we showed that bone-marrow macrophages (BMMs) derived from C57BL/6, BALB/c and ddY mice exhibited considerably distinct morphological characteristics and cell differentiation into OCLs. The differentiation of BMMs into OCLs was comparatively quicker in the C57BL/6 and ddY mice, while that of BALB/c mice was rather slow. Morphologically, ddY OCLs showed a giant cell with a round shape, C57BL/6 OCLs were of a moderate size with many protrusions and BALB/c OCLs had the smallest size with fewer nuclei. The intracellular signaling of differentiation and expression levels of marker proteins of OCLs were different in the respective strains. Treatment of BMMs from the three different strains with the reducing agent N-acetylcysteine (NAC) or with the oxidation agent hydrogen peroxide (H2O2) induced changes in the shape and sizes of the cells and caused distinct patterns of cell differentiation and survival. Thus, genetic backgrounds and redox conditions regulate the morphological characteristics and cell differentiation of OCLs.












Similar content being viewed by others
Abbreviations
- OCLs:
-
Osteoclasts
- RANKL:
-
Receptor activator of NF-κB ligand
- TRAP:
-
Tartrate-resistant acid phosphatase
- BMMs:
-
Bone-marrow macrophages
- NAC:
-
N-acetylcysteine
- H2O2 :
-
Hydrogen peroxide
- M-CSF:
-
Macrophage-colony-stimulating factor
- NFATc1:
-
Nuclear factor of activated T cells cytoplasmic-1
- DC-STAM:
-
Dendritic cell-specific transmembrane protein
- OC-STAMP:
-
Osteoclast stimulatory transmembrane protein
- MMP:
-
Matrix metalloproteinase
- TRAF6:
-
TNF receptor associated factor-6
References
Autenrieth IB, Beer M, Bohn E, Kaufmann SH, Heesemann J (1994) Immune responses to Yersinia enterocolitica in susceptible BALB/c and resistant C57BL/6 mice: an essential role for gamma interferon. Infect Immun 62:2590–2599
Baker PJ, Dixon M, Roopenian DC (2000) Genetic control of susceptibility to Porphyromonas gingivalis-induced alveolar bone loss in mice. Infect Immun 68:5864–5868
Boyle WJ, Simonet WS, Lacey DL (2003) Osteoclast differentiation and activation. Nature 423:337–342
Brett SJ, Butler R (1986) Resistance to Mycobacterium lepraemurium is correlated with the capacity to generate macrophage activating factor(s) in response to mycobacterial antigens in vitro. Immunology 59:339–345
Choi HG, Kim JM, Kim BJ, Yoo YJ, Cha JH (2007) Mouse strain-dependent osteoclastogenesis in response to lipopolysaccharide. J Microbiol 45:566–571
Delaisse JM, Andersen TL, Engsig MT, Henriksen K, Troen T, Blavier L (2003) Matrix metalloproteinases (MMP) and cathepsin K contribute differently to osteoclastic activities. Microsc Res Tech 61:504–513
Fairweather D, Cihakova D (2009) Alternatively activated macrophages in infection and autoimmunity. J Autoimmun 33:222–230
Gerstenfeld LC, McLean J, Healey DS, Stapleton SN, Silkman LJ, Price C, Jepsen KJ (2010) Genetic variation in the structural pattern of osteoclast activity during post-natal growth of mouse femora. Bone 46:1546–1554
Guler ML, Gorham JD, Hsieh CS, Mackey AJ, Steen RG, Dietrich WF, Murphy KM (1996) Genetic susceptibility to Leishmania: IL-12 responsiveness in TH1 cell development. Science 271:984–987
Ha H, Kwak HB, Lee SW, Jin HM, Kim HM, Kim HH, Lee ZH (2004) Reactive oxygen species mediate RANK signaling in osteoclasts. Exp Cell Res 301:119–127
Hoft DF, Lynch RG, Kirchhoff LV (1993) Kinetic analysis of antigen-specific immune responses in resistant and susceptible mice during infection with Trypanosoma cruzi. J Immunol 151:7038–7047
Hotokezaka H, Sakai E, Kanaoka K, Saito K, Matsuo K, Kitaura H, Yoshida N, Nakayama K (2002) U0126 and PD98059, specific inhibitors of MEK, accelerate differentiation of RAW264.7 cells into osteoclast-like cells. J Biol Chem 277:47366–47372
Hu JP, Nishishita K, Sakai E, Yoshida H, Kato Y, Tsukuba T, Okamoto K (2008) Berberine inhibits RANKL-induced osteoclast formation and survival through suppressing the NF-kappaB and Akt pathways. Eur J Pharmacol 580:70–79
Kamiya T, Kobayashi Y, Kanaoka K, Nakashima T, Kato Y, Mizuno A, Sakai H (1998) Fluorescence microscopic demonstration of cathepsin K activity as the major lysosomal cysteine proteinase in osteoclasts. J Biochem 123:752–759
Kim MS, Yang YM, Son A, Tian YS, Lee SI, Kang SW, Muallem S, Shin DM (2010) RANKL-mediated reactive oxygen species pathway that induces long lasting Ca2+ oscillations essential for osteoclastogenesis. J Biol Chem 285:6913–6921
Kukita T, Wada N, Kukita A, Kakimoto T, Sandra F, Toh K, Nagata K, Iijima T, Horiuchi M, Matsusaki H, Hieshima K, Yoshie O, Nomiyama H (2004) RANKL-induced DC-STAMP is essential for osteoclastogenesis. J Exp Med 200:941–946
Kuroda E, Noguchi J, Doi T, Uematsu S, Akira S, Yamashita U (2007) IL-3 is an important differentiation factor for the development of prostaglandin E2-producing macrophages between C57BL/6 and BALB/c mice. Eur J Immunol 37:2185–2195
Lacey DL, Timms E, Tan HL, Kelley MJ, Dunstan CR, Burgess T, Elliott R, Colombero A, Elliott G, Scully S, Hsu H, Sullivan J, Hawkins N, Davy E, Capparelli C, Eli A, Qian YX, Kaufman S, Sarosi I, Shalhoub V, Senaldi G, Guo J, Delaney J, Boyle WJ (1998) Osteoprotegerin ligand is a cytokine that regulates osteoclast differentiation and activation. Cell 93:165–176
Launois P, Maillard I, Pingel S, Swihart KG, Xenarios I, Acha-Orbea H, Diggelmann H, Locksley RM, MacDonald HR, Louis JA (1997) IL-4 rapidly produced by V beta 4 V alpha 8 CD4+ T cells instructs Th2 development and susceptibility to Leishmania major in BALB/c mice. Immunity 6:541–549
Leakey AK, Ulett GC, Hirst RG (1998) BALB/c and C57Bl/6 mice infected with virulent Burkholderia pseudomallei provide contrasting animal models for the acute and chronic forms of human melioidosis. Microb Pathog 24:269–275
Lee NK, Choi YG, Baik JY, Han SY, Jeong DW, Bae YS, Kim N, Lee SY (2005) A crucial role for reactive oxygen species in RANKL-induced osteoclast differentiation. Blood 106:852–859
Linkhart TA, Linkhart SG, Kodama Y, Farley JR, Dimai HP, Wright KR, Wergedal JE, Sheng M, Beamer WG, Donahue LR, Rosen CJ, Baylink DJ (1999) Osteoclast formation in bone marrow cultures from two inbred strains of mice with different bone densities. J Bone Miner Res 14:39–46
Lomaga MA, Yeh WC, Sarosi I, Duncan GS, Furlonger C, Ho A, Morony S, Capparelli C, Van G, Kaufman S, van der Heiden A, Itie A, Wakeham A, Khoo W, Sasaki T, Cao Z, Penninger JM, Paige CJ, Lacey DL, Dunstan CR, Boyle WJ, Goeddel DV, Mak TW (1999) TRAF6 deficiency results in osteopetrosis and defective interleukin-1, CD40, and LPS signaling. Genes Dev 13:1015–1024
Mantovani A, Sica A, Locati M (2007) New vistas on macrophage differentiation and activation. Eur J Immunol 37:14–16
Mills CD, Kincaid K, Alt JM, Heilman MJ, Hill AM (2000) M-1/M-2 macrophages and the Th1/Th2 paradigm. J Immunol 164:6166–6173
Murata Y, Shimamura T, Hamuro J (2002) The polarization of T(h)1/T(h)2 balance is dependent on the intracellular thiol redox status of macrophages due to the distinctive cytokine production. Int Immunol 14:201–212
Raggatt LJ, Partridge NC (2010) Cellular and molecular mechanisms of bone remodeling. J Biol Chem 285:25103–25108
Rho J, Takami M, Choi Y (2004) Osteoimmunology: interactions of the immune and skeletal systems. Mol Cells 17:1–9
Rivera J, Tessarollo L (2008) Genetic background and the dilemma of translating mouse studies to humans. Immunity 28:1–4
Suda T, Takahashi N, Martin TJ (1992) Modulation of osteoclast differentiation. Endocr Rev 13:66–80
Suda T, Takahashi N, Udagawa N, Jimi E, Gillespie MT, Martin TJ (1999) Modulation of osteoclast differentiation and function by the new members of the tumor necrosis factor receptor and ligand families. Endocr Rev 20:345–357
Takayanagi H (2009) Osteoimmunology and the effects of the immune system on bone. Nat Rev Rheumatol 5:667–676
Takayanagi H, Kim S, Koga T, Nishina H, Isshiki M, Yoshida H, Saiura A, Isobe M, Yokochi T, Inoue J, Wagner EF, Mak TW, Kodama T, Taniguchi T (2002) Induction and activation of the transcription factor NFATc1 (NFAT2) integrate RANKL signaling in terminal differentiation of osteoclasts. Dev Cell 3:889–901
Tanaka S, Wakeyama H, Akiyama T, Takahashi K, Amano H, Nakayama KI, Nakamura K (2010) Regulation of osteoclast apoptosis by bcl-2 family protein bim and caspase-3. Adv Exp Med Biol 658:111–116
Teitelbaum SL, Ross FP (2003) Genetic regulation of osteoclast development and function. Nat Rev Genet 4:638–649
Udagawa N, Takahashi N, Akatsu T, Tanaka H, Sasaki T, Nishihara T, Koga T, Martin TJ, Suda T (1990) Origin of osteoclasts: mature monocytes and macrophages are capable of differentiating into osteoclasts under a suitable microenvironment prepared by bone marrow-derived stromal cells. Proc Natl Acad Sci U S A 87:7260–7264
Udagawa N, Takahashi N, Jimi E, Matsuzaki K, Tsurukai T, Itoh K, Nakagawa N, Yasuda H, Goto M, Tsuda E, Higashio K, Gillespie MT, Martin TJ, Suda T (1999) Osteoblasts/stromal cells stimulate osteoclast activation through expression of osteoclast differentiation factor/RANKL but not macrophage colony-stimulating factor: receptor activator of NF-kappa B ligand. Bone 25:517–523
Yang M, Birnbaum MJ, MacKay CA, Mason-Savas A, Thompson B, Odgren PR (2008) Osteoclast stimulatory transmembrane protein (OC-STAMP), a novel protein induced by RANKL that promotes osteoclast differentiation. J Cell Physiol 215:497–505
Yasuda H, Shima N, Nakagawa N, Yamaguchi K, Kinosaki M, Mochizuki S, Tomoyasu A, Yano K, Goto M, Murakami A, Tsuda E, Morinaga T, Higashio K, Udagawa N, Takahashi N, Suda T (1998) Osteoclast differentiation factor is a ligand for osteoprotegerin/osteoclastogenesis-inhibitory factor and is identical to TRANCE/RANKL. Proc Natl Acad Sci U S A 95:3597–3602
Yoshida H, Hayashi S, Kunisada T, Ogawa M, Nishikawa S, Okamura H, Sudo T, Shultz LD (1990) The murine mutation osteopetrosis is in the coding region of the macrophage colony stimulating factor gene. Nature 345:442–444
Zhang C, Tang T, Ren W, Zhang X, Dai K (2008) Influence of mouse genetic background on wear particle-induced in vivo inflammatory osteolysis. Inflamm Res 57:211–215
Acknowledgments
This work was supported in part by grants-in-aid for Scientific Research from the Ministry of Education, Science and Culture of Japan (E.S, T.T).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Shun Narahara and Haruna Matsushima contributed equally to this work.
Rights and permissions
About this article
Cite this article
Narahara, S., Matsushima, H., Sakai, E. et al. Genetic backgrounds and redox conditions influence morphological characteristics and cell differentiation of osteoclasts in mice. Cell Tissue Res 348, 81–94 (2012). https://doi.org/10.1007/s00441-012-1325-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-012-1325-8