Abstract
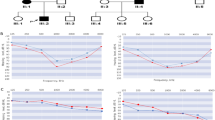
Autosomal dominant nonsyndromic hearing loss (ADNSHL) is a highly genetically heterogeneous disorder. Up to date only approximately 37 ADNSHL-causing genes have been identified. The goal of this study was to determine the causative gene in a five-generation Chinese family with ADNSHL. A Chinese family was ascertained. Simultaneously, two affected individuals and one normal hearing control from the family were analyzed by whole exome capture sequencing. To assess the functional effect of the identified variant, in-vitro studies were performed. novel missense variant, c.512A>G (p.His171Arg) in exon 8 of the ELMO domain-containing 3 (ELMOD3) gene, was identified as a causative variant in this family affected by late-onset and progressive ADNSHL. The variant was validated by Sanger sequencing and found to co-segregate with the phenotype within the pedigree and was absent in 500 ethnically matched unrelated normal hearing control subjects. To our knowledge, this is the first report of a family with ADNSHL caused by ELMOD3 mutation. Western blots and immunofluorescence staining demonstrated that p.His171Arg resulted in abnormal expression levels of ELMOD3 and abnormal subcellular localization. Furthermore, the analysis of the stability of the wild-type (WT) and mutant ELMOD3 protein shows that the decay of p.His171Arg is faster than that of the WT, suggesting a shorter halflife of the c.512A > G variant. A novel variant in the ELMOD3 gene, encoding a member of the engulfment and cell motility (ELMO) family of GTPase-activating proteins, was identified for the first time as responsible for ADNSHL.





Similar content being viewed by others
References
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Gallo Cassarino T, Bertoni M, Bordoli L, Schwede T (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42:W252-8. https://doi.org/10.1093/nar/gku340
Bom SJ, Kemperman MH, De Kok YJ, Huygen PL, Verhagen WI, Cremers FP, Cremers CW (1999) Progressive cochleovestibular impairment caused by a point mutation in the COCH gene at DFNA9. Laryngoscope 109:1525–1530. https://doi.org/10.1097/00005537-199909000-00031
Bowzard JB, Cheng D, Peng J, Kahn RA (2007) ELMOD2 is an Arl2 GTPase-activating protein that also acts on Arfs. J Biol Chem 282:17568–17580. https://doi.org/10.1074/jbc.M701347200
Chakchouk I, Grati M, Bademci G, Bensaid M, Ma Q, Chakroun A, Foster J, Yan D, Duman D, Diaz-Horta O, Ghorbel A, Mittal R, Farooq A, Tekin M, Masmoudi S, Liu XZ (2015) Novel mutations confirm that COL11A2 is responsible for autosomal recessive non-syndromic hearing loss DFNB53. Mol Genet Genom 290:1327–1334. https://doi.org/10.1007/s00438-015-0995-9
Chen H, Jiang L, Xie Z, Mei L, He C, Hu Z, Xia K, Feng Y (2010) Novel mutations of PAX3, MITF, and SOX10 genes in Chinese patients with type I or type II Waardenburg syndrome. Biochem Biophys Res Commun 397:70–74. https://doi.org/10.1016/j.bbrc.2010.05.066
Chung S, Gumienny TL, Hengartner MO, Driscoll M (2000) A common set of engulfment genes mediates removal of both apoptotic and necrotic cell corpses in C. elegans. Nat Cell Biol 2:931–937. https://doi.org/10.1038/35046585
Drummond MC, Belyantseva IA, Friderici KH, Friedman TB (2012) Actin in hair cells and hearing loss. Hear Res 288:89–99. https://doi.org/10.1016/j.heares.2011.12.003
East MP, Bowzard JB, Dacks JB, Kahn RA (2012) ELMO domains, evolutionary and functional characterization of a novel GTPase-activating protein (GAP) domain for Arf protein family GTPases. J Biol Chem 287:39538–39553. https://doi.org/10.1074/jbc.M112.417477
Elliott MR, Zheng S, Park D, Woodson RI, Reardon MA, Juncadella IJ, Kinchen JM, Zhang J, Lysiak JJ, Ravichandran KS (2010) Unexpected requirement for ELMO1 in clearance of apoptotic germ cells in vivo. Nature 467:333–337. https://doi.org/10.1038/nature09356
Gao X, Huang SS, Yuan YY, Wang GJ, Xu JC, Ji YB, Han MY, Yu F, Kang DY, Lin X, Dai P (2015) Targeted gene capture and massively parallel sequencing identify TMC1 as the causative gene in a six-generation Chinese family with autosomal dominant hearing loss. Am J Med Genet A 167A:2357–2365. https://doi.org/10.1002/ajmg.a.37206
Gumienny TL, Brugnera E, Tosello-Trampont AC, Kinchen JM, Haney LB, Nishiwaki K, Walk SF, Nemergut ME, Macara IG, Francis R, Schedl T, Qin Y, Van Aelst L, Hengartner MO, Ravichandran KS (2001) CED-12/ELMO, a novel member of the CrkII/Dock180/Rac pathway, is required for phagocytosis and cell migration. Cell 107:27–41
Hildebrand MS, Thorne NP, Bromhead CJ, Kahrizi K, Webster JA, Fattahi Z, Bataejad M, Kimberling WJ, Stephan D, Najmabadi H, Bahlo M, Smith RJ (2010) Variable hearing impairment in a DFNB2 family with a novel MYO7A missense mutation. Clin Genet 77:563–571. https://doi.org/10.1111/j.1399-0004.2009.01344.x
Huang J, MacKerell AD (2013) CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J Comput Chem 34:2135–2145. https://doi.org/10.1002/jcc.23354
Imtiaz A, Maqsood A, Rehman AU, Morell RJ, Holt JR, Friedman TB, Naz S (2016) Recessive mutations of TMC1 associated with moderate to severe hearing loss. Neurogenetics 17:115–123. https://doi.org/10.1007/s10048-016-0477-1
Jaworek TJ, Richard EM, Ivanova AA, Giese AP, Choo DI, Khan SN, Riazuddin S, Kahn RA, Riazuddin S (2013) An alteration in ELMOD3, an Arl2 GTPase-activating protein, is associated with hearing impairment in humans. PLoS Genet 9:e1003774. https://doi.org/10.1371/journal.pgen.1003774
Johnson KR, Longo-Guess CM, Gagnon LH (2012) Mutations of the mouse ELMO domain containing 1 gene (Elmod1) link small GTPase signaling to actin cytoskeleton dynamics in hair cell stereocilia. PLoS One 7:e36074. https://doi.org/10.1371/journal.pone.0036074
Kitajiri S, Sakamoto T, Belyantseva IA, Goodyear RJ, Stepanyan R, Fujiwara I, Bird JE, Riazuddin S, Riazuddin S, Ahmed ZM, Hinshaw JE, Sellers J, Bartles JR, Hammer JA, Richardson GP, Griffith AJ, Frolenkov GI, Friedman TB (2010) Actin-bundling protein TRIOBP forms resilient rootlets of hair cell stereocilia essential for hearing. Cell 141:786–798. https://doi.org/10.1016/j.cell.2010.03.049
Konings A, Van Camp G, Goethals A, Van Eyken E, Vandevelde A, Ben Azza J, Peeters N, Wuyts W, Smeets H, Van Laer L (2008) Mutation analysis of mitochondrial DNA 12SrRNA and tRNASer(UCN) genes in non-syndromic hearing loss patients. Mitochondrion 8:377–382. https://doi.org/10.1016/j.mito.2008.08.001
LeMasurier M, Gillespie PG (2005) Hair-cell mechanotransduction and cochlear amplification. Neuron 48:403–415. https://doi.org/10.1016/j.neuron.2005.10.017
Morell RJ, Kim HJ, Hood LJ, Goforth L, Friderici K, Fisher R, Van Camp G, Berlin CI, Oddoux C, Ostrer H, Keats B, Friedman TB (1998) Mutations in the connexin 26 gene (GJB2) among Ashkenazi Jews with nonsyndromic recessive deafness. N Engl J Med 339:1500–1505. https://doi.org/10.1056/NEJM199811193392103
Park HJ, Shaukat S, Liu XZ, Hahn SH, Naz S, Ghosh M, Kim HN, Moon SK, Abe S, Tukamoto K, Riazuddin S, Kabra M, Erdenetungalag R, Radnaabazar J, Khan S, Pandya A, Usami SI, Nance WE, Wilcox ER, Riazuddin S, Griffith AJ (2003) Origins and frequencies of SLC26A4 (PDS) mutations in east and south Asians: global implications for the epidemiology of deafness. J Med Genet 40:242–248
Petersen MB (2002) Non-syndromic autosomal-dominant deafness. Clin Genet 62:1–13
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kale L, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802. https://doi.org/10.1002/jcc.20289
Santos RL, Wajid M, Khan MN, McArthur N, Pham TL, Bhatti A, Lee K, Irshad S, Mir A, Yan K, Chahrour MH, Ansar M, Ahmad W, Leal SM (2005) Novel sequence variants in the TMC1 gene in Pakistani families with autosomal recessive hearing impairment. Hum Mutat 26:396. https://doi.org/10.1002/humu.9374
Sherry ST, Ward M, Sirotkin K (2000) Use of molecular variation in the NCBI dbSNP database. Hum Mutat 15(1):68–75
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, Sirotkin K (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29: 308–311
Tekin D, Yan D, Bademci G, Feng Y, Guo S, Foster J, Blanton S, Tekin M, Liu X (2016) A next-generation sequencing gene panel (MiamiOtoGenes) for comprehensive analysis of deafness genes. Hear Res 333:179–184. https://doi.org/10.1016/j.heares.2016.01.018
Urrutia RA, Kalinec F (2015) Biology and pathobiology of lipid droplets and their potential role in the protection of the organ of Corti. Hear Res 330:26–38. https://doi.org/10.1016/j.heares.2015.04.015
Wang H, Wang X, He C, Li H, Qing J, Grati M, Hu Z, Li J, Hu Y, Xia K, Mei L, Wang X, Yu J, Chen H, Jiang L, Liu Y, Men M, Zhang H, Guan L, Xiao J, Zhang J, Liu X, Feng Y (2015a) Exome sequencing identifies a novel CEACAM16 mutation associated with autosomal dominant nonsyndromic hearing loss DFNA4B in a Chinese family. J Hum Genet 60:119–126. https://doi.org/10.1038/jhg.2014.114
Wang HY, Zhao YL, Liu Q, Yuan H, Gao Y, Lan L, Yu L, Wang DY, Guan J, Wang QJ (2015b) Identification of two disease-causing Genes TJP2 and GJB2 in a Chinese family with unconditional autosomal dominant nonsyndromic hereditary hearing impairment. Chin Med J (Engl) 128:3345–3351. https://doi.org/10.4103/0366-6999.171440
Wu YC, Tsai MC, Cheng LC, Chou CJ, Weng NY (2001) C. elegans CED-12 acts in the conserved crkII/DOCK180/Rac pathway to control cell migration and cell corpse engulfment. Dev Cell 1:491–502
Xia JH, Liu CY, Tang BS, Pan Q, Huang L, Dai HP, Zhang BR, Xie W, Hu DX, Zheng D, Shi XL, Wang DA, Xia K, Yu KP, Liao XD, Feng Y, Yang YF, Xiao JY, Xie DH, Huang JZ (1998) Mutations in the gene encoding gap junction protein beta-3 associated with autosomal dominant hearing impairment. Nat Genet 20:370–373. https://doi.org/10.1038/3845
Xia H, Xu H, Deng X, Yuan L, Xiong W, Yang Z, Deng H (2016) Compound heterozygous GJB2 mutations associated to a consanguineous Han family with autosomal recessive non-syndromic hearing loss. Acta Otolaryngol 136:782–785. https://doi.org/10.3109/00016489.2016.1157727
Zhang H, Chen H, Luo H, An J, Sun L, Mei L, He C, Jiang L, Jiang W, Xia K, Li JD, Feng Y (2012) Functional analysis of Waardenburg syndrome-associated PAX3 and SOX10 mutations: report of a dominant-negative SOX10 mutation in Waardenburg syndrome type II. Hum Genet 131:491–503. https://doi.org/10.1007/s00439-011-1098-2
Zhou Z, Caron E, Hartwieg E, Hall A, Horvitz HR (2001) The C. elegans PH domain protein CED-12 regulates cytoskeletal reorganization via a Rho/Rac GTPase signaling pathway. Dev Cell 1:477–489
Acknowledgements
The authors would like to thank the family members for their invaluable cooperation and participation. The study was supported by a grant from the National Natural Science Foundation of China (Grants no. 81470705, 81771023, 81771024, 81500803), the Major State Basic Research Development Program of China (973 Program) (Grants no. 2014CB541702, 2014CB943003), National Institutes of Health/National Institute on Deafness and Other Communication Disorders (Grants no. R01 DC012115, R01 DC01246, R01 DC05575) and in part by the China Postdoctoral Science Foundation (Grants no. 2017M620359).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Patient consent
Obtained.
Ethical approval
The study was approved by the Medical Ethics Committee of Xiangya Hospital, Central South university, in Hunan Province, China.
Data sharing
The raw data can be made available on reasonable request to researchers, after publication.
Additional information
The reported Nucleotide sequence data are available in the GenBank databases under the accession number SUB3343116.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, W., Sun, J., Ling, J. et al. ELMOD3, a novel causative gene, associated with human autosomal dominant nonsyndromic and progressive hearing loss. Hum Genet 137, 329–342 (2018). https://doi.org/10.1007/s00439-018-1885-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-018-1885-0