Abstract
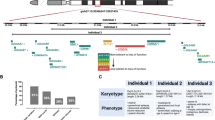
Williams–Beuren syndrome is a neurodevelopmental disorder mainly characterized by dysmorphic features, vascular stenoses, abnormalities of calcium and glucose metabolism, and mental retardation with visuospatial deficits, caused by de novo deletion of 26–28 genes at 7q11.23. Clinical–molecular correlations have defined critically deleted genes as likely responsible for several aspects of the phenotype, but the precise biological pathways affected are mostly unknown. We performed comparative transcriptome profiling of lymphoblastoid cell lines from four Williams–Beuren syndrome patients and two patients with smaller deletions and partial phenotypes. We detected 92 genes deregulated in all patients and 47 genes deregulated only in Williams–Beuren syndrome, with two additional clusters differentially expressed between both groups. Glycolysis and neuronal migration were the most significantly affected pathways by over-representation analyses. In addition, several genes involved in microtubule formation were specifically deregulated in patients with the common deletion. In summary, comparative expression profiling in lymphoblasts has revealed abnormal regulation of gene pathways potentially related to relevant aspects of the Williams–Beuren syndrome phenotype, including the cognitive, visuospatial and metabolic disturbances.



Similar content being viewed by others
References
Allen E, Ding J, Wang W, Pramanik S, Chou J, Yau V, Yang Y (2005) Gigaxonin-controlled degradation of MAP1B light chain is critical to neuronal survival. Nature 438:224–228
Antonell A, Del Campo M, Magano LF, Kaufmann L, Martínez de la Iglesia J, Gallastegui F, Flores R, Schweigmann U, Fauth C, Kotzot D et al (2009) Partial 7q11.23 deletions further implicate GTF2I and GTF2IRD1 as the main genes responsible for the Williams–Beuren syndrome neurocognitive profile. J Med Genet. doi:10.1136/jmg.2009.071712
Baron CA, Liu SY, Hicks C, Gregg JP (2006) Utilization of lymphoblastoid cell lines as a system for the molecular modeling of autism. J Autism Dev Disord 36:973–982
Barrett T, Edgar R (2006) Gene expression omnibus: microarray data storage, submission, retrieval, and analysis. Methods Enzymol 411:352–369
Bayarsaihan D, Bitchevaia N, Enkhmandakh B, Tussie-Luna MI, Leckman JF, Roy A, Ruddle F (2003) Expression of BEN, a member of TFII-I family of transcription factors, during mouse pre- and post-implantation development. Gene Expr Patterns 3:579–589
Bayes M, Magano LF, Rivera N, Flores R, Perez Jurado LA (2003) Mutational mechanisms of Williams–Beuren syndrome deletions. Am J Hum Genet 73:131–151
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300
Billups D, Hanley JG, Orme M, Attwell D, Moss SJ (2000) GABAC receptor sensitivity is modulated by interaction with MAP1B. J Neurosci 20:8643–8650
Bittel DC, Kibiryeva N, Sell SM, Strong TV, Butler MG (2007) Whole genome microarray analysis of gene expression in Prader–Willi syndrome. Am J Med Genet A 143:430–442
Blass JP, Sheu RK, Cedarbaum JM (1988) Energy metabolism in disorders of the nervous system. Rev Neurol (Paris) 144:543–563
Brown V, Jin P, Ceman S, Darnell JC, O’Donnell WT, Tenenbaum SA, Jin X, Feng Y, Wilkinson KD, Keene JD et al (2001) Microarray identification of FMRP-associated brain mRNAs and altered mRNA translational profiles in fragile X syndrome. Cell 107:477–487
Cai Q, Sheng ZH (2009) Molecular motors and synaptic assembly. Neuroscientist 15:78–89
Carleton M, Mao M, Biery M, Warrener P, Kim S, Buser C, Marshall CG, Fernandes C, Annis J, Linsley PS (2006) RNA interference-mediated silencing of mitotic kinesin KIF14 disrupts cell cycle progression and induces cytokinesis failure. Mol Cell Biol 26:3853–3863
Castelo-Branco M, Mendes M, Sebastiao AR, Reis A, Soares M, Saraiva J, Bernardes R, Flores R, Perez-Jurado L, Silva E (2007) Visual phenotype in Williams–Beuren syndrome challenges magnocellular theories explaining human neurodevelopmental visual cortical disorders. J Clin Invest 117:3720–3729
Cherniske EM, Carpenter TO, Klaiman C, Young E, Bregman J, Insogna K, Schultz RT, Pober BR (2004) Multisystem study of 20 older adults with Williams syndrome. Am J Med Genet A 131:255–264
Cheung VG, Conlin LK, Weber TM, Arcaro M, Jen KY, Morley M, Spielman RS (2003) Natural variation in human gene expression assessed in lymphoblastoid cells. Nat Genet 33:422–425
Cusco I, del Campo M, Vilardell M, Gonzalez E, Gener B, Galan E, Toledo L, Perez-Jurado LA (2008) Array-CGH in patients with Kabuki-like phenotype: identification of two patients with complex rearrangements including 2q37 deletions and no other recurrent aberration. BMC Med Genet 9:27
De Zeeuw CI, Hoogenraad CC, Goedknegt E, Hertzberg E, Neubauer A, Grosveld F, Galjart N (1997) CLIP-115, a novel brain-specific cytoplasmic linker protein, mediates the localization of dendritic lamellar bodies. Neuron 19:1187–1199
Durinck S, Moreau Y, Kasprzyk A, Davis S, De Moor B, Brazma A, Huber W (2005) BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21:3439–3440
Eckert MA, Galaburda AM, Mills DL, Bellugi U, Korenberg JR, Reiss AL (2006) The neurobiology of Williams syndrome: cascading influences of visual system impairment? Cell Mol Life Sci 63:1867–1875
Edelmann W, Zervas M, Costello P, Roback L, Fischer I, Hammarback JA, Cowan N, Davies P, Wainer B, Kucherlapati R (1996) Neuronal abnormalities in microtubule-associated protein 1B mutant mice. Proc Natl Acad Sci USA 93:1270–1275
Edelmann L, Prosnitz A, Pardo S, Bhatt J, Cohen N, Lauriat T, Ouchanov L, Gonzalez PJ, Manghi ER, Bondy P et al (2007) An atypical deletion of the Williams–Beuren syndrome interval implicates genes associated with defective visuospatial processing and autism. J Med Genet 44:136–143
Fiala JC, Spacek J, Harris KM (2002) Dendritic spine pathology: cause or consequence of neurological disorders? Brain Res Brain Res Rev 39:29–54
Garcia-Marcos M, Ghosh P, Farquhar MG (2009) GIV is a nonreceptor GEF for G alpha i with a unique motif that regulates Akt signaling. Proc Natl Acad Sci USA 106:3178–3183
Gerber SH, Rah JC, Min SW, Liu X, de Wit H, Dulubova I, Meyer AC, Rizo J, Arancillo M, Hammer RE et al (2008) Conformational switch of syntaxin-1 controls synaptic vesicle fusion. Science 321:1507–1510
Gruneberg U, Neef R, Li X, Chan EH, Chalamalasetty RB, Nigg EA, Barr FA (2006) KIF14 and citron kinase act together to promote efficient cytokinesis. J Cell Biol 172:363–372
Hartigan JA, Wong MA (1979) Algorithm AS 136: a k-means clustering algorithm. Appl Stat 28:100–108
Hocking JC, Hehr CL, Bertolesi G, Funakoshi H, Nakamura T, McFarlane S (2009) LIMK1 acts downstream of BMP signaling in developing retinal ganglion cell axons but not dendrites. Dev Biol 330(2):273–285
Hoogenraad CC, Eussen BH, Langeveld A, van Haperen R, Winterberg S, Wouters CH, Grosveld F, De Zeeuw CI, Galjart N (1998) The murine CYLN2 gene: genomic organization, chromosome localization, and comparison to the human gene that is located within the 7q11.23 Williams syndrome critical region. Genomics 53:348–358
Hoogenraad CC, Akhmanova A, Galjart N, De Zeeuw CI (2004) LIMK1 and CLIP-115: linking cytoskeletal defects to Williams syndrome. Bioessays 26:141–150
Horike S, Cai S, Miyano M, Cheng JF, Kohwi-Shigematsu T (2005) Loss of silent-chromatin looping and impaired imprinting of DLX5 in Rett syndrome. Nat Genet 37:31–40
Hu K, Carroll J, Fedorovich S, Rickman C, Sukhodub A, Davletov B (2002) Vesicular restriction of synaptobrevin suggests a role for calcium in membrane fusion. Nature 415:646–650
Iwamoto K, Kakiuchi C, Bundo M, Ikeda K, Kato T (2004) Molecular characterization of bipolar disorder by comparing gene expression profiles of postmortem brains of major mental disorders. Mol Psychiatry 9:406–416
Kamburov A, Wierling C, Lehrach H, Herwig R (2009) ConsensusPathDB—a database for integrating human functional interaction networks. Nucleic Acids Res 37:623–628
Kerr MK (2003) Design considerations for efficient and effective microarray studies. Biometrics 59:822–828
Kuo TY, Hong CJ, Hsueh YP (2009) Bcl11A/CTIP1 regulates expression of DCC and MAP1b in control of axon branching and dendrite outgrowth. Mol Cell Neurosci 42:195–207
Martens MA, Wilson SJ, Reutens DC (2008) Research review: Williams syndrome: a critical review of the cognitive, behavioral, and neuroanatomical phenotype. J Child Psychol Psychiatry 49:576–608
Meixner A, Haverkamp S, Wassle H, Fuhrer S, Thalhammer J, Kropf N, Bittner RE, Lassmann H, Wiche G, Propst F (2000) MAP1B is required for axon guidance and is involved in the development of the central and peripheral nervous system. J Cell Biol 151:1169–1178
Meloni I, Muscettola M, Raynaud M, Longo I, Bruttini M, Moizard MP, Gomot M, Chelly J, des Portes V, Fryns JP et al (2002) FACL4, encoding fatty acid-CoA ligase 4, is mutated in nonspecific X-linked mental retardation. Nat Genet 30:436–440
Meng Y, Zhang Y, Tregoubov V, Janus C, Cruz L, Jackson M, Lu WY, MacDonald JF, Wang JY, Falls DL et al (2002) Abnormal spine morphology and enhanced LTP in LIMK-1 knockout mice. Neuron 35:121–133
Merla G, Howald C, Antonarakis SE, Reymond A (2004) The subcellular localization of the ChoRE-binding protein, encoded by the Williams–Beuren syndrome critical region gene 14, is regulated by 14-3-3. Hum Mol Genet 13:1505–1514
Merla G, Howald C, Henrichsen CN, Lyle R, Wyss C, Zabot MT, Antonarakis SE, Reymond A (2006) Submicroscopic deletion in patients with Williams–Beuren syndrome influences expression levels of the nonhemizygous flanking genes. Am J Hum Genet 79:332–341
Meyer-Lindenberg A, Kohn P, Mervis CB, Kippenhan JS, Olsen RK, Morris CA, Berman KF (2004) Neural basis of genetically determined visuospatial construction deficit in Williams syndrome. Neuron 43:623–631
Moss SJ, Smart TG (2001) Constructing inhibitory synapses. Nat Rev Neurosci 2:240–250
Nagamatsu S, Nakamichi Y, Katahira H (1997) Syntaxin, but not soluble NSF attachment protein (SNAP), biosynthesis by rat pancreatic islets is regulated by glucose in parallel with proinsulin biosynthesis. Diabetologia 40:1396–1402
Nakagawa T, Tanaka Y, Matsuoka E, Kondo S, Okada Y, Noda Y, Kanai Y, Hirokawa N (1997) Identification and classification of 16 new kinesin superfamily (KIF) proteins in mouse genome. Proc Natl Acad Sci USA 94:9654–9659
Nakajima H, Hirata A, Ogawa Y, Yonehara T, Yoda K, Yamasaki M (1991) A cytoskeleton-related gene, uso1, is required for intracellular protein transport in Saccharomyces cerevisiae. J Cell Biol 113:245–260
Nishimura Y, Martin CL, Vazquez-Lopez A, Spence SJ, Alvarez-Retuerto AI, Sigman M, Steindler C, Pellegrini S, Schanen NC, Warren ST et al (2007) Genome-wide expression profiling of lymphoblastoid cell lines distinguishes different forms of autism and reveals shared pathways. Hum Mol Genet 16:1682–1698
Nothwang HG, Kim HG, Aoki J, Geisterfer M, Kubart S, Wegner RD, van Moers A, Ashworth LK, Haaf T, Bell J et al (2001) Functional hemizygosity of PAFAH1B3 due to a PAFAH1B3-CLK2 fusion gene in a female with mental retardation, ataxia and atrophy of the brain. Hum Mol Genet 10:797–806
Olshen AB, Venkatraman ES, Lucito R, Wigler M (2004) Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5:557–572
Pérez Jurado LA (2003) Williams–Beuren syndrome: a model of recurrent genomic mutation. Horm Res 59(Suppl 1):106–113
Pober BR (2010) Williams–Beuren syndrome. N Engl J Med 362:239–252
Pober BR, Morris CA (2007) Diagnosis and management of medical problems in adults with Williams–Beuren syndrome. Am J Med Genet 145C:280–290
Riederer BM (2007) Microtubule-associated protein 1B, a growth-associated and phosphorylated scaffold protein. Brain Res Bull 71:541–558
Romeo S, Sentinelli F, Cavallo MG, Leonetti F, Fallarino M, Mariotti S, Baroni MG (2008) Search for genetic variants of the SYNTAXIN 1A (STX1A) gene: the −352 A>T variant in the STX1A promoter associates with impaired glucose metabolism in an Italian obese population. Int J Obes (Lond) 32:413–420
Roy AL (2007) Signal-induced functions of the transcription factor TFII-I. Biochim Biophys Acta 1769:613–621
Scharpf RB, Iacobuzio-Donahue CA, Sneddon JB, Parmigiani G (2007) When should one subtract background fluorescence in 2-color microarrays? Biostatistics 8:695–707
Schwender H, Krause A, Ickstadt K (2006) Identifying interesting genes with siggenes. RNews 6:45–50
Stranger BE, Forrest MS, Clark AG, Minichiello MJ, Deutsch S, Lyle R, Hunt S, Kahl B, Antonarakis SE, Tavare S et al (2005) Genome-wide associations of gene expression variation in humans. PLoS Genet 1:e78
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Subramanian A, Kuehn H, Gould J, Tamayo P, Mesirov JP (2007) GSEA-P: a desktop application for gene set enrichment analysis. Bioinformatics 23:3251–3253
Svaasand EK, Aasly J, Landsem VM, Klungland H (2007) Altered expression of PGK1 in a family with phosphoglycerate kinase deficiency. Muscle Nerve 36:679–684
Tam E, Young EJ, Morris CA, Marshall CR, Loo W, Scherer SW, Mervis CB, Osborne LR (2008) The common inversion of the Williams–Beuren syndrome region at 7q11.23 does not cause clinical symptoms. Am J Med Genet A 146A:1797–1806
Traer CJ, Rutherford AC, Palmer KJ, Wassmer T, Oakley J, Attar N, Carlton JG, Kremerskothen J, Stephens DJ, Cullen PJ (2007) SNX4 coordinates endosomal sorting of TfnR with dynein-mediated transport into the endocytic recycling compartment. Nat Cell Biol 9:1370–1380
Vaillend C, Poirier R, Laroche S (2008) Genes, plasticity and mental retardation. Behav Brain Res 192:88–105
Vastrik I, D’Eustachio P, Schmidt E, Gopinath G, Croft D, de Bono B, Gillespie M, Jassal B, Lewis S, Matthews L et al (2007) Reactome: a knowledge base of biologic pathways and processes. Genome Biol 8:R39
Vilardell M, Vives S, Lozano J, Cusco I, Gonzalez E, García M, Sumoy L, Perez Jurado LA (2008) Statistical assessment of sources of systematic variation in aCGH experiments. JP J Biostat 2:81–168
Yan X, Zhao X, Qian M, Guo N, Gong X, Zhu X (2000) Characterization and gene structure of a novel retinoblastoma-protein-associated protein similar to the transcription regulator TFII-I. Biochem J 345(Pt 3):749–757
Zhang R, Maksymowych AB, Simpson LL (1995) Cloning and sequence analysis of a cDNA encoding human syntaxin 1A, a polypeptide essential for exocytosis. Gene 159:293–294
Acknowledgments
We thank patients and families for their collaboration and Drs Dieter Kotzot and Jorge Martínez de la Iglesia for referring cases with atypical deletions. LCLs from controls and two WBS patients were kindly provided by Drs Xavier Estivill and Lucy Osborne, respectively. We also thank Drs Benjamín Rodriguez-Santiago and Andrés Medrano for helpful discussion and Annabelle Parent and Anna Carreras for technical assistance, as well as Charlotte Henrichsen, Alexandre Reymond and Giuseppe Merla for sharing unpublished transcriptomic data from WBS fibroblast cell lines. This work was funded by the European Commission AnEUploidy integrated project (FP6-2005-LIFESCIHEALTH-7) to L.P.J. The experiments comply with the current laws of the country in which they were performed.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Antonell, A., Vilardell, M. & Pérez Jurado, L.A. Transcriptome profile in Williams–Beuren syndrome lymphoblast cells reveals gene pathways implicated in glucose intolerance and visuospatial construction deficits. Hum Genet 128, 27–37 (2010). https://doi.org/10.1007/s00439-010-0817-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-010-0817-4