Abstract
Purpose
Epidermal growth factor receptor (EGFR) mutation testing has several limitations. Therefore, we built predictive models to determine the EGFR mutation status of patients and guide therapeutic decision-making.
Methods
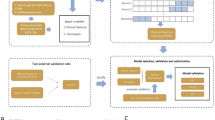
We collected data from 320 patients with lung carcinoma, including sex, age, smoking history, serum tumour marker levels, maximum standardized uptake value, pathological results, computed tomography images, and EGFR mutation status. Artificial neural network (ANN) models based on multiple clinical characteristics were proposed to predict EGFR mutation status.
Results
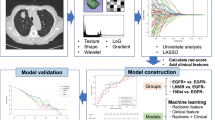
A training set (n = 200) was used to develop predictive models of the EGFR mutation status (Model 1: area under the receiver operating characteristic curve [AUROC] = 0.910, 95% CI 0.861–0.945; Model 2: AUROC = 0.859, 95% CI 0.803–0.904; Model 3: AUROC = 0.711, 95% CI 0.643–0.773). A testing set (n = 50) and temporal validation data set (n = 70) were used to evaluate the generalisation performance of the established models (testing set: Model 1, AUROC = 0.845, 95% CI 0.715–0.932; Model 2, AUROC = 0.882, 95% CI 0.759–0.956; Model 3, AUROC = 0.817, 95% CI 0.682–0.912; temporal validation dataset: Model 1, AUROC = 0.909, 95% CI 0.816–0.964; Model 2, AUROC = 0.855, 95% CI 0.751–0.928; Model 3, AUROC = 0.831, 95% CI 0.723–0.910). The predictive abilities of the three ANN models were superior to that of a previous logistic regression model (P < 0.001, 0.027, and 0.050, respectively).
Conclusions
ANN models provide a non-invasive and readily available method for EGFR mutation status prediction.


Similar content being viewed by others
Data availability
The data sets analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- ANN:
-
Artificial neural network
- AUROC:
-
Area under receiver operating characteristic curve
- CI:
-
Confidence interval
- CT:
-
Computed tomography
- CEA:
-
Carcinoembryonic antigen
- CYFRA21-1:
-
Cytokeratin 19 fragment
- EGFR:
-
Epidermal growth factor receptor
- EGFR-TKIs:
-
Epidermal growth factor receptor tyrosine kinase inhibitors
- 18F-FDG:
-
18F-fluorodeoxyglucose
- NSE:
-
Neuron-specific enolase
- OR:
-
Odds ratio
- PET/CT:
-
Positron emission tomography/computed tomography
- ROC:
-
Receiver operating characteristic
- SCCA:
-
Squamous cell carcinoma antigen
- SUVmax :
-
Maximum standardized uptake value
References
Bray F, Ferlay J, Soerjomataram I et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Brinkman GL, Coates EO Jr (1963) The effect of bronchitis, smoking, and occupation on ventilation. Am Rev Respir Dis 87:684–693
Cai Z (2016) Relationship between serum carcinoembryonic antigen level and epidermal growth factor receptor mutations with the influence on the prognosis of non-small-cell lung cancer patients. Oncol Targets Ther 9:3873–3878. https://doi.org/10.2147/OTT.S102199
Cho A, Hur J, Moon YW et al (2016) Correlation between EGFR gene mutation, cytologic tumor markers, 18F-FDG uptake in non-small cell lung cancer. BMC Cancer 16:224. https://doi.org/10.1186/s12885-016-2251-z
Cross SS, Harrison RF, Kennedy RL (1995) Introduction to neural networks. Lancet 346:1075–1079. https://doi.org/10.1016/s0140-6736(95)91746-2
Cucchetti A, Piscaglia F, Grigioni AD et al (2010) Preoperative prediction of hepatocellular carcinoma tumour grade and micro-vascular invasion by means of artificial neural network: a pilot study. J Hepatol 52:880–888. https://doi.org/10.1016/j.jhep.2009.12.037
Fei Y, Hu J, Gao K et al (2017) Predicting risk for portal vein thrombosis in acute pancreatitis patients: a comparison of radical basis function artificial neural network and logistic regression models. J Crit Care 39:115–123. https://doi.org/10.1016/j.jcrc.2017.02.032
Gazdar AF (2009) Personalized medicine and inhibition of EGFR signaling in lung cancer. N Engl J Med 361:1018–1020. https://doi.org/10.1056/NEJMe0905763
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Gu J, Xu S, Huang L et al (2018) Value of combining serum carcinoembryonic antigen and PET/CT in predicting EGFR mutation in non-small cell lung cancer. J Thorac Dis 10:723–731. https://doi.org/10.21037/jtd.2017.12.143
Hong SJ, Kim TJ, Choi YW et al (2016) Radiogenomic correlation in lung adenocarcinoma with epidermal growth factor receptor mutations: imaging features and histological subtypes. Eur Radiol 26:3660–3668. https://doi.org/10.1007/s00330-015-4196-z
Jorge SE, Kobayashi SS, Costa DB (2014) Epidermal growth factor receptor (EGFR) mutations in lung cancer: preclinical and clinical data. Braz J Med Biol Res 47:929–939. https://doi.org/10.1590/1414-431X20144099
Kim TJ, Lee CT, Jheon SH et al (2016) Radiologic characteristics of surgically resected non-small cell lung cancer with ALK rearrangement or EGFR mutations. Ann Thorac Surg 101:473–480. https://doi.org/10.1016/j.athoracsur.2015.07.062
Lai Y, Zhang Z, Li J et al (2013) EGFR mutations in surgically resected fresh specimens from 697 consecutive Chinese patients with non-small cell lung cancer and their relationships with clinical features. Int J Mol Sci 14:24549–24559. https://doi.org/10.3390/ijms141224549
Landry AP, Ting WKC, Zador Z et al (2018) Using artificial neural networks to identify patients with concussion and postconcussion syndrome based on antisaccades. J Neurosurg 1:1–8. https://doi.org/10.3171/2018.6.JNS18607
Lee SM, Bae SK, Jung SJ et al (2015) FDG uptake in non-small cell lung cancer is not an independent predictor of EGFR or KRAS mutation status: a retrospective analysis of 206 patients. Clin Nucl Med 40:950–958. https://doi.org/10.1097/RLU.0000000000000975
Loughran CF, Keeling CR (2011) Seeding of tumour cells following breast biopsy: a literature review. Br J Radiol 84:869–874. https://doi.org/10.1259/bjr/77245199
Lv Z, Fan J, Xu J et al (2018) Value of 18F-FDG PET/CT for predicting EGFR mutations and positive ALK expression in patients with non-small cell lung cancer: a retrospective analysis of 849 Chinese patients. Eur J Nucl Med Mol Imaging 45:735–750. https://doi.org/10.1007/s00259-017-3885-z
Lynch TJ, Bell DW, Sordella R et al (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 350:2129–2139. https://doi.org/10.1056/NEJMoa040938
McHugh ML (2012) Interrater reliability: the kappa statistic. Biochem Med (Zagreb) 22:276–282. https://doi.org/10.11613/BM.2012.031
Na II, Byun BH, Kim KM et al (2010) 18F-FDG uptake and EGFR mutations in patients with non-small cell lung cancer: a single-institution retrospective analysis. Lung Cancer 67:76–80. https://doi.org/10.1016/j.lungcan.2009.03.010
Paez JG, Jänne PA, Lee JC et al (2004) EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304:1497–1500. https://doi.org/10.1126/science.1099314
Paydar K, Niakan Kalhori SR, Akbarian M et al (2017) A clinical decision support system for prediction of pregnancy outcome in pregnant women with systemic lupus erythematosus. Int J Med Inform 97:239–246. https://doi.org/10.1016/j.ijmedinf.2016.10.018
Rashidi Khazaee P, Bagherzadeh J, Niazkhani Z et al (2018) A dynamic model for predicting graft function in kidney recipients’ upcoming follow up visits: a clinical application of artificial neural network. Int J Med Inform 119:125–133. https://doi.org/10.1016/j.ijmedinf.2018.09.012
Rizzo S, Petrella F, Buscarino V et al (2016) CT radiogenomic characterization of EGFR, K-RAS, and ALK mutations in non-small cell lung cancer. Eur Radiol 26:32–42. https://doi.org/10.1007/s00330-015-3814-0
Sacher AG, Dahlberg SE, Heng J et al (2016) Association between younger age and targetable genomic alterations and prognosis in non-small-cell lung cancer. JAMA Oncol 2:313–320. https://doi.org/10.1001/jamaoncol.2015.4482
Shi HY, Hwang SL, Lee KT et al (2013) In-hospital mortality after traumatic brain injury surgery: a nationwide population-based comparison of mortality predictors used in artificial neural network and logistic regression models. J Neurosurg 118:746–752. https://doi.org/10.3171/2013.1.JNS121130
Torre LA, Siegel RL, Jemal A (2016) Lung cancer statistics. Adv Exp Med Biol 893:1–19. https://doi.org/10.1007/978-3-319-24223-1_1
Wang S, Shi J, Ye Z et al (2019) Predicting EGFR mutation status in lung adenocarcinoma on computed tomography image using deep learning. Eur Respir J. https://doi.org/10.1183/13993003.00986-2018
Xiong JF, Jia TY, Li XY et al (2018) Identifying epidermal growth factor receptor mutation status in patients with lung adenocarcinoma by three-dimensional convolutional neural networks. Br J Radiol 91:20180334. https://doi.org/10.1259/bjr.20180334
Yang X, Lin D (2016) Changes of 2015 WHO histological classification of lung cancer and the clinical significance. Zhongguo Fei Ai Za Zhi 19:332–336. https://doi.org/10.3779/j.issn.1009-3419.2016.06.06
Zain AM, Haron H, Sharif S (2010) Prediction of surface roughness in the end milling machining using Artificial Neural Network. Expert Syst Appl 37:1755–1768. https://doi.org/10.1016/j.eswa.2009.07.033
Acknowledgements
We appreciate the help from two respiratory physicians (L. Yang and Y. Lu) for CT image interpretation, and the staff of the record retrieval department from the First Affiliated Hospital of Wenzhou Medical University for their efforts.
Funding
No external funding for this research need be declared.
Author information
Authors and Affiliations
Contributions
XYQ and WZ conceived and designed this study; HLW, XH and XLG helped with the collection and assembly of data. All authors contributed toward data analysis, drafting and critically revising the paper and agree to be accountable for all aspects of the work. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval and consent to participate
This study was approved by the Ethical Committee of the First Affiliated Hospital of Wenzhou Medical University. All study participants provided written informed consent, and their data confidentiality were protected.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
432_2019_3103_MOESM3_ESM.tif
Supplementary material 3 Fig. S2. The confusion matrices for the three artificial neural network (ANN) models to predict epidermal growth factor receptor (EGFR) mutation status in the training set and testing set of modelling data set. a, all-variable model in the training set; b, significant-variable model in the training set; c, traditional-variable model in the training set; d, all-variable model in the testing set; e, significant-variable model in the testing set; f, traditional-variable model in the testing set (TIFF 4377 kb)
432_2019_3103_MOESM4_ESM.tif
Supplementary material 4 Fig. S3. The confusion matrices for the three artificial neural network (ANN) models to predict epidermal growth factor receptor (EGFR) mutation status in the temporal validation data set. a, all-variable model; b, significant-variable model; c, traditional-variable model (TIFF 2177 kb)
Rights and permissions
About this article
Cite this article
Qin, X., Wang, H., Hu, X. et al. Predictive models for patients with lung carcinomas to identify EGFR mutation status via an artificial neural network based on multiple clinical information. J Cancer Res Clin Oncol 146, 767–775 (2020). https://doi.org/10.1007/s00432-019-03103-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-019-03103-x