Abstract
Purpose
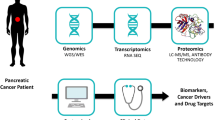
Pancreatic cancer is commonly detected at advanced stages when the tumor is no longer amenable to surgical resection. Therefore, finding biomarkers for early stage disease is urgent. Here, we show that high-definition mass spectrometry (HDMSE) can be used to identify serum protein alterations associated with early stage pancreatic cancer.
Methods
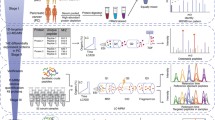
We analyzed serum samples from patients with resectable pancreatic cancer, benign pancreatic disease, and healthy controls. The SYNAPT G2-Si platform was used in a data-independent manner coupled with ion mobility. The dilution of the samples with yeast alcohol dehydrogenase tryptic digest of known concentration allowed the estimated amounts of each identified protein to be calculated (Silva et al. in Anal Chem 77:2187–2200, 2005; Silva et al. in Mol Cell Proteomics 5:144–156, 2006). A global protein expression comparison of the three study groups was made using label-free quantification and bioinformatic analyses.
Results
Two-way unsupervised hierarchical clustering revealed 134 proteins that successfully classified pancreatic cancer patients from the controls, and identified 40 proteins that showed a significant up-regulation in the pancreatic cancer group. This discrimination reliability was further confirmed by principal component analysis. The differentially expressed candidates were aligned with protein network analyses and linked to biological pathways related to pancreatic tumorigenesis. Pancreatic disease link associations could be made for BAZ2A, CDK13, DAPK1, DST, EXOSC3, INHBE, KAT2B, KIF20B, SMC1B, and SPAG5, by pathway network linkages to p53, the most frequently altered tumor suppressor in pancreatic cancer.
Conclusion
These pancreatic cancer study candidates may provide new avenues of research for a noninvasive blood-based diagnosis for pancreatic tumor stratification.






Similar content being viewed by others
References
Ansari D, Aronsson L, Sasor A, Welinder C, Rezeli M, Marko-Varga G et al (2014) The role of quantitative mass spectrometry in the discovery of pancreatic cancer biomarkers for translational science. J Transl Med 12:87. doi:10.1186/1479-5876-12-87
Balbas-Martinez C, Sagrera A, Carrillo-de-Santa-Pau E, Earl J, Marquez M, Vazquez M et al (2013) Recurrent inactivation of STAG2 in bladder cancer is not associated with aneuploidy. Nat Genet 45:1464–1469. doi:10.1038/ng.2799
Bond NJ, Shliaha PV, Lilley KS, Gatto L (2013) Improving qualitative and quantitative performance for MS(E)-based label-free proteomics. J Proteome Res 12:2340–2353. doi:10.1021/pr300776t
Bruning A, Matsingou C, Brem GJ, Rahmeh M, Mylonas I (2012) Inhibin beta E is upregulated by drug-induced endoplasmic reticulum stress as a transcriptional target gene of ATF4. Toxicol Appl Pharmacol 264:300–304. doi:10.1016/j.taap.2012.08.011
Charrier A, Chen R, Chen L, Kemper S, Hattori T, Takigawa M et al (2014) Connective tissue growth factor (CCN2) and microRNA-21 are components of a positive feedback loop in pancreatic stellate cells (PSC) during chronic pancreatitis and are exported in PSC-derived exosomes. Journal of cell communication and signaling. doi:10.1007/s12079-014-0220-3
Chen R, Pan S, Brentnall TA, Aebersold R (2005) Proteomic profiling of pancreatic cancer for biomarker discovery. Mol Cell Proteomics 4:523–533. doi:10.1074/mcp.R500004-MCP200
Clayton A (2012) Cancer cells use exosomes as tools to manipulate immunity and the microenvironment. Oncoimmunology 1:78–80. doi:10.4161/onci.1.1.17826
Dangi-Garimella S, Sahai V, Ebine K, Kumar K, Munshi HG (2013) Three-dimensional collagen I promotes gemcitabine resistance in vitro in pancreatic cancer cells through HMGA2-dependent histone acetyltransferase expression. PLoS ONE 8:e64566. doi:10.1371/journal.pone.0064566
Decker GA, Batheja MJ, Collins JM, Silva AC, Mekeel KL, Moss AA et al (2010) Risk factors for pancreatic adenocarcinoma and prospects for screening. Gastroenterol Hepatol (NY) 6:246–254
Domon B, Aebersold R (2010) Options and considerations when selecting a quantitative proteomics strategy. Nat Biotechnol 28:710–721. doi:10.1038/nbt.1661
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A et al (2013) STRING v9.1: protein–protein interaction networks, with increased coverage and integration. Nucleic Acids Res 41:808–815. doi:10.1093/nar/gks1094
Gaspar NJ, Li L, Kapoun AM, Medicherla S, Reddy M, Li G et al (2007) Inhibition of transforming growth factor beta signaling reduces pancreatic adenocarcinoma growth and invasiveness. Mol Pharmacol 72:152–161. doi:10.1124/mol.106.029025
Goonetilleke KS, Siriwardena AK (2007) Systematic review of carbohydrate antigen (CA 19-9) as a biochemical marker in the diagnosis of pancreatic cancer. Eur J Surg Oncol 33:266–270. doi:10.1016/j.ejso.2006.10.004
Hanash SM, Pitteri SJ, Faca VM (2008) Mining the plasma proteome for cancer biomarkers. Nature 452:571–579. doi:10.1038/nature06916
Harrison B, Kraus M, Burch L, Stevens C, Craig A, Gordon-Weeks P et al (2008) DAPK-1 binding to a linear peptide motif in MAP1B stimulates autophagy and membrane blebbing. J Biol Chem 283:9999–10014. doi:10.1074/jbc.M706040200
Hashimoto O, Ushiro Y, Sekiyama K, Yamaguchi O, Yoshioka K, Mutoh K et al (2006) Impaired growth of pancreatic exocrine cells in transgenic mice expressing human activin betaE subunit. Biochem Biophys Res Commun 341:416–424. doi:10.1016/j.bbrc.2005.12.205
Hsiao YH, Su YA, Tsai HD, Mason JT, Chou MC, Man YG (2010) Increased invasiveness and aggressiveness in breast epithelia with cytoplasmic p63 expression. Int J Biol Sci 6:428–442
Jones S, Zhang X, Parsons DW, Lin JC, Leary RJ, Angenendt P et al (2008) Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321:1801–1806. doi:10.1126/science.1164368
Kahlert C, Melo SA, Protopopov A, Tang J, Seth S, Koch M et al (2014) Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J Biol Chem 289:3869–3875. doi:10.1074/jbc.C113.532267
Kim KH, Yoo HY, Joo KM, Jung Y, Jin J, Kim Y et al (2011) Time-course analysis of DNA damage response-related genes after in vitro radiation in H460 and H1229 lung cancer cell lines. Exp Mol Med 43:419–426. doi:10.3858/emm.2011.43.7.046
Kim HE, Kim DG, Lee KJ, Son JG, Song MY, Park YM et al (2012) Frequent amplification of CENPF, GMNN and CDK13 genes in hepatocellular carcinomas. PLoS ONE 7:e43223. doi:10.1371/journal.pone.0043223
Kim MS, Pinto SM, Getnet D, Nirujogi RS, Manda SS, Chaerkady R et al (2014) A draft map of the human proteome. Nature 509:575–581. doi:10.1038/nature13302
Langley SR, Dwyer J, Drozdov I, Yin X, Mayr M (2013) Proteomics: from single molecules to biological pathways. Cardiovasc Res 97:612–622. doi:10.1093/cvr/cvs346
Liggett T, Melnikov A, Yi QL, Replogle C, Brand R, Kaul K et al (2010) Differential methylation of cell-free circulating DNA among patients with pancreatic cancer versus chronic pancreatitis. Cancer 116:1674–1680. doi:10.1002/cncr.24893
Liu L, Scolnick DM, Trievel RC, Zhang HB, Marmorstein R, Halazonetis TD et al (1999) p53 sites acetylated in vitro by PCAF and p300 are acetylated in vivo in response to DNA damage. Mol Cell Biol 19:1202–1209
Liu M, Yu H, Huo L, Liu J, Li M, Zhou J (2008) Validating the mitotic kinesin Eg5 as a therapeutic target in pancreatic cancer cells and tumor xenografts using a specific inhibitor. Biochem Pharmacol 76:169–178. doi:10.1016/j.bcp.2008.04.018
Mi H, Muruganujan A, Casagrande JT, Thomas PD (2013) Large-scale gene function analysis with the PANTHER classification system. Nat Protoc 8:1551–1566. doi:10.1038/nprot.2013.092
Missiaglia E, Blaveri E, Terris B, Wang YH, Costello E, Neoptolemos JP et al (2004) Analysis of gene expression in cancer cell lines identifies candidate markers for pancreatic tumorigenesis and metastasis. Int J Cancer 112:100–112. doi:10.1002/ijc.20376
Morton JP, Timpson P, Karim SA, Ridgway RA, Athineos D, Doyle B et al (2010) Mutant p53 drives metastasis and overcomes growth arrest/senescence in pancreatic cancer. Proc Natl Acad Sci USA 107:246–251. doi:10.1073/pnas.0908428107
Oshima M, Okano K, Muraki S, Haba R, Maeba T, Suzuki Y et al (2013) Immunohistochemically detected expression of 3 major genes (CDKN2A/p16, TP53, and SMAD4/DPC4) strongly predicts survival in patients with resectable pancreatic cancer. Ann Surg 258:336–346. doi:10.1097/SLA.0b013e3182827a65
Polanski M, Anderson NL (2007) A list of candidate cancer biomarkers for targeted proteomics. Biomark Insights 1:1–48
Postepska-Igielska A, Krunic D, Schmitt N, Greulich-Bode KM, Boukamp P, Grummt I (2013) The chromatin remodelling complex NoRC safeguards genome stability by heterochromatin formation at telomeres and centromeres. EMBO Rep 14:704–710. doi:10.1038/embor.2013.87
Rodriguez-Suarez E, Hughes C, Gethings L, Giles K, Wildgoose J, Stapels M et al (2013) An ion mobility assisted data independent LC-MS strategy for the analysis of complex biological samples. Curr Anal Chem 9:199–211
Rosenfeldt MT, O’Prey J, Morton JP, Nixon C, MacKay G, Mrowinska A et al (2013) p53 status determines the role of autophagy in pancreatic tumour development. Nature 504:296–300. doi:10.1038/nature12865
Ryan SD, Ferrier A, Kothary R (2012) A novel role for the cytoskeletal linker protein dystonin in the maintenance of microtubule stability and the regulation of ER-Golgi transport. Bioarchitecture 2:2–5
Schnelldorfer T, Ware AL, Sarr MG, Smyrk TC, Zhang L, Qin R et al (2008) Long-term survival after pancreatoduodenectomy for pancreatic adenocarcinoma: is cure possible? Ann Surg 247:456–462. doi:10.1097/SLA.0b013e3181613142
Silva JC, Denny R, Dorschel CA, Gorenstein M, Kass IJ, Li GZ et al (2005) Quantitative proteomic analysis by accurate mass retention time pairs. Anal Chem 77:2187–2200. doi:10.1021/ac048455k
Silva JC, Gorenstein MV, Li GZ, Vissers JP, Geromanos SJ (2006) Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol Cell Proteomics 5:144–156. doi:10.1074/mcp.M500230-MCP200
Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, Barber TD et al (2006) The consensus coding sequences of human breast and colorectal cancers. Science 314:268–274. doi:10.1126/science.1133427
Suzuki N, Hazama S, Ueno T, Matsui H, Shindo Y, Iida M et al (2014) A phase I clinical trial of vaccination with KIF20A-derived peptide in combination with gemcitabine for patients with advanced pancreatic cancer. J Immunother 37:36–42. doi:10.1097/CJI.0000000000000012
Taniuchi K, Nakagawa H, Nakamura T, Eguchi H, Ohigashi H, Ishikawa O et al (2005) Down-regulation of RAB6KIFL/KIF20A, a kinesin involved with membrane trafficking of discs large homologue 5, can attenuate growth of pancreatic cancer cell. Cancer Res 65:105–112
Tessitore A, Gaggiano A, Cicciarelli G, Verzella D, Capece D, Fischietti M et al (2013) Serum biomarkers identification by mass spectrometry in high-mortality tumors. Int J Proteomics 2013:125858. doi:10.1155/2013/125858
Thedieck K, Holzwarth B, Prentzell MT, Boehlke C, Klasener K, Ruf S et al (2013) Inhibition of mTORC1 by astrin and stress granules prevents apoptosis in cancer cells. Cell 154:859–874. doi:10.1016/j.cell.2013.07.031
Ting DT, Lipson D, Paul S, Brannigan BW, Akhavanfard S, Coffman EJ et al (2011) Aberrant overexpression of satellite repeats in pancreatic and other epithelial cancers. Science 331:593–596. doi:10.1126/science.1200801
Tingstedt B, Johansson P, Andersson B, Andersson R (2007) Predictive factors in pancreatic ductal adenocarcinoma: role of the inflammatory response. Scand J Gastroenterol 42:754–759. doi:10.1080/00365520601058452
Verdone L, Caserta M, Di Mauro E (2005) Role of histone acetylation in the control of gene expression. Biochem Cell Biol 83:344–353. doi:10.1139/o05-041
Vincent A, Herman J, Schulick R, Hruban RH, Goggins M (2011) Pancreatic cancer. Lancet 378:607–620. doi:10.1016/S0140-6736(10)62307-0
Wilhelm M, Schlegl J, Hahne H, Gholami AM, Lieberenz M, Savitski MM et al (2014) Mass-spectrometry-based draft of the human proteome. Nature 509:582–587. doi:10.1038/nature13319
Yachida S, Jones S, Bozic I, Antal T, Leary R, Fu B et al (2010) Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467:1114–1117. doi:10.1038/nature09515
Acknowledgments
We wish to thank all subjects participating in this study for providing their clinical data and serum samples. We appreciate the assistance of Katarzyna Said Hilmersson for collecting samples and organizing the biobank. Protein sequencing was completed at Waters Corporation, Manchester, UK. We would also like to acknowledge the Kamprad Foundation, the Swedish Research Council, Vinnova, and the Crafoord Foundation.
Conflict of interest
JBC is employed by Waters Corporation, Manchester, UK. The remaining authors have no conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ansari, D., Andersson, R., Bauden, M.P. et al. Protein deep sequencing applied to biobank samples from patients with pancreatic cancer. J Cancer Res Clin Oncol 141, 369–380 (2015). https://doi.org/10.1007/s00432-014-1817-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-014-1817-x