Abstract
Although great efforts to characterize the embryonic phase of brain microvascular system development have been made, its postnatal maturation has barely been described. Here, we compared the molecular and functional properties of brain vascular cells on postnatal day (P)5 vs. P15, via a transcriptomic analysis of purified mouse cortical microvessels (MVs) and the identification of vascular-cell-type-specific or -preferentially expressed transcripts. We found that endothelial cells (EC), vascular smooth muscle cells (VSMC) and fibroblasts (FB) follow specific molecular maturation programs over this time period. Focusing on VSMCs, we showed that the arteriolar VSMC network expands and becomes contractile resulting in a greater cerebral blood flow (CBF), with heterogenous developmental trajectories within cortical regions. Samples of the human brain cortex showed the same postnatal maturation process. Thus, the postnatal phase is a critical period during which arteriolar VSMC contractility required for vessel tone and brain perfusion is acquired and mature.






Similar content being viewed by others
Data availability
Transcriptomic data are openly available in the GEO repository (www.ncbi.nlm.nih.gov/geo/) under the accession number GSE173844. Oher data that support the findings of this study are available on request from the corresponding author.
References
Anderson AW, Marois R, Colson ER, Peterson BS, Duncan CC, Ehrenkranz RA, Schneider KC, Gore JC, Ment LR (2001) Neonatal auditory activation detected by functional magnetic resonance imaging. Magn Reson Imaging 19(1):1–5. https://doi.org/10.1016/s0730-725x(00)00231-9
Ando K, Tong L, Peng D, Vazquez-Liebanas E, Chiyoda H, He L, Liu J, Kawakami K, Mochizuki N, Fukuhara S, Grutzendler J, Betsholtz C (2022) KCNJ8/ABCC9-containing K-ATP channel modulates brain vascular smooth muscle development and neurovascular coupling. Dev Cell 57(11):1383–1399. https://doi.org/10.1016/j.devcel.2022.04.019
Arichi T, Moraux A, Melendez A, Doria V, Groppo M, Merchant N, Combs S, Burdet E, Larkman DJ, Counsell SJ, Beckmann CF, Edwards AD (2010) Somatosensory cortical activation identified by functional MRI in preterm and term infants. Neuroimage 49(3):2063–2071. https://doi.org/10.1016/j.neuroimage.2009.10.038
Attwell D, Mishra A, Hall CN, O’Farrell FM, Dalkara T (2016) What is a pericyte? J Cereb Blood Flow Metab 36(2):451–455. https://doi.org/10.1177/0271678X15610340
Bankhead P, Loughrey MB, Fernandez JA, Dombrowski Y, McArt DG, Dunne PD, McQuaid S, Gray RT, Murray LJ, Coleman HG, James JA, Salto-Tellez M, Hamilton PW (2017) QuPath: open source software for digital pathology image analysis. Sci Rep 7(1):16878. https://doi.org/10.1038/s41598-017-17204-5
Ben-Zvi A, Liebner S (2021) Developmental regulation of barrier- and non-barrier blood vessels in the CNS. J Intern Med. https://doi.org/10.1111/joim.13263
Bondjers C, He L, Takemoto M, Norlin J, Asker N, Hellstrom M, Lindahl P, Betsholtz C (2006) Microarray analysis of blood microvessels from PDGF-B and PDGF-Rbeta mutant mice identifies novel markers for brain pericytes. FASEB J 20(10):1703–1705
Bonney SK, Sullivan LT, Cherry TJ, Daneman R, Shih AY (2022) Distinct features of brain perivascular fibroblasts and mural cells revealed by in vivo two-photon imaging. J Cereb Blood Flow Metab 42(6):966–978. https://doi.org/10.1177/0271678X211068528
Boulay AC, Saubamea B, Decleves X, Cohen-Salmon M (2015) Purification of Mouse Brain Vessels. J vis Exp. https://doi.org/10.3791/53208
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R (2018) Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36(5):411–420. https://doi.org/10.1038/nbt.4096
Chalouhi N, Ali MS, Jabbour PM, Tjoumakaris SI, Gonzalez LF, Rosenwasser RH, Koch WJ, Dumont AS (2012) Biology of intracranial aneurysms: role of inflammation. J Cereb Blood Flow Metab 32(9):1659–1676. https://doi.org/10.1038/jcbfm.2012.84
Chasseigneaux S, Moraca Y, Cochois-Guegan V, Boulay AC, Gilbert A, Le Crom S, Blugeon C, Firmo C, Cisternino S, Laplanche JL, Curis E, Decleves X, Saubamea B (2018) Isolation and differential transcriptome of vascular smooth muscle cells and mid-capillary pericytes from the rat brain. Sci Rep 8(1):12272. https://doi.org/10.1038/s41598-018-30739-5
Coelho-Santos V, Shih AY (2020) Postnatal development of cerebrovascular structure and the neurogliovascular unit. Wiley Interdiscip Rev Dev Biol 9(2):e363. https://doi.org/10.1002/wdev.363
Coelho-Santos V, Berthiaume AA, Ornelas S, Stuhlmann H, Shih AY (2021) Imaging the construction of capillary networks in the neonatal mouse brain. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.2100866118
Colonnese MT, Phillips MA, Constantine-Paton M, Kaila K, Jasanoff A (2008) Development of hemodynamic responses and functional connectivity in rat somatosensory cortex. Nat Neurosci 11(1):72–79. https://doi.org/10.1038/nn2017
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. https://doi.org/10.1038/nprot.2008.211
Daneman R, Zhou L, Kebede AA, Barres BA (2010) Pericytes are required for blood-brain barrier integrity during embryogenesis. Nature 468(7323):562–566. https://doi.org/10.1038/nature09513
Detre JA, Rao H, Wang DJ, Chen YF, Wang Z (2012) Applications of arterial spin labeled MRI in the brain. J Magn Reson Imaging 35(5):1026–1037. https://doi.org/10.1002/jmri.23581
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21. https://doi.org/10.1093/bioinformatics/bts635
Dore-Duffy P (2008) Pericytes: pluripotent cells of the blood brain barrier. Curr Pharm Des 14(16):1581–1593
Dorrier CE, Jones HE, Pintaric L, Siegenthaler JA, Daneman R (2022) Emerging roles for CNS fibroblasts in health, injury and disease. Nat Rev Neurosci 23(1):23–34. https://doi.org/10.1038/s41583-021-00525-w
Ezan P, Andre P, Cisternino S, Saubamea B, Boulay AC, Doutremer S, Thomas MA, Quenech’du N, Giaume C, Cohen-Salmon M (2012) Deletion of astroglial connexins weakens the blood-brain barrier. J Cereb Blood Flow Metab 32(8):1457–1467. https://doi.org/10.1038/jcbfm.2012.45
Frosen J, Joutel A (2018) Smooth muscle cells of intracranial vessels: from development to disease. Cardiovasc Res 114(4):501–512
Hadjiargyrou M (2018) Mustn1: A Developmentally regulated pan-musculoskeletal cell marker and regulatory gene. Int J Mol Sci. https://doi.org/10.3390/ijms19010206
Hall CN, Reynell C, Gesslein B, Hamilton NB, Mishra A, Sutherland BA, O’Farrell FM, Buchan AM, Lauritzen M, Attwell D (2014) Capillary pericytes regulate cerebral blood flow in health and disease. Nature 508(7494):55–60. https://doi.org/10.1038/nature13165
Hartmann DA, Underly RG, Grant RI, Watson AN, Lindner V, Shih AY (2015) Pericyte structure and distribution in the cerebral cortex revealed by high-resolution imaging of transgenic mice. Neurophotonics 2(4):041402
Hartmann DA, Berthiaume AA, Grant RI, Harrill SA, Koski T, Tieu T, McDowell KP, Faino AV, Kelly AL, Shih AY (2021) Brain capillary pericytes exert a substantial but slow influence on blood flow. Nat Neurosci 24(5):633–645
He L, Vanlandewijck M, Raschperger E, Andaloussi Mae M, Jung B, Lebouvier T, Ando K, Hofmann J, Keller A, Betsholtz C (2016) Analysis of the brain mural cell transcriptome. Sci Rep 6:35108. https://doi.org/10.1038/srep35108
Hellstrom M, Kalen M, Lindahl P, Abramsson A, Betsholtz C (1999) Role of PDGF-B and PDGFR-beta in recruitment of vascular smooth muscle cells and pericytes during embryonic blood vessel formation in the mouse. Development 126(14):3047–3055
Huang DW, Sherman BT, Tan Q, Kir J, Liu D, Bryant D, Guo Y, Stephens R, Baseler MW, Lane HC, Lempicki RA (2007) DAVID bioinformatics resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. https://doi.org/10.1093/nar/gkm415
Iadecola C, Nedergaard M (2007) Glial regulation of the cerebral microvasculature. Nat Neurosci 10(11):1369–1376. https://doi.org/10.1038/nn2003
Kelly KK, MacPherson AM, Grewal H, Strnad F, Jones JW, Yu J, Pierzchalski K, Kane MA, Herson PS, Siegenthaler JA (2016) Col1a1+ perivascular cells in the brain are a source of retinoic acid following stroke. BMC Neurosci 17(1):49. https://doi.org/10.1186/s12868-016-0284-5
Kozberg MG, Chen BR, DeLeo SE, Bouchard MB, Hillman EM (2013) Resolving the transition from negative to positive blood oxygen level-dependent responses in the developing brain. Proc Natl Acad Sci U S A 110(11):4380–4385. https://doi.org/10.1073/pnas.1212785110
Kranias EG, Hajjar RJ (2012) Modulation of cardiac contractility by the phospholamban/SERCA2a regulatome. Circ Res 110(12):1646–1660. https://doi.org/10.1161/CIRCRESAHA.111.259754
Kusaka T, Kawada K, Okubo K, Nagano K, Namba M, Okada H, Imai T, Isobe K, Itoh S (2004) Noninvasive optical imaging in the visual cortex in young infants. Hum Brain Mapp 22(2):122–132. https://doi.org/10.1002/hbm.20020
Lendahl U, Nilsson P, Betsholtz C (2019) Emerging links between cerebrovascular and neurodegenerative diseases-a special role for pericytes. EMBO Rep 20(11):e48070. https://doi.org/10.15252/embr.201948070
Manberg A, Skene N, Sanders F, Trusohamn M, Remnestal J, Szczepinska A, Aksoylu IS, Lonnerberg P, Ebarasi L, Wouters S, Lehmann M, Olofsson J, von Gohren AI, Domaniku A, De Schaepdryver M, De Vocht J, Poesen K, Uhlen M, Anink J, Mijnsbergen C, Vergunst-Bosch H, Hubers A, Klappe U, Rodriguez-Vieitez E, Gilthorpe JD, Hedlund E, Harris RA, Aronica E, Van Damme P, Ludolph A, Veldink J, Ingre C, Nilsson P, Lewandowski SA (2021) Altered perivascular fibroblast activity precedes ALS disease onset. Nat Med 27(4):640–646. https://doi.org/10.1038/s41591-021-01295-9
Nakagomi T, Nakano-Doi A, Kawamura M, Matsuyama T (2015) Do Vascular pericytes contribute to neurovasculogenesis in the central nervous system as multipotent vascular stem cells? Stem Cells Dev. https://doi.org/10.1089/scd.2015.0039
Nelson PT, Jicha GA, Wang WX, Ighodaro E, Artiushin S, Nichols CG, Fardo DW (2015) ABCC9/SUR2 in the brain: Implications for hippocampal sclerosis of aging and a potential therapeutic target. Ageing Res Rev 24(Pt B):111–125. https://doi.org/10.1016/j.arr.2015.07.007
Nippert AR, Biesecker KR, Newman EA (2018) Mechanisms mediating functional hyperemia in the brain. Neuroscientist 24(1):73–83. https://doi.org/10.1177/1073858417703033
Noli L, Capalbo A, Ogilvie C, Khalaf Y, Ilic D (2015) Discordant growth of monozygotic twins starts at the blastocyst stage: a case study. Stem Cell Reports 5(6):946–953. https://doi.org/10.1016/j.stemcr.2015.10.006
Ollion J, Cochennec J, Loll F, Escude C, Boudier T (2013) TANGO: a generic tool for high-throughput 3D image analysis for studying nuclear organization. Bioinformatics 29(14):1840–1841. https://doi.org/10.1093/bioinformatics/btt276
Oudart M, Tortuyaux R, Mailly P, Mazare N, Boulay AC, Cohen-Salmon M (2020) Astrodot - a new method for studying the spatial distribution of mRNA in astrocytes. J Cell Sci. https://doi.org/10.1242/jcs.239756
Perrenoud Q, Rossier J, Ferezou I, Geoffroy H, Gallopin T, Vitalis T, Rancillac A (2012) Activation of cortical 5-HT(3) receptor-expressing interneurons induces NO mediated vasodilatations and NPY mediated vasoconstrictions. Front Neural Circuits 6:50. https://doi.org/10.3389/fncir.2012.00050
Renier N, Wu Z, Simon DJ, Yang J, Ariel P, Tessier-Lavigne M (2014) iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159(4):896–910. https://doi.org/10.1016/j.cell.2014.10.010
Rueden CT, Schindelin J, Hiner MC, DeZonia BE, Walter AE, Arena ET, Eliceiri KW (2017) Image J2: imagej for the next generation of scientific image data. BMC Bioinformatics 18(1):529. https://doi.org/10.1186/s12859-017-1934-z
Rungta RL, Chaigneau E, Osmanski BF, Charpak S (2018) Vascular compartmentalization of functional hyperemia from the synapse to the pia. Neuron 99(2):362–375. https://doi.org/10.1016/j.neuron.2018.06.012
Saunders A, Macosko EZ, Wysoker A, Goldman M, Krienen FM, de Rivera H, Bien E, Baum M, Bortolin L, Wang S, Goeva A, Nemesh J, Kamitaki N, Brumbaugh S, Kulp D, McCarroll SA (2018) Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174(4):1015–1030. https://doi.org/10.1016/j.cell.2018.07.028
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7):676–682. https://doi.org/10.1038/nmeth.2019
Sullender CT, Richards LM, He F, Luan L, Dunn AK (2022) Dynamics of isoflurane-induced vasodilation and blood flow of cerebral vasculature revealed by multi-exposure speckle imaging. J Neurosci Methods 366:109434. https://doi.org/10.1016/j.jneumeth.2021.109434
Supek F, Bosnjak M, Skunca N, Smuc T (2011) REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6(7):e21800. https://doi.org/10.1371/journal.pone.0021800
Taga G, Asakawa K, Maki A, Konishi Y, Koizumi H (2003) Brain imaging in awake infants by near-infrared optical topography. Proc Natl Acad Sci U S A 100(19):10722–10727. https://doi.org/10.1073/pnas.1932552100
Toth-Heyn P, Cataldi L (2012) Vasoactive compounds in the neonatal period. Curr Med Chem 19(27):4633–4639. https://doi.org/10.2174/092986712803306330
Uranishi R, Baev NI, Kim JH, Awad IA (2001) Vascular smooth muscle cell differentiation in human cerebral vascular malformations. Neurosurgery 49(3):671–679. https://doi.org/10.1097/00006123-200109000-00027
Vanlandewijck M, He L, Mae MA, Andrae J, Ando K, Del Gaudio F, Nahar K, Lebouvier T, Lavina B, Gouveia L, Sun Y, Raschperger E, Rasanen M, Zarb Y, Mochizuki N, Keller A, Lendahl U, Betsholtz C (2018) A molecular atlas of cell types and zonation in the brain vasculature. Nature 554(7693):475–480. https://doi.org/10.1038/nature25739
Wang J, Vasaikar S, Shi Z, Greer M, Zhang B (2017) WebGestalt 2017: a more comprehensive, powerful, flexible and interactive gene set enrichment analysis toolkit. Nucleic Acids Res 45(W1):W130–W137. https://doi.org/10.1093/nar/gkx356
Yu Y, Cai Z, Cui M, Nie P, Sun Z, Sun S, Chu S, Wang X, Hu L, Yi J, Shen L, He B (2015) The orphan nuclear receptor Nur77 inhibits low shear stress-induced carotid artery remodeling in mice. Int J Mol Med 36(6):1547–1555. https://doi.org/10.3892/ijmm.2015.2375
Zehendner CM, Tsohataridis S, Luhmann HJ, Yang JW (2013) Developmental switch in neurovascular coupling in the immature rodent barrel cortex. PLoS ONE 8(11):e80749. https://doi.org/10.1371/journal.pone.0080749
Zheng M, Cheng H, Banerjee I, Chen J (2010) ALP/Enigma PDZ-LIM domain proteins in the heart. J Mol Cell Biol 2(2):96–102. https://doi.org/10.1093/jmcb/mjp038
Acknowledgements
This work was funded by grants from the Association Européenne contre les Leucodystrophies (ELA) (ELA2012-014C2B), the Fondation pour la Recherche Médicale (FRM) (AJE20171039094) and the Fondation Maladies Rares (20170603). A. Gilbert’s PhD fellowship was funded by the FRM (PLP20170939025p60) and ELA (ELA2012-014C2B). L. Slaoui’s fellowship PhD was funded by the Ecole Normale Supérieure. A.-C. Boulay’s work was funded by the FRM (AJE20171039094) and the Foundation pour la recherche sur la sclérose en plaques (ARSEP). Despite our efforts, our work has not received any support from the French National Agency for Research (ANR).
Author information
Authors and Affiliations
Contributions
Conceptualization, MC-S. and A-CB; Methodology, MC-S, A-CB, PM, GL, AG, NR, DV, AR; Investigation, LS, AG, BD-P, MBL, LF, SC, AR, AC, QG, MF, KD, LJ, SA, A-CB, MC-S; Writing—Original Draft, MC-S; Funding Acquisition, MC-S.; Supervision, MC-S. and A-CB.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare no competing financial interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
429_2022_2592_MOESM1_ESM.xlsx
Supplementary Table S1 Dataset of the comparison of cortical MV transcriptomes on P5 and P15 Selected mRNAs have a mean number of reads ≥ 50 in at least one condition. The fold-change (FC) between expression on P5 and P15 and associated adjusted p-values (padj) are indicated. Base mean: mean reads for each transcript. n=3 libraries for each stage. file1 (XLSX 2599 KB)
429_2022_2592_MOESM2_ESM.xlsx
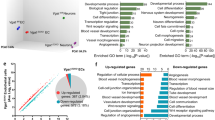
Supplementary Table S2 Pathway/Gene ontology data sets In cortical MVs, the “biological process” GO, KEGG and Reactome pathways significantly changed between P5 and P15. file2 (XLSX 69 KB)
429_2022_2592_MOESM3_ESM.xlsx
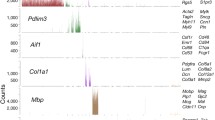
Supplementary Table S3 Identification of vascular-cell-type-specific or -preferentially expressed transcripts Cell-type-specific or -preferentially expressed transcripts are listed for each cluster. Left-hand columns: identification of transcripts -preferentially expressed in or specific to each cell. Middle columns: % of single-cells within and outside each cluster expressing the transcript logFC of expression difference between cells of the cluster and outside and the associated adjusted p-values. Right-hand columns: cortical MV RNA-Seq analysis. Log2FC on P15 versus P5 and the associated adjusted p-values. EC, endothelial cell; PC, pericyte; FB, fibroblast: VSMC, vascular smooth muscle cell file3 (XLSX 34 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Slaoui, L., Gilbert, A., Rancillac, A. et al. In mice and humans, brain microvascular contractility matures postnatally. Brain Struct Funct 228, 475–492 (2023). https://doi.org/10.1007/s00429-022-02592-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00429-022-02592-w