Abstract
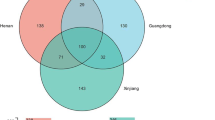
Vaginal fluid and saliva are of great importance in forensic sciences. The identification of vaginal fluid or saliva is especially important in criminal cases. Microbes are considered as a promising marker for the identification of body fluids. In this study, 18 salivary fluids and 18 vaginal fluid samples were collected from 18 healthy women of the Han population in Guangdong province, China. The microbes of the above samples were analyzed by 16S rDNA high-throughput sequencing. The results showed that the microbes whose proportions are over 1% in saliva samples distributed across 12 genera and 57 operational taxonomic units (OTUs), and in vaginal fluid distributed across 4 genera and 9 OTUs. The microbes that dominated in saliva were quite different from those dominated in vaginal fluids. The linear discriminant analysis (LDA) effect size (LEfSe) algorithm was used to screen out the specific microbes of the studied samples, and the results showed that the specific microbes in saliva samples are Haemophilus parainfluenzae, Veillonella parvula, and Aggregatibacter segnis, while in vaginal fluid is Lactobacillus iners.









Similar content being viewed by others
References
Sijen T (2015) Molecular approaches for forensic cell type identification: on mRNA, miRNA, DNA methylation and microbial markers. Forensic Sci Int Genet 18:21–32
Harbison SA, Fleming RI (2016) Forensic body fluid identification: state of the art. Res Rep Forensic Med Sci 6(2016):11–23. https://doi.org/10.2147/RRFMS.S57994
van den Berge M, Carracedo A, Gomes I, Graham EAM, Haas C, Hjort B, Hoff-Olsen P, Maroñas O, Mevåg B, Morling N, Niederstätter H, Parson W, Schneider PM, Court DS, Vidaki A, Sijen T (2014) A collaborative European exercise on mRNA-based body fluid/skin typing and interpretation of DNA and RNA results. Forensic Sci Int Genet 10:40–48
Haas C, Hanson E, Anjos MJ, Bär W, Banemann R, Berti A, Borges E, Bouakaze C, Carracedo A, Carvalho M, Castella V, Choma A, de Cock G, Dötsch M, Hoff-Olsen P, Johansen P, Kohlmeier F, Lindenbergh PA, Ludes B, Maroñas O, Moore D, Morerod ML, Morling N, Niederstätter H, Noel F, Parson W, Patel G, Popielarz C, Salata E, Schneider PM, Sijen T, Sviežena B, Turanská M, Zatkalíková L, Ballantyne J (2012) RNA/DNA co-analysis from blood stains--results of a second collaborative EDNAP exercise. Forensic Sci Int Genet 6(1):70–80
Haas C, Hanson E, Anjos MJ, Ballantyne KN, Banemann R, Bhoelai B, Borges E, Carvalho M, Courts C, de Cock G, Drobnic K, Dötsch M, Fleming R, Franchi C, Gomes I, Hadzic G, Harbison SA, Harteveld J, Hjort B, Hollard C, Hoff-Olsen P, Hüls C, Keyser C, Maroñas O, McCallum N, Moore D, Morling N, Niederstätter H, Noël F, Parson W, Phillips C, Popielarz C, Roeder AD, Salvaderi L, Sauer E, Schneider PM, Shanthan G, Court DS, Turanská M, van Oorschot RAH, Vennemann M, Vidaki A, Zatkalíková L, Ballantyne J (2014) RNA/DNA co-analysis from human menstrual blood and vaginal secretion stains: results of a fourth and fifth collaborative EDNAP exercise. Forensic Sci Int Genet 8(1):203–212
Haas C, Hanson E, Anjos MJ, Banemann R, Berti A, Borges E, Carracedo A, Carvalho M, Courts C, de Cock G, Dötsch M, Flynn S, Gomes I, Hollard C, Hjort B, Hoff-Olsen P, Hríbiková K, Lindenbergh A, Ludes B, Maroñas O, McCallum N, Moore D, Morling N, Niederstätter H, Noel F, Parson W, Popielarz C, Rapone C, Roeder AD, Ruiz Y, Sauer E, Schneider PM, Sijen T, Court DS, Sviežená B, Turanská M, Vidaki A, Zatkalíková L, Ballantyne J (2013) RNA/DNA co-analysis from human saliva and semen stains--results of a third collaborative EDNAP exercise. Forensic Sci Int Genet 7(2):230–239
Haas C, Hanson E, Banemann R, Bento AM, Berti A, Carracedo Á, Courts C, Cock GD, Drobnic K, Fleming R, Franchi C, Gomes I, Hadzic G, Harbison SA, Hjort B, Hollard C, Hoff-Olsen P, Keyser C, Kondili A, Maroñas O, McCallum N, Miniati P, Morling N, Niederstätter H, Noël F, Parson W, Porto MJ, Roeder AD, Sauer E, Schneider PM, Shanthan G, Sijen T, Syndercombe Court D, Turanská M, van den Berge M, Vennemann M, Vidaki A, Zatkalíková L, Ballantyne J (2015) RNA/DNA co-analysis from human skin and contact traces--results of a sixth collaborative EDNAP exercise. Forensic Sci Int Genet 16:139–147
Hanson EK, Lubenow H, Ballantyne J (2009) Identification of forensically relevant body fluids using a panel of differentially expressed microRNAs. Anal Biochem 387(2):303–314
Sauer E, Reinke AK, Courts C (2016) Differentiation of five body fluids from forensic samples by expression analysis of four microRNAs using quantitative PCR. Forensic Sci Int Genet 22:89–99
Zubakov D, Boersma AWM, Choi Y, van Kuijk PF, Wiemer EAC, Kayser M (2010) MicroRNA markers for forensic body fluid identification obtained from microarray screening and quantitative RT-PCR confirmation. Int J Legal Med 124(3):217–226
An JH, Choi A, Shin KJ, Yang WI, Lee HY (2013) DNA methylation-specific multiplex assays for body fluid identification. Int J Legal Med 127(1):35–43
Lee HY, Park MJ, Choi A, An JH, Yang WI, Shin KJ (2012) Potential forensic application of DNA methylation profiling to body fluid identification. Int J Legal Med 126(1):55–62
Bahn JH, Zhang Q, Li F, Chan TM, Lin X, Kim Y, Wong DTW, Xiao X (2015) The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem 61(1):221–230
Dong WW, Li HM, Qing XR, Huang DH, Li HG (2016) Identification and characterization of human testis derived circular RNAs and their existence in seminal plasma. Sci Rep 6:39080
Hanssen EN, Avershina E, Rudi K, Gill P, Snipen L (2017) Body fluid prediction from microbial patterns for forensic application. Forensic Sci Int Genet 30:10–17
Choi A, Shin KJ, Yang WI, Lee HY (2014) Body fluid identification by integrated analysis of DNA methylation and body fluid-specific microbial DNA. Int J Legal Med 128(1):33–41
Quaak FCA, van Duijn T, Hoogenboom J, Kloosterman AD, Kuiper I (2018 Sep) Human-associated microbial populations as evidence in forensic casework. Forensic Sci Int Genet 36:176–185
Penning R, Betz P (1992) Physical examination of the victim of alleged rape. Geburtshilf Frauenheilkd 52(1):59–61 German
Grunbaum BW (1989) Admissibility of biochemical analyses results from sexual assault evidence in the United States courts. Med Law 8(5):485–492
Jones S, Scott K, Lewis J, Davidson G, Allard JE, Lowrie C, McBride BM, McKenna L, Teppett G, Rogers C, Clayson N (2016) Baird a. DNA transfer through nonintimate social contact[J]. Sci Justice 56(2):90–95
Gönültaş BM, Sahin B (2018) Event locations in extra-familial child sexual molestation cases: the Istanbul example. Int J Offender Ther Comp Criminol 62(5):1164–1178
Seto MC, Babchishin KM, Pullman LE, McPhail IV (2015) The puzzle of intrafamilial child sexual abuse: a meta-analysis comparing intrafamilial and extrafamilial offenders with child victims. Clin Psychol Rev 39:42–57
Stoltenborgh M, van IJzendoorn MH, Euser EM, Bakermans-Kranenburg MJ (2011) A global perspective on child sexual abuse: meta-analysis of prevalence around the world. Child Maltreatment 16:79–101. https://doi.org/10.1177/1077559511403920
Benschop CC et al (2012) Vaginal microbial flora analysis by next generation sequencing and microarrays; can microbes indicate vaginal origin in a forensic context? Int J Legal Med 126(2):303–310
Nasidze I, Li J, Quinque D, Tang K, Stoneking M (2009) Global diversity in the human salivary microbiome. Genome Res 19(4):636–643
Kang JG, Kim SH, Ahn TY (2006) Bacterial diversity in the human saliva from different ages. J Microbiol 44(5):572–576
Ali MM, Shokry DA, Zaghloul HS, Rashed LA (2013) Nada MG.PCR applications in identification of saliva samples exposed to different conditions (streptococci detection based). Pak J Biol Sci 16(12):575–579
Nakanishi H, Ohmori T, Hara M, Takada A, Shojo H, Adachi N, Saito K (2011) A simple identification method of saliva by detecting Streptococcus salivarius using loop-mediated isothermal amplification. J Forensic Sci 56(Suppl 1):S158–S161
Fleming RI, Harbison S (2010) The use of bacteria for the identification of vaginal secretions. Forensic Sci Int Genet 4(5):311–315
Zou KN, Hu M, Huang JP, Zhou HG (2016) Identification of vaginal fluid using microbial signatures. Fa Yi Xue Za Zhi 32(4):254–256
Zou KN, Ren LJ, Ping Y, Ma K, Li H, Cao Y, Zhou HG, Wei YL (2016) Identification of vaginal fluid, saliva, and feces using microbial signatures in a Han Chinese population. J Forensic Legal Med 43:126–131
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
SILVA: https://www.arb-silva.de/
Amabebe E, Anumba DOC (2018 Jun 13) The vaginal microenvironment: the physiologic role of lactobacilli. Front Med (Lausanne) 5:181
Dan Z, Linlin M et al (2015) Diversity of Lactobacillus in vagina of vulvovaginal candidiasis. Zhonghua Yi Xue Za Zhi 95(13):1012–1016 Chinese
Petrova MI, Reid G, Vaneechoutte M, Lebeer S (2017) Lactobacillus iners: friend or foe?[J]. Trends Microbiol 25(3):182–191
Jakobsson T, Forsum U (2007) Laetobacillus iners:a marker of changes in the vaginal flora? J Clin Microbial 45(9):3145
Fettweis JM, Brooks JP, Serrano MG, Sheth NU, Girerd PH, Edwards DJ, Strauss JF, the Vaginal Microbiome Consortium, Jefferson KK, Buck GA (2014) Differences in vaginal microbiome in African American women versus women of European ancestry. Microbiology 160:2272–2282. https://doi.org/10.1099/mic.0.081034-0.
Cobo F, Jiménez G, Rodríguez-Granger J, Sampedro A, Aliaga-Martínez L, Navarro-Marí JM (2017) Clinical and microbiological findings of septic arthritis caused by Haemophilus parainfluenzae. Med Mal Infect 47(8):526–531. https://doi.org/10.1016/j.medmal.2017.08.002
Li J, Chen P, Li J, Gao X, Chen X, Chen J (2017) A new treatment of sepsis caused by veillonella parvula: a case report and literature review. J Clin Pharm Ther 42(5):649–652. https://doi.org/10.1111/jcpt.12559
Nørskov-Lauritsen N (2014) Classification, identification, and clinical significance of Haemophilus and Aggregatibacter species with host specificity for humans. Clin Microbiol Rev 27(2):214–240. https://doi.org/10.1128/CMR.00103-13.
Seerangaiyan K, van Winkelhoff AJ, Harmsen HJM, Rossen JWA, Winkel EG (2017) The tongue microbiome in healthy subjects and patients with intra-oral halitosis. J Breath Res 11(3):036010
Mason MR, Nagaraja HN, Camerlengo T, Joshi V, Kumar PS (2013) Deep sequencing identifies ethnicity-specific bacterial signatures in the oral microbiome. PLoS One 8:e77287. https://doi.org/10.1371/journal.pone.0077287
Adler CJ, Malik R, Browne GV, Norris JM (2016) Diet may influence the oral microbiome composition in cats. Microbiome 4(1):23. https://doi.org/10.1186/s40168-016-0169-y.
Lee CC, Tang JH et al (2015) Effect of meteorological and geographical factors on the epidemics of hand, foot, and mouth disease in island-type territory. East Asia Biomed Res Int 2015:805039
Belstrøm D, Sembler-Møller ML, Grande MA, Kirkby N, Cotton SL, Paster BJ, Twetman S, Holmstrup P (2018) Impact of Oral hygiene discontinuation on Supragingival and salivary microbiomes. JDR Clin Trans Res 3(1):57–64
Takeshita T, Kageyama S, Furuta M, Tsuboi H, Takeuchi K, Shibata Y, Shimazaki Y, Akifusa S, Ninomiya T, Kiyohara Y, Yamashita Y (2016) Bacterial diversity in saliva and oral health-related conditions: the Hisayama study. Sci Rep 6:22164. https://doi.org/10.1038/srep22164
Funding
This work was supported by the Natural Science Foundation of China (Grant no. 81501627) and Innovative training program for College Students (Grant no. 201612121083 and Grant no. 201812121123).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The study was approved by the Ethics Committee of Southern Medical University.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlight
• Both saliva and vaginal fluid from each participant in Guangdong province of China were collected and detected by 16S rDNA high-throughput sequencing for the study.
• The specific microbes in saliva samples of the studied population are Haemophilus parainfluenzae, Veillonella parvula, and Aggregatibacter segnis, while in vaginal fluid is Lactobacillus iners.
• Compared with other studies, the results showed that there may be some different specific microbes in the same kind of body fluid for different populations.
Rights and permissions
About this article
Cite this article
Huang, H., Yao, T., Wu, W. et al. Specific microbes of saliva and vaginal fluid of Guangdong Han females based on 16S rDNA high-throughput sequencing. Int J Legal Med 133, 699–710 (2019). https://doi.org/10.1007/s00414-018-1986-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-018-1986-2