Abstract
During meiosis, pairing of homologous chromosomes and their synapsis are essential prerequisites for normal male gametogenesis. Even limited autosomal asynapsis often leads to spermatogenic impairment, the mechanism of which is not fully understood. The present study was aimed at deliberately increasing the size of partial autosomal asynapsis and analysis of its impact on male meiosis. For this purpose, we studied the effect of t 12 haplotype encompassing four inversions on chromosome 17 on mouse autosomal translocation T(16;17)43H (abbreviated T43H). The T43H/T43H homozygotes were fully fertile in both sexes, while +/T43H heterozygous males, but not females, were sterile with meiotic arrest at late pachynema. Inclusion of the t 12 haplotype in trans to the T43H translocation resulted in enhanced asynapsis of the translocated autosome, ectopic phosphorylation of histone H2AX, persistence of RAD51 foci, and increased gene silencing around the translocation break. Increase was also on colocalization of unsynapsed chromatin with sex body. Remarkably, we found that transcriptional silencing of the unsynapsed autosomal chromatin precedes silencing of sex chromosomes. Based on the present knowledge, we conclude that interference of meiotic silencing of unsynapsed autosomes with meiotic sex chromosome inactivation is the most likely cause of asynapsis-related male sterility.
Similar content being viewed by others
Introduction
Prophase of the first meiotic division is initiated by formation of DNA double-strand breaks (DSBs) at leptotene stage and continues in zygotene with genetic recombination and gradual building of the synaptonemal complex. This multiprotein scaffold (Moses 1969) mediates pairing and synapsis of homologous chromosomes culminating at pachynema (Page and Hawley 2004). Chromosomes and chromosomal parts lacking a pairing partner remain unsynapsed and are transcriptionally silenced. In normal meiosis, the unsynapsed chromatin is restricted to sex chromosomes that pair solely in the short pseudoautosomal homologous region. The asynapsed sex chromosomes in pachytene spermatocytes are morphologically manifested as sex body, a separate nuclear compartment comprised of condensed X and Y chromosomes (Solari 1974; McKee and Handel 1993).
The transcriptional silencing of sex chromosomes, called meiotic sex chromosome inactivation (MSCI), first proposed by Monesi (1965), is accompanied by substantial chromatin remodeling and embarkation of specific histone variants (Motzkus et al. 1999; Turner et al. 2006; van der Heijden et al. 2007; Baarends et al. 2005). The testes-abundant histone H2AX and its phosphorylated form γH2AX play an indispensable role in this process (Mahadevaiah et al. 2001; Redon et al. 2002). At early leptotene over 200 DSBs are induced throughout the genome by the topoisomerase II-like enzyme SPO11 (Keeney 2001) turning on phosphorylation of H2AX on Ser-139 in the vicinity of DNA lesions to form γH2AX (Rogakou et al. 1998). The phosphorylation is performed by ATM kinase and eventually leads to recruitment of other members of the DNA repair machinery. As the DSBs are successively repaired, the γH2AX gradually disappears from the chromatin. Later, during zygotene–pachytene transition, the second wave of massive H2AX phosphorylation takes place on unsynapsed sex chromosomes (Mahadevaiah et al. 2001). This time, the phosphorylation is performed by ataxia telangiectasia and Rad3 related (ATR) kinase (Turner et al. 2004; Bellani et al. 2005) apparently recruited to the asynapsed chromosomes by tumor suppressor protein BRCA1 (Turner et al. 2004). Thus, γH2AX is involved in both DSB repair and silencing of asynapsed chromatin (Fernandez-Capetillo et al. 2003). All these events are crucial for subsequent sex body formation accompanied by transcriptional silencing of the sex chromatin.
The existence and essential role of MSCI was first proposed in the hypothesis based on the sterilizing effect of X-autosomal translocations by Lifschytz and Lindsley (1972) and further extended by Forejt to explain autosomal, male-sterile rearrangements in mice and men (Forejt 1982, 1985, 1996). More recently, the chromosomally induced impairment of MSCI was studied on XYY males and male carriers of the X-autosome translocation T(X;16)16H (Turner et al. 2005; Royo et al. 2010). In these models, segments of Chrs Y and X escaped inactivation because of aberrant synapsis and pachytene spermatocytes with impaired MSCI then underwent stage IV apoptosis (see Ahmed and de Rooij (2009) for staging mouse seminiferous tubules cross sections). Although the importance of MSCI for progression of spermatogenesis has been well established (Fernandez-Capetillo et al. 2003), its biological role is not entirely understood. MSCI might have evolved for several reasons: (a) to mask the unsynapsed parts of sex chromosomes and prevent their recognition by pachytene checkpoint (Jablonka and Lamb 1988; Odorisio et al. 1998), (b) to preclude the meiotic recombination between non-homologous DNA sequences (McKee and Handel 1993), or (c) to hinder transcription from damaged DNA (Inagaki et al. 2010). Moreover, the silencing of sex-linked genes might also be important due to (d) their “toxicity” for spermatogenesis (Royo et al. 2010; Lifschytz and Lindsley 1972). MSCI is evolutionarily an older phenomenon than X-inactivation in somatic tissues. Besides mammals, MSCI was identified in opossum (Namekawa et al. 2007; Hornecker et al. 2007), in Z chromosome of female birds (Schoenmakers et al. 2009), and in nematode worm (Kelly et al. 2002). The controversy persists about MSCI in fruit fly (Vibranovski et al. 2009; Hense et al. 2007—but see Mikhaylova and Nurminsky 2011; Meiklejohn et al. 2011). Important differences were reported between mouse, opossum, and chicken in chronological order of the silencing relative to chromosomal synapsis. Unlike in mouse, where MSCI is concordant with asynapsis of sex chromosomes, MSCI in chicken and opossum precedes the chromosomal synapsis (Schoenmakers et al. 2009; Namekawa et al. 2007). Early beginning of MSCI in the latter species suggests that it might be triggered by a homology search mechanism rather than by asynapsis (Namekawa and Lee 2009).
Previously, it was found that BRCA1, ATR, and γH2AX are associated not only with sex chromosomes but also with unsynapsed autosomes in the process called meiotic silencing of unsynapsed chromatin (MSUC) (Turner et al. 2005; Baarends et al. 2005; Manterola et al. 2009). It parallels the meiotic silencing of unpaired DNA in filamentous fungus Neurospora crassa (Shiu et al. 2001); however, the mechanism is different. MSUC acts in male and female meiosis, and several histone modifications were found in asynapsed autosomal and sex chromatin, namely H2A ubiquitination (Baarends et al. 2005), H3-K9 deacetylation and dimethylation (Khalil et al. 2004) or incorporation of specific H3.3 histone variant (van der Heijden et al. 2007). In this perspective, MSCI can be considered a specific case of MSUC. Gene knockouts leading to extensive autosomal asynapsis, such as Spo11 −/−, can cause sequestration of the silencing machinery to a fraction of asynapsed autosomes and creation of pseudo-sex body accompanied by aberrant expression of sex chromosomes and stage IV pachytene arrest (Mahadevaiah et al. 2008). We reasoned that even the MSUC brought about by partial asynapsis of a rearranged autosome might be detrimental to male meiosis due to the interference with X-chromosome inactivation (Forejt 1982, 1984, 1985, 1996). According to an alternative, mutually not exclusive interpretation, persistent DNA DSBs per se (Inagaki et al. 2010; Lemmens and Tijsterman 2011) or silencing of essential spermatogenesis genes (Turner et al. 2005) might be detrimental to meiosis. Previously we have shown that silencing of genes in the unsynapsed portion of the rearranged mouse chromosome 17 (Chr 17) involved in T(16;17)43H translocation (hereafter T43H) is accompanied by incomplete silencing of Chr X in mid–late pachytene spermatocytes (Homolka et al. 2007). Here we used t 12 haplotype, a naturally occurring variant of chromosome 17, spanning over 25 Mb in the centromeric end of mouse Chr 17 and carrying four tandem inversions (Lyon 2005; Bauer et al. 2007), to deliberately extend the interval of asynapsis caused by T43H. Evidence is presented for the first time, on positive correlation between the extent of asynapsis and MSUC, aberrant MSCI responses, and stringency of the pachytene block. Moreover, we show that MSUC of Chr 17 also occurs in pachytene oocytes, without apparently affecting female fertility. Remarkably, MSUC of the asynapsed part of Chr 17 precedes the inactivation of sex chromosomes and leads to frequent intrusion of partially asynapsed translocation quadrivalent into the sex body. The latter phenomenon coincides with the escape of X-linked genes from transcriptional inactivation and indicates interference of MSUC with MSCI as the most likely cause of asynapsis-related male-limited sterility.
Results
Meiotic consequences of genetically enhanced autosomal asynapsis
More than three decades ago we disclosed non-random association of rearranged autosomes with XY chromosomes in pachytene-metaphase I spermatocytes of sterile male carriers of some chromosome translocations (Forejt 1974; Forejt and Gregorova 1977; Forejt et al. 1981). We hypothesized that incomplete meiotic synapsis and intrusion of autosomal chromatin into the forming sex body could interfere with meiotic sex chromosome inactivation and result in meiotic arrest (Forejt 1982, 1985, 1996). More recently, we proved transcriptional downregulation of genes in the unsynapsed region of the translocated Chr 17 and dysregulation of the X-linked genes in pachytene spermatocytes of sterile males heterozygous for T(16;17)43H translocation (hereafter T43H) (Homolka et al. 2007).
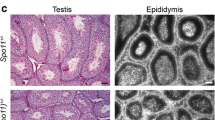
Here we asked whether extending partial asynapsis of the translocation quadrivalent can influence the sterilizing effect of the translocation and what molecular mechanisms are associated with asynapsis-driven male sterility. To extend the unsynapsed chromatin, we used the t 12 haplotype harboring four adjacent non-overlapping inversions that encompass over 25 Mb proximal to the T43H translocation breakpoint (Schimenti 2000; Lyon 2005; Bauer et al. 2007). The male translocation heterozygotes with or without the t 12 haplotype were completely sterile when outcrossed to C57BL/6J (B6), but their female sibs of the same genotype were fertile (Fig. 1a and Table S1). The t 12 haplotype significantly influenced spermatogenic block caused by T43H. The mean relative testes weight, a sensitive indicator of early spermatogenic failure, was significantly decreased in the presence of t 12 (0.25 in t 12 +/+T43H males, 0.34 in T+/+T43H males, P < 2.2e−16; t test). These sterile genotypes had about two-fold lower relative testes weight compared to fertile controls (T/t 12 × B10) t 12/+, T43H/T43H, and B6, 0.83, 0.69, and 0.72, respectively (Fig. 1b and Table S2). Testicular histology showed incomplete spermatogenic block with significantly reduced numbers of round and elongated spermatids in T+/+T43H males (Fig. 1c) while the t 12 +/+T43H males displayed complete lack of postmeiotic cell types, smaller tubule diameter and no progression beyond epithelial stage IV (Fig. 1c).
T43H males with t 12 haplotype display stronger meiotic arrest in pachynema. a Scheme of translocation quadrivalents at pachytene stage in the T+/+T43H (T/T43H) and t 12 +/+T43H (t 12/T43H) males. The t 12 haplotype with four genomic inversions (blue arrows) and the T43H translocation break are situated within the centromeric 30 Mb of Chr 17. The translocation break on Chr 16 localizes to centromeric chromatin. Chr 17 is shown in red, Chr 16 in blue. Centromeres are shown as circles. b Comparison of relative testes weights. The difference between T+/+T43H and t 12 +/+T43H males is significant (***P < 2.2e−16; t test). c The histological sections of seminiferous tubules demonstrate predominant pachytene arrest at epithelial stage IV in both T+/+T43H and t 12 +/+T43H males and complete loss of postmeiotic cells in t 12 +/+T43H males. eS elongated spermatids, rS round spermatids, aPS apoptotic pachytene spermatocytes. Scale bar = 50 μm
To examine the proportions of individual cell types, we prepared chromosome spreads from testicular cell suspensions and immunostained them for three markers of meiotic prophase I, namely synaptonemal complex protein 3 (SYCP3), a structural component of axial and lateral parts of synaptonemal complex; γH2AX, the phosphorylated histone variant, a mark of DSBs in leptotene/zygotene stage and of unsynapsed chromosomes in late pachytene spermatocytes (Mahadevaiah et al. 2001); and H1t, the testis-specific histone variant first appearing in mid–late pachytene spermatocytes (Inselman et al. 2003). Based on the patterns of these markers and DAPI staining of nuclei, we observed significantly lower frequency of late pachytene cells in sterile T+/+T43H and t 12 +/+T43H males when compared to the B6 and t 12 /+ fertile controls (0.18 and 0.12 vs 0.29 and 0.24, respectively, Fig. S1 and Table S3). The number of diplotene spermatocytes and postmeiotic cells in T+/+T43H males was further reduced compared to late pachynema. Only a few abnormal diplotene spermatocytes containing some fully synapsed homologs and no spermatids were present in t 12 +/+T43H cell spreads. In conclusion, the spermatogenic failure occurred in both genotypes of sterile males at mid–late pachytene stage, with more stringent pachytene arrest affecting t 12 +/+T43H males.
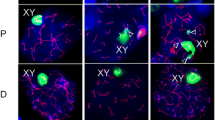
In the next step, we evaluated the effect of t 12 haplotype on pachytene synapsis of translocated Chr 17. To monitor synaptonemal complexes, we visualized SYCP3 and SYCP1 and immunostained γH2AX to mark unsynapsed chromatin at pachynema (Fig. 2a, b). In fertile males, phosphorylation of H2AX on sex chromosomes and formation of sex body is a gradual process, which begins at the late zygotene and is fully developed as a single nuclear domain at mid–late pachynema. In sterile males, we observed sex body and additional γH2AX domain covering the autosomal unsynapsed chromatin within the translocation quadrivalent. This autosomal γH2AX domain was observed in 81% (25 of 31) of pachytene nuclei of t 12 +/+T43H males and in 54% (41 of 76) pachynemas of T+/+T43H males (Fig. 2a, d) with the difference being statistically significant (P = 0.008; one-sided Fisher’s exact test). Strikingly, the two domains of the unsynapsed chromatin often colocalized and formed an enlarged γH2AX structure apparent in 68% and 38% (21 of 31 and 29 of 76) of pachynemas of t 12 +/+T43H and T+/+T43H males (Fig. 2c) (P = 0.005; one-sided Fisher’s exact test). Also immunostaining of SYCP3 and SYCP1 revealed increased frequency of pachynemas with unsynapsed translocated quadrivalent in the presence of t 12 in T43H translocation heterozygotes (Fig. 2b, d).
The abundance and the extent of autosomal asynapsis are elevated by t 12 haplotype in pachytene spermatocytes of T43H males. The local asynapsis of translocation quadrivalent was detected in mid–late pachytene spermatocytes with antibodies against the synaptonemal complex proteins and γH2AX. a The γH2AX (red) is restricted to the sex body (arrowheads) in spermatocytes of fertile males (B6, t 12/+). In T+/+T43H (T/T43H) and t 12+/+T43H (t 12/T43H) sterile males, it marks XY and, in addition the partially unsynapsed, translocated autosomes (arrows). The axial elements of the synaptonemal complex are detected by antibody against SYCP3 (green), and DNA is stained by DAPI (blue). b Co-immunostaining of SYCP1 (central element of SC, green) and SYCP3 (red) discriminates the unsynapsed chromatin associated only with SYCP3 from the synapsed chromosomes labeled by both antibodies (yellow). c The proportions of the T+/+T43H (n = 76) and t 12+/+T43H (n = 31) mid–late pachynemas with γH2AX restricted either to XY bivalent only, or to XY and T43H quadrivalent as two separate foci, or to XY and T43H as enlarged, fused focus. d The frequency of mid–late pachynemas with local asynapsis of the translocation quadrivalent estimated by two independent combinations of antibodies
Female oocytes were examined to find out whether the asynapsis per se might explain the male-limited character of sterility. In the majority of oocyte pachynema of T+/+T43H, the translocated quadrivalent was clearly asynapsed as detected by immunostaining of SYCP3 and CEN (Fig. S5). Thus, in spite of the autosomal asynapsis during oogenesis, the T43H does not affect female fertility, at least not in a manner comparable to spermatogenic failure of +/T43H heterozygotes (Table S1).
The repair of meiotic DSBs requires homologous synapsis because, unlike in somatic cells, in pachynema, the DSBs repair via sister chromatid recombination is suppressed (Inagaki et al. 2010; Lemmens and Tijsterman 2011). To find out whether T43H autosomal asynapsis affects DNA DSBs repair, we assessed the average frequency and distribution of RAD51 foci in H1t-positive mid–late pachytene nuclei (Fig. S2). Besides the sex chromosomes, a significant portion of persistent DSBs, 2.6 and 3.4 foci per nucleus resided within the unsynapsed arm of the translocation quadrivalent in T+/+T43H and t 12 +/+T43H males, respectively. The difference between both sterile genotypes was statistically significant (P = 0.016; t test). Also the proportion of the cells with at least one unrepaired DSB on translocated quadrivalent was significantly higher (P = 0.024; one-sided Fisher’s exact test) in t 12 +/+T43H males (91 of 150; 61%) than in T+/+T43H males (73 of 150; 49%).
Altogether, our results demonstrate that the t 12 haplotype increases the extent of meiotic asynapsis of T43H translocation, the recruitment of γH2AX to the translocated chromosome, and the association of unsynapsed autosomal chromatin with the sex body. The t 12 haplotype might also increase the frequency of persisting unrepaired DSBs seen as RAD51 foci in unsynapsed autosomal chromatin. The meiotic changes are accompanied by the absence of postmeiotic stages and complete arrest of spermatogenesis. Female oogenesis shows comparable cytological signs of MSUC affecting T43H quadrivalent, but without apparent effect on fertility.
Genetically extended asynapsis increases gene silencing
We searched for silenced chromatin domains by comparing transcription profiles of pre–mid-pachytene spermatocytes isolated by fluorescence-activated cell sorting (FACS) from sterile t 12 +/+T43H and T+/+T43H males and from fertile B10 controls using Affymetrix GeneChip MG_430 2.0. Based on control immunofluorescent staining with prophase I markers, the sorted cells consisted of leptotene, zygotene, and early pachytene spermatocytes with only traces of late pachynemas. Comparison of gene expression levels (relative to B10 controls) among individual chromosomes revealed the excess of downregulated genes on translocated Chr 17 in T+/+T43H (6%, 19 of 317 P < 0.05) and in t 12 +/+T43H (15.8%, 50 of 317) males. The gene silencing on Chr 17 was significant in T+/+T43H males (P < =0.011; one-tailed Fisher exact test) when compared to the rest of the genome with the exception of Chr 16 (P = 0.063), the other translocated chromosome involved in T43H. In the presence of t 12 haplotype, the abundance of downregulated genes on Chr 17 was even more obvious compared to all other chromosomes (P ≤ 7.82e−05) (Fig. 3a).
Genome-wide mRNA profiling reveals stronger MSUC response in the T43H males with t 12 haplotype. a Percentage of downregulated and upregulated genes in the population of pre–mid-pachytene spermatocytes is shown for Chrs 1 to 19 and Chr X. The T+/+T43H (T/T43H) and t 12+/+T43H (t 12/T43H) genotypes were compared to the fertile B10 controls and the proportion of genes with significantly dysregulated expression (P < 0.05) was estimated. Chr 17 (red) exhibits the highest proportion of downregulated genes in agreement with its asynapsed status. b Significantly dysregulated genes of Chr 17 tend to be more strongly downregulated in t 12+/+T43H males. Expression change is shown in logarithmic scale of base 2. c Distribution of expressed genes along Chr 17. The genes close to the translocation breakpoint (blue dashed line) are predominantly suppressed. Genes with significant expression differences are plotted in red (P < 0.01) and green (0.01 < P < 0.05)
To evaluate the effect of t 12 haplotype on the strength of silencing of Chr 17 genes, we selected all Chr 17 genes in T+/+T43H and t 12 +/+T43H males differentially expressed relative to B10 controls and plotted their log-fold changes (Fig. 3b). The silencing of Chr 17 was stronger in the presence of t 12 (P = 4.97e−06; paired t test), obviously in correlation with higher proportion of the cells with asynapsed Chr 17. The effect was confirmed by qRT-PCR for selected genes (Fig. S3). Downregulated genes predominantly clustered around the T43H translocation breakpoint at 30 Mb (Vacik et al. 2005), in agreement with the observed asynapsis in the region. However, the extent of downregulation differed between individual genes, perhaps due to the differences in local chromatin modification. When the sterile genotypes were compared, the t 12 +/+T43H showed more downregulated genes spread in the larger interval of proximal Chr 17 (Fig. 3c), but still predominantly located around the translocation break. We confirmed the effect of t 12 haplotype on silencing of Chr 17 genes also in the testes of 15 days postpartum (pp) males. The prepubertal testis contains spermatocytes of the first wave of spermatogenesis with mid-pachynema as the most advanced stage and only minor traces of late pachynemas. While downregulation of Chr 17 genes was clearly detected and more pronounced in the presence of t 12 haplotype (Fig. 4a), the X-linked genes were still active without signs of MSCI (Fig. 4b).
Downregulation of Chr 17 genes in the prepubertal testes of T43H males with or without t 12. a Percentage of significantly downregulated genes (P < 0.01) in the testes of 15 days pp males is shown for Chrs 1 to 19 and Chr X. The gene expression was compared between sterile T+/+T43H (T/T43H) and t 12 +/+T43H (t 12/T43H) males and fertile B6 controls. The abundance of downregulated genes on Chr 17 (red) is shown in T+/+T43H and t 12 +/+T43H males. b Comparison of average gene expression between autosomes and Chr X for all samples. Similar X-chromosomal and autosomal activity indicates absence of MSCI at 15 days pp
MSUC precedes and interferes with the MSCI
Downregulation of autosomal genes in unsynapsed chromatin at pre–mid-pachytene stage highlighted different timing of MSUC and MSCI, which is a suggested manifestation of MSUC on sex chromosomes (Turner et al. 2006). To get further insight into the chronological order of MSUC and MSCI during the first meiotic prophase, we compared the activity of autosomal and X-linked genes in the cell-sorted populations of pre–mid-pachytene spermatocytes with observed MSUC together with mid–late pachytene spermatocytes of B10 fertile males. Because of the early meiotic arrest in t 12 +/+T43H males, we could collect only pre–mid-pachytene cells from sterile genotypes. Gene expression profiling revealed that Chr X in the pre–mid-pachytene spermatocytes of translocation heterozygotes as well as of B10 controls was still active, in contrast to B10 mid–late pachynemas, which showed dramatic reduction of mRNA levels (Fig. 5).
Transcriptional silencing of Chr X does not occur in the population of pre–mid-pachytene spermatocytes. Boxplots of expressions of autosomal and X chromosomal genes are shown. Autosomal gene expressions are similar in the populations of pre–mid-pachytene spermatocytes for all samples of T+/+T43H (T/T43H), t 12 +/+T43H (t 12/T43H), and B10 males, as well as for the population of mid–late pachytene spermatocytes of B10 males. Average Chr X expression is similar to autosomes in all samples of pre–mid late pachytene spermatocytes, indicating its active state. Transcriptional silencing of Chr X occurs later, as seen in the population of mid–late pachytene spermatocytes of B10 males
To address the possibility that the results could be skewed by cellular heterogeneity, we validated MSUC by RNA fluorescence in situ hybridization (RNA FISH) on single cells. Tcp1 in the proximal part of Chr 17 (chr17:13,109,331–13,117,933) and Tsga2 located in close proximity of the T43H translocation breakpoint (chr17:31,391,965–31,414,301) were selected based on their high expression in the pre–mid-pachynema in fertile males and their apparent downregulation in T+/+T43H and t 12 +/+T43H sterile males. In accord with the microarray data, Tcp1 displayed positive signal in 100% of late zygotene to early pachytene nuclei of B6 and in 88% of t 12/+ samples, confirming its expression in fertile males at this stage. Significantly lower numbers of Tcp1-positive cells at the same meiotic stage were detected in sterile males, 44% in T+/+T43H (P = 4.75E−06; comparison to B6; one-sided Fisher’s exact test) and 28% in t 12 +/+T43H (P = 2.66E−08) (Fig. 6a). The observation is in agreement with MSUC initiation before mid-pachynema. Later, in mid–late pachynema, silencing still increased as slightly lower frequency of Tcp1-positive nuclei was observed: 36% for T+/+T43H (P = 2.39E−10) and 20% for t 12 +/+T43 (P = 6.62E−14). Similarly, for Tsga2, the signal was detectable only in 37% of late zygotene to early pachytene cells in T+/+T43H (P = 3.1E−06; compared to B6; one-sided Fisher’s exact test) and in 23% of t 12 +/+T43H samples (P = 1.37E−08), whereas the majority of the nuclei of fertile males were positive, 91% of B6 nuclei and 83% of t 12/+ nuclei (Fig. 6b). At mid–late pachynema, the number of positive nuclei in sterile males further decreased to 28% for T+/+T43H samples (P = 7.44E−10) and to 8% for t 12 +/+T43H genotype (P = 7.01E−15). Altogether, the data indicate that in contrast to γH2AX, which marks both unsynapsed T43H chromatin and XY chromosomes from late zygotene stage, transcriptional silencing observed by RNA FISH and mRNA expression profiling (Fig. 4a, b) is different, starting at late zygotene/early pachytene on Chr 17 and at mid–late pachytene stage on sex chromosomes. RNA FISH results are summarized in Table S4.
Tcp1, Tsga2, and Scml2 expression by RNA FISH of control and T43H pachytene spermatocytes. a Expression of Tcp1 (green), (chr17:13,109,331–13,117,933) in late zygotene–early pachytene (upper panel), and in mid–late pachytene cells (lower panel) of B6 and t 12/B10 fertile males and T+/+T43H (T/T43H) and t 12 +/+T43H (t 12/T43H) sterile males. b Expression of Tsga2 (chr17:31,391,965–31,414,301) (green). c Expression of Scml2 (chrX:157,555,125–157,696,145) (green). The substages of zygotene–pachytene and mid–late pachytene spermatocytes were discriminated by pattern of γH2AX immunostaining (red). At transition of late zygonema to early leptonema, the γH2AX disappears from autosomal bivalents and simultaneously elevates its content on the condensing sex chromatin; in mid–late pachynema, γH2AX is associated with the condensed chromatin of the sex body. However, it is completely absent on autosomes except of unsynapsed translocation quadrivalent in sterile males. The chromatin is stained by DAPI (blue). Positive RNA FISH signals are marked by arrows
To further verify the chronological order of MSUC and MSCI and to see whether autosomal transcriptional silencing interferes with sex chromosome inactivation, we analyzed the expression of the X-linked gene Scml2. As predicted, the FISH signal of Scml2 transcript was visible in the late zygotene–early pachytene cells in fertile and sterile males. Later, at mid–late pachytene stage, the fertile and sterile males differed. The Scml2 transcript was absent in all nuclei of fertile males, consistent with efficient MSCI, but present in 14% of T+/+T43H and 26% of t 12 +/+T43H mid–late pachynemas (Fig. 6c)—significant enrichment in comparison to B6 (P = 6.24E−03 for T+/+T43H and P = 4.99E−05 for t 12 +/+T43H; one-sided Fisher’s exact test). The signal was often found at the border or outside the γH2AX foci, indicating that parts of sex chromosomes might be devoid of this silencing mark. The finding indicates local MSCI impairment in a fraction of pachytene cells.
The extent of local MSCI failure in the late pachytene spermatocytes was also inferred from the Cot1 signal referring to the presence of nascent RNA transcripts. It was absent over the sex body of pachynemas in fertile males but overlapped a limited portion of the XY chromosomes in the majority of late pachytene spermatocytes of sterile males (Fig. S4).
Discussion
The relationship between autosomal asynapsis in meiotic prophase and male sterility has been known for decades, but the mechanism of sterilizing effect has remained elusive (Miklos 1974; Forejt et al. 1981). Asynapsis can reflect pairing difficulties of chromosomes involved in certain rearrangements, such as translocations, inversions, or other structural mutations, or can be caused by mutations in certain meiotic genes. The local consequences of asynapsis include persistence of DNA double-strand breaks (Schoenmakers et al. 2008), silencing of genes in the unsynapsed chromosomal segment—MSUC (Turner et al. 2005; Baarends et al. 2005), and intrusion of the rearranged autosome in the sex body (Forejt 1996). The ultimate physiological consequence of asynapsis consists in male subfertility or sterility due to partial or complete meiotic arrest at the first meiotic prophase. However, in many instances, as in multiple heterozygotes for Robertsonian translocations, pachytene block can be leaky or missing and spermatogenesis may fail at metaphase I or later (Forejt 1979; Manterola et al. 2009; Mahadevaiah et al. 2008; Burgoyne et al. 2009). In normal development, the only unsynapsed chromatin is that of the heterologous portions of male sex chromosomes, which are transcriptionally silenced by MSCI. Thus MSCI in spermatogenesis and Chr X reactivation in oogenesis are the two major epigenetic variables of male and female gametogenesis. We have argued that MSCI may function as a general male-specific surveillance mechanism disrupted by various unrelated male-sterile autosomal rearrangements in mice, humans, and other species. This hypothesis was based on repeated observations of asynapsed autosomes involved in male-sterile rearrangements, which colocalized with the sex body. The sex chromosomes displayed morphologically discernible changes of chromatin compaction (Forejt 1982, 1984; Forejt 1985, 1996). Possible alternative but not mutually exclusive explanations include MSUC affecting genes indispensable for male meiosis or a failure of MSCI due to asynapsis caused primarily by meiotic gene mutations (Mihola et al. 2009; Mahadevaiah et al. 2008; Burgoyne et al. 2009; Hayashi et al. 2005).
To bring a new insight into the possible mechanism of asynapsis-related male-specific meiotic arrest, we asked whether, by genetically increasing the size of an unsynapsed autosomal segment, we can enhance the spermatogenic arrest and which, if any, meiotic phenotypes will be affected. To enhance the asynapsis of the proximal part of Chr 17 in T43H translocation quadrivalent, we constructed a genotype with t 12 haplotype in trans to T43H. The effect of mouse autosomal reciprocal translocation T43H on spermatogenesis was described in some detail previously (Homolka et al. 2007; Forejt et al. 1980). The t-haplotypes are naturally occurring variants harboring four non-overlapping tandem genomic inversions spanning ~25 Mb of proximal Chr 17 (Schimenti 2000; Lyon 2005; Bauer et al. 2007). The t-inversions suppress genetic recombination in the region and prevent homologous synapsis at the sequence level as evidenced by the absence of inversion loops of synaptonemal complexes in pachytene nuclei (Tres and Erickson 1982). The normal male fertility and absence of unsynapsed chromatin in t 12/+ males can be explained by heterologous synapsis via synaptic adjustment (Henzel et al. 2011). Analyzing the γH2AX signal on translocated chromosomes, we found the asynapsis more frequent in the presence of t 12-haplotype. Moreover, the decreased mRNA levels of genes in the region indicated stronger MSUC response to the more pronounced pairing problems in the proximal part of Chr 17. In accord with other models of pachytene asynapsis (Burgoyne et al. 2009), the T43H heterozygotes revealed similar chronological order of H2AX phosphorylation on asynapsed autosomes and sex chromosomes, starting in late zygonema and gradually increasing until late pachynema. However, the transcriptional silencing was different. In contrast to the conspicuous downregulation of mRNA levels in the unsynapsed autosomal chromatin in early pachytene spermatocytes, the X-linked genes were normally active in the same cells. The conclusion based on the transcriptional profiling of FACS-sorted populations of primary spermatocytes was confirmed by RNA FISH experiments on individual cells. MSUC was confirmed at zygotene–early pachytene stages while the first signs of MSCI appeared later, at mid–late pachynema. The early commencement of MSUC reported here formally parallels timing of MSCI in the opossum Monodelphis domestica (Namekawa et al. 2007). The X and Y chromosomes of the opossum lack any obvious homology and show only limited association at mid-pachytene, though their silencing starts already at early pachynema before they come into contact. More recently, Z and W chromosomes in chicken oogenesis were also shown to undergo MSCI, which predates their synapsis (Schoenmakers et al. 2009). At present we cannot determine the exact start of MSUC with respect to the homologous synapsis at zygotene/early pachytene stages. If specific silencing begins already at zygotene, then the question arises how the MSUC can recognize chromosomal regions unable to synapse later in pachytene. Interestingly, the spermatogenesis fails at late pachytene, several days after formation of MSUC, showing that MSUC per se is an unlikely reason for elimination of late pachytene spermatocytes (see also Manterola et al. 2009). Moreover, we observed MSUC also in female oogenesis of T43H heterozygotes with no apparent effect on female fertility.
Our original disclosure of colocalization of sex chromosomes with translocated autosomes causing male-specific sterility (Forejt 1974) had been confirmed and extended for various male-sterile rearrangements in several mammalian species (Forejt and Gregorova 1977; de Boer and Groen 1974; Gabriel-Robez and Rumpler 1994; Oliver-Bonet et al. 2005; Sciurano et al. 2007). More recently, we linked the colocalization of the translocated autosome and the sex body with breakdown of MSCI (Homolka et al. 2007). In this study, we provide new evidence for spermatogenic impairment being proportional to the extent of asynapsis and to the strength of MSUC response. Both colocalization of the translocated chromosome with sex chromosomes and aberrant transcription of the X-linked gene responded to the presence of t 12-haplotype.
At present the MSCI is considered an important surveillance mechanism controlling the progression of pachytene spermatocytes into diplotene (Royo et al. 2010). We propose that its disturbance is the likely common cause of meiotic arrest in various unrelated male-sterile chromosomal rearrangements.
Materials and methods
Ethics statement
The principles of laboratory animal care obeyed the Czech Republic Act for Experimental Work with Animals (Decree No. 207/2004 Sb. and Acts Nos. 246/92 Sb. and 77/2004 Sb.), fully compatible with the corresponding EU regulations and standards, namely Council Directive 806/609/EEC and Appendix A of the Council of Europe Convention ETS123.
Mice
The mice of B10-T43/T43H congenic strain, carrying T(16;17)43H translocation on the genetic background of inbred strain C57BL/10ScSnPh (B10) (Forejt et al. 1980), were crossed with T/t 12 mice, carrying t 12 haplotype with four inversions in the proximal part of Chr 17 in trans to Brachyury (T) mutation also mapping within the t-complex region. The obtained progeny consisted of t 12 +/+T43H and T+/+T43H mice. All mice were kept at the Specific Pathogen-Free Facility of the Institute of Molecular Genetics, AS CR.
Fertility test and histology
To test the fertility of the T+/+T43H and t 12 +/+T43H males, two sexually mature (2-month-old) males of both genotypes were mated with 2-month-old C57BL/6J females for 3 months. Similarly, T+/+T43H and t 12 +/+T43H females were tested by mating with C57BL/6J males. The number of offspring per mating unit (OMU) was calculated. One OMU = 1 male mated to two females for 1 month. Wet weight of both testes was estimated to the nearest mg immediately after dissection. Testes of mice were dissected and fixed in Bouin’s solution (Sigma-Aldrich), embedded in paraffin, serially sectioned at 2 to 3 μm thickness, and mounted on slides. The periodic acid Schiff stained sections were observed in a Nikon Eclipse 200 microscope. The images were captured using a Penguin 150CL CCD color camera (Pixera) and processed with Adobe Photoshop (Adobe Systems).
Immunostaining of spread spermatocytes and RNA FISH
Testes were processed to obtain spread meiocytes either by isotonic or hypotonic protocol (Turner et al. 2005; Anderson et al. 1999). For immunocytochemistry, the spread nuclei were immunolabeled with the following antibodies: rat polyclonal anti-SYCP3 (Abcam, #15092), mouse monoclonal anti-SYCP1 (Abcam, #15087), guinea pig anti-histone linker H1t (Inselman et al. 2003), mouse monoclonal anti-γH2AX (Upstate, #05-636), human autoantibody from autoimmune serum anti-centromere (AB-Incorporated, #15-235), rabbit polyclonal antibody anti-SCP3 (Santa Cruz, #74569), rabbit polyclonal antibody anti-Rad51 (Santa Cruz, H-92, #8349), and the secondary antibodies: goat anti-Rabbit IgG-AlexaFluor488 (MolecularProbes, A-11034), goat anti-Mouse IgG-AlexaFluor568 (MolecularProbes, A-11031), goat anti-Rabbit IgG-AlexaFluor568 (MolecularProbes, A-11036), goat anti-Mouse IgG-AlexaFluor350 (MolecularProbes, A-21049), and goat anti-Guinea pig IgG-Cy3 (Chemicon, #AP108C).
RNA FISH was performed on spread meiocytes prepared by isotonic protocol (Turner et al. 2005): Individual probes for RNA FISH listed in Table S5 were generated by long range PCR (Expand Long Range dNTPack, Roche, cat# 11811002001) on BAC DNA carrying the genomic region of interest. The PCR products were labeled with biotin by a nick translation kit (Biotin-Nick Translation Mix, Roche, cat# 11745824910) according to the manufacturer’s instructions. RNA FISH was performed according to the protocol of Mahadevaiah et al. (2009). The immunocytochemistry was performed directly after RNA FISH, and the images were acquired. The images for immunostain and immunostain combined with RNA FISH were examined in a Nikon Eclipse 400 (Tokyo, Japan) microscope with motorized stage control using Plan Fluor objective, 60× (Nikon, MRH00601) and captured using a DS-QiMc monochrome CCD camera (Nikon) and NIS elements program. The images were adjusted with Adobe Photoshop CS software (Adobe Systems).
FACS characterization and isolation of spermatogenic populations
Spermatogenic populations were isolated as previously described (Homolka et al. 2007) using fluorescence-activated cell sorting according to Bastos et al. (2005). Briefly, the spermatogenic tubules of mice euthanized by cervical dislocation were incubated in enriched Krebs–Ringer bicarbonate medium (EKRB) with collagenase (0.5 mg/mL; Sigma) for 20 min at 32°C in a shaker. The tubules were filtered with a cell strainer (BD Falcon) and incubated with collagenase under the same conditions. The filtered suspension was washed twice by EKRB containing 1% FCS. Finally, the cells were diluted in 1 mL of EKRB with 1% FCS and stained with Hoechst 33342 (5 μg/mL) for 1 h at 32°C. Propidium iodide was added just before FACS analysis to a concentration of 2 μg/mL. Individual populations were sorted according to red and blue Hoechst emission (Bastos et al. 2005) directly into the RLT buffer of the RNeasy Micro isolation kit (QIAGEN) and RNA was isolated. Cell aliquots were sorted to EKRB medium for parallel immunofluorescence analysis. Based on SYCP3, SYCP1, and γH2AX immunostaining and morphology of the cells, we have determined the composition of FACS-isolated populations. The sorted population of pre–mid-pachytene spermatocytes consisted of leptotene, zygotene, and early pachytene spermatocytes. Three independent isolations were made for both B10-t 12 +/+T43H and B10-T+/+T43H pachytene spermatocytes. Contrary to our previous study (Homolka et al. 2007), the isolation of the mid–late pachytene spermatocytes in sufficient purity was impossible from males of B10-t 12 +/+T43H genetic background due to ~30% reduction of late pachytene spermatocytes.
Testicular RNA isolation from 15 days pp males
Testes of juvenile mice were dissected, testicular tunica was removed, and the testes were placed into RLT buffer of the RNeasy Mini isolation kit (QIAGEN) and homogenized using syringe and needle. RNA was then isolated according to the manufacturer’s instructions.
Microarray analysis
The total RNA (20–30 ng) was converted in cRNA using the Affymetrix Two-Cycle Target Labeling kit according to the manufacturer’s instructions (sorted cell populations) or using the Affymetrix 3′ IVT Express Kit (15 days pp testes). Affymetrix GeneChip Mouse Genome 430 2.0 Arrays were hybridized with cRNA. The data obtained for the cell populations were analyzed together with the previously hybridized samples from pre–mid-pachytene spermatocytes and mid–late pachytene spermatocytes of B10 males (Homolka et al. 2007). The data from the testis of 15 days pp males were analyzed independently but in a similar way. The analysis used Bioconductor (Gentleman et al. 2004) (http://www.bioconductor.org/) and the R project for statistical computing (version 2.12; http://www.r-project.org/). The probes were annotated to Entrez gene identifiers using the custom chip description file “mm430mmentrezg” version 10 (Dai et al. 2005), which is based on NCBI build 37. The data were normalized using gcRMA. We used Linear Models for Microarray Data Package, limma version 3.6 (Smyth 2005) for statistical evaluations of expression differences. A linear model was fitted for each gene in a given series of arrays by using the lmFit function. To rank the differential expression of genes, we applied the eBayes function of the empirical Bayes method. A correction for multiple testing was performed using the Benjamini and Hochberg false discovery rate method (Benjamini and Hochberg 1995). Genes were considered to be expressed if average expression in all samples was ≥100. To assess the significance of enrichment of downregulated genes on Chr 17, we compared their incidence on Chr 17 and on other chromosomes by a series of one-tailed Fisher exact tests. Chr Y was omitted because only seven genes mapped to the probe sets. The microarray dataset is deposited in the NCBI Gene Expression Omnibus (GEO) with series accession number GSE29177.
Real-time quantitative RT-PCR
Total RNA isolated from spermatogenic populations and whole testis of 15 days pp males was reverse-transcribed using Mu-MLV Reverse Transcriptase (Invitrogen). Quantitative real-time PCR was performed with the Light-Cycler DNA Fast Start Master SYBR green I kit (Roche) in a Light Cycler 2000 machine at T m = 61°C. The sequences of primers are in Table S6.
References
Ahmed EA, de Rooij DG (2009) Staging of mouse seminiferous tubule cross-sections. In: Keeney S (ed) Meiosis, vol 2, Methods Mol Biol (558). Humana, New York, pp 263–277
Anderson LK, Reeves A, Webb LM, Ashley T (1999) Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151(4):1569–1579
Baarends WM, Wassenaar E, van der Laan R, Hoogerbrugge J, Sleddens-Linkels E, Hoeijmakers JH, de Boer P, Grootegoed JA (2005) Silencing of unpaired chromatin and histone H2A ubiquitination in mammalian meiosis. Mol Cell Biol 25(3):1041–1053
Bastos H, Lassalle B, Chicheportiche A, Riou L, Testart J, Allemand I, Fouchet P (2005) Flow cytometric characterization of viable meiotic and postmeiotic cells by Hoechst 33342 in mouse spermatogenesis. Cytometry A 65(1):40–499
Bauer H, Veron N, Willert J, Herrmann BG (2007) The t-complex-encoded guanine nucleotide exchange factor Fgd2 reveals that two opposing signaling pathways promote transmission ratio distortion in the mouse. Genes Dev 21(2):143–147
Bellani MA, Romanienko PJ, Cairatti DA, Camerini-Otero RD (2005) SPO11 is required for sex-body formation, and Spo11 heterozygosity rescues the prophase arrest of Atm−/− spermatocytes. J Cell Sci 118(Pt 15):3233–3245
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate—a practical and powerful approach to multiple testing. J Royal Stat Soc Ser B-Methodol 57(1):289–300
Burgoyne PS, Mahadevaiah SK, Turner JM (2009) The consequences of asynapsis for mammalian meiosis. Nat Rev Genet 10(3):207–216
Dai M, Wang P, Boyd AD, Kostov G, Athey B, Jones EG, Bunney WE, Myers RM, Speed TP, Akil H, Watson SJ, Meng F (2005) Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res 33(20):e175
de Boer P, Groen A (1974) Fertility and meiotic behavior of male T70H tertiary trisomics of the mouse (Mus musculus). A case of preferential telomeric meiotic pairing in a mammal. Cytogenet Cell Genet 13(6):489–510
Fernandez-Capetillo O, Mahadevaiah SK, Celeste A, Romanienko PJ, Camerini-Otero RD, Bonner WM, Manova K, Burgoyne P, Nussenzweig A (2003) H2AX is required for chromatin remodeling and inactivation of sex chromosomes in male mouse meiosis. Dev Cell 4(4):497–508
Forejt J (1974) Nonrandom association between a specific autosome and X-chromosome in meiosis of the male mouse: possible consequence of the homologous centromeres separation. Cytogenet Cell Genet 13:369–383
Forejt J (1979) Meiotic studies of translocations causing male sterility in the mouse. II. Double heterozygotes for Robertsonian translocations. Cytogenet Cell Genet 23(3):163–170
Forejt J (1982) X-Y involvement in male sterility caused by autosome translocations—a hypothesis. In: Fraccaro M, Rubin B (eds) Genetic control of gamete production and function. Academic, New York, pp 135–151
Forejt J (1984) X-inactivation and its role in male sterility. In: Bennett M, Gropp A, Wolf U (eds) Chromosomes today, vol 8. George Allen and Unwin, London, pp 117–127
Forejt J (1985) Chromosomal and genic sterility of hybrid type in mice and men. Exp Clin Immunogenet 2(2):106–119
Forejt J (1996) Hybrid sterility in the mouse. Trends Genet 12(10):412–417
Forejt J, Gregorova S (1977) Meiotic studies of translocations causing male sterility in the mouse. I. Autosomal reciprocal translocations. Cytogenet Cell Genet 19(2–3):159–179
Forejt J, Capkova J, Gregorova S (1980) T(16: 17)43H translocation as a tool in analysis of the proximal part of chromosome 17 (including T–t gene complex) of the mouse. Genet Res 35(2):165–177
Forejt J, Gregorova S, Goetz P (1981) XY pair associates with the synaptonemal complex of autosomal male-sterile translocations in pachytene spermatocytes of the mouse (Mus musculus). Chromosoma 82(1):41–53
Gabriel-Robez O, Rumpler Y (1994) The meiotic pairing behaviour in human spermatocytes carrier of chromosome anomalies and their repercussions on reproductive fitness. I: Inversions and insertions. A European collaborative study. Ann Genet 37(1):3–10
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JY, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5(10):R80
Hayashi K, Yoshida K, Matsui Y (2005) A histone H3 methyltransferase controls epigenetic events required for meiotic prophase. Nature 438(7066):374–378
Hense W, Baines JF, Parsch J (2007) X chromosome inactivation during Drosophila spermatogenesis. PLoS Biol 5(10):e273
Henzel JV, Nabeshima K, Schvarzstein M, Turner BE, Villeneuve AM, Hillers KJ (2011) An asymmetric chromosome pair undergoes synaptic adjustment and crossover redistribution during Caenorhabditis elegans meiosis: implications for sex chromosome evolution. Genetics 187(3):685–699
Homolka D, Ivanek R, Capkova J, Jansa P, Forejt J (2007) Chromosomal rearrangement interferes with meiotic X chromosome inactivation. Genome Res 17(10):1431–1437
Hornecker JL, Samollow PB, Robinson ES, Vandeberg JL, McCarrey JR (2007) Meiotic sex chromosome inactivation in the marsupial Monodelphis domestica. Genesis 45(11):696–708
Inagaki A, Schoenmakers S, Baarends WM (2010) DNA double strand break repair, chromosome synapsis and transcriptional silencing in meiosis. Epigenetics 5(4):255–266
Inselman A, Eaker S, Handel MA (2003) Temporal expression of cell cycle-related proteins during spermatogenesis: establishing a timeline for onset of the meiotic divisions. Cytogenet Genome Res 103(3–4):277–284
Jablonka E, Lamb MJ (1988) Meiotic pairing constraints and the activity of sex chromosomes. J Theor Biol 133(1):23–36
Keeney S (2001) Mechanism and control of meiotic recombination initiation. Curr Top Dev Biol 52:1–53
Kelly WG, Schaner CE, Dernburg AF, Lee MH, Kim SK, Villeneuve AM, Reinke V (2002) X-chromosome silencing in the germline of C. elegans. Development 129(2):479–492
Khalil AM, Boyar FZ, Driscoll DJ (2004) Dynamic histone modifications mark sex chromosome inactivation and reactivation during mammalian spermatogenesis. Proc Natl Acad Sci U S A 101(47):16583–16587
Lemmens BB, Tijsterman M (2011) DNA double-strand break repair in Caenorhabditis elegans. Chromosoma 120(1):1–21
Lifschytz E, Lindsley DL (1972) The role of X-chromosome inactivation during spermatogenesis (Drosophila-allocycly-chromosome evolution-male sterility-dosage compensation). Proc Natl Acad Sci U S A 69(1):182–186
Lyon MF (2005) Elucidating mouse transmission ratio distortion. Nat Genet 37(9):924–925
Mahadevaiah SK, Turner JM, Baudat F, Rogakou EP, de Boer P, Blanco-Rodriguez J, Jasin M, Keeney S, Bonner WM, Burgoyne PS (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27(3):271–276
Mahadevaiah SK, Bourc’his D, de Rooij DG, Bestor TH, Turner JM, Burgoyne PS (2008) Extensive meiotic asynapsis in mice antagonises meiotic silencing of unsynapsed chromatin and consequently disrupts meiotic sex chromosome inactivation. J Cell Biol 182(2):263–276
Mahadevaiah SK, Costa Y, Turner JM (2009) Using RNA FISH to study gene expression during mammalian meiosis. In: Keeney S (ed) Meiosis, vol volume 2, cytological methods. Methods in molecular biology. Humana, New York, pp 433–444
Manterola M, Page J, Vasco C, Berrios S, Parra MT, Viera A, Rufas JS, Zuccotti M, Garagna S, Fernandez-Donoso R (2009) A high incidence of meiotic silencing of unsynapsed chromatin is not associated with substantial pachytene loss in heterozygous male mice carrying multiple simple robertsonian translocations. PLoS Genet 5(8):e1000625
McKee BD, Handel MA (1993) Sex chromosomes, recombination, and chromatin conformation. Chromosoma 102(2):71–80
Meiklejohn CD, Landeen EL, Cook JM, Kingan SB, Presgraves DC (2011) Sex chromosome-specific regulation in the Drosophila male germline but little evidence for chromosomal dosage compensation or meiotic inactivation. PLoS Biol 9(8):e1001126
Mihola O, Trachtulec Z, Vlcek C, Schimenti JC, Forejt J (2009) A mouse speciation gene encodes a meiotic histone H3 methyltransferase. Science 323(5912):373–375
Mikhaylova LM, Nurminsky DI (2011) Lack of global meiotic sex chromosome inactivation, and paucity of tissue-specific gene expression on the Drosophila X chromosome. BMC Biol 9:29
Miklos GL (1974) Sex–chromosome pairing and male fertility. Cytogenet Cell Genet 13(6):558–577
Monesi V (1965) Differential rate of ribonucleic acid synthesis in the autosomes and sex chromosomes during male meiosis in the mouse. Chromosoma 17(1):11–21
Moses MJ (1969) Structure and function of the synaptonemal complex. Genetics 61(1):41–51
Motzkus D, Singh PB, Hoyer-Fender S (1999) M31, a murine homolog of Drosophila HP1, is concentrated in the XY body during spermatogenesis. Cytogenet Cell Genet 86(1):83–88
Namekawa SH, Lee JT (2009) XY and ZW: is meiotic sex chromosome inactivation the rule in evolution? PLoS Genet 5(5):e1000493
Namekawa SH, VandeBerg JL, McCarrey JR, Lee JT (2007) Sex chromosome silencing in the marsupial male germ line. Proc Natl Acad Sci U S A 104(23):9730–9735
Odorisio T, Rodriguez TA, Evans EP, Clarke AR, Burgoyne PS (1998) The meiotic checkpoint monitoring synapsis eliminates spermatocytes via p53-independent apoptosis. Nat Genet 18(3):257–261
Oliver-Bonet M, Benet J, Sun F, Navarro J, Abad C, Liehr T, Starke H, Greene C, Ko E, Martin RH (2005) Meiotic studies in two human reciprocal translocations and their association with spermatogenic failure. Hum Reprod 20(3):683–688
Page SL, Hawley RS (2004) The genetics and molecular biology of the synaptonemal complex. Annu Rev Cell Dev Biol 20:525–558
Redon C, Pilch D, Rogakou E, Sedelnikova O, Newrock K, Bonner W (2002) Histone H2A variants H2AX and H2AZ. Curr Opin Genet Dev 12(2):162–169
Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM (1998) DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem 273(10):5858–5868
Royo H, Polikiewicz G, Mahadevaiah SK, Prosser H, Mitchell M, Bradley A, de Rooij DG, Burgoyne PS, Turner JM (2010) Evidence that meiotic sex chromosome inactivation is essential for male fertility. Curr Biol 20(23):2117–2123
Schimenti J (2000) Segregation distortion of mouse t haplotypes the molecular basis emerges. Trends Genet 16(6):240–243
Schoenmakers S, Wassenaar E, van Cappellen WA, Derijck AA, de Boer P, Laven JS, Grootegoed JA, Baarends WM (2008) Increased frequency of asynapsis and associated meiotic silencing of heterologous chromatin in the presence of irradiation-induced extra DNA double strand breaks. Dev Biol 317(1):270–281
Schoenmakers S, Wassenaar E, Hoogerbrugge JW, Laven JS, Grootegoed JA, Baarends WM (2009) Female meiotic sex chromosome inactivation in chicken. PLoS Genet 5(5):e1000466
Sciurano R, Rahn M, Rey-Valzacchi G, Solari AJ (2007) The asynaptic chromatin in spermatocytes of translocation carriers contains the histone variant gamma-H2AX and associates with the XY body. Hum Reprod 22(1):142–150
Shiu PK, Raju NB, Zickler D, Metzenberg RL (2001) Meiotic silencing by unpaired DNA. Cell 107(7):905–916
Smyth G (2005) Limma: linear models for microarray data. In: Gentleman R, Carey V, Dudoit S, Irizzary K, Huber W (eds) Bioinformatics and computational biology solutions using R and bioconductor. Springer, New York, pp 397–420
Solari AJ (1974) The behavior of the XY pair in mammals. Int Rev Cytol 38:273–317
Tres LL, Erickson RP (1982) Electron microscopy of t-allele synaptonemal complexes discloses no inversions. Nature 299(5885):752–754
Turner JM, Aprelikova O, Xu X, Wang R, Kim S, Chandramouli GV, Barrett JC, Burgoyne PS, Deng CX (2004) BRCA1, histone H2AX phosphorylation, and male meiotic sex chromosome inactivation. Curr Biol 14(23):2135–2142
Turner JM, Mahadevaiah SK, Fernandez-Capetillo O, Nussenzweig A, Xu X, Deng CX, Burgoyne PS (2005) Silencing of unsynapsed meiotic chromosomes in the mouse. Nat Genet 37(1):41–47
Turner JM, Mahadevaiah SK, Ellis PJ, Mitchell MJ, Burgoyne PS (2006) Pachytene asynapsis drives meiotic sex chromosome inactivation and leads to substantial postmeiotic repression in spermatids. Dev Cell 10(4):521–529
Vacik T, Ort M, Gregorova S, Strnad P, Blatny R, Conte N, Bradley A, Bures J, Forejt J (2005) Segmental trisomy of chromosome 17: a mouse model of human aneuploidy syndromes. Proc Natl Acad Sci U S A 102(12):4500–4505
van der Heijden GW, Derijck AA, Posfai E, Giele M, Pelczar P, Ramos L, Wansink DG, van der Vlag J, Peters AH, de Boer P (2007) Chromosome-wide nucleosome replacement and H3.3 incorporation during mammalian meiotic sex chromosome inactivation. Nat Genet 39(2):251–258
Vibranovski MD, Lopes HF, Karr TL, Long M (2009) Stage-specific expression profiling of Drosophila spermatogenesis suggests that meiotic sex chromosome inactivation drives genomic relocation of testis-expressed genes. PLoS Genet 5(11):e1000731
Acknowledgments
We thank the members of our laboratory for helpful discussions, M.A. Handel for anti-H1t antibody, M. Flemr for his help with oocyte preparation, and S. Takacova for reading the manuscript. This work was supported by the Academy of Sciences of the Czech Republic (Praemium Academiae to JF) and by the Ministry of Education, Youth and Sports of the Czech Republic (COST LD11079). DH is a Ph.D. student supported in part by the Faculty of Science, Charles University, Prague. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Fertility testing (DOC 2410 kb)
Table S2
Testes and body weights (DOC 763 kb)
Table S3
Frequencies of meiotic stages of primary spermatocytes and haploid cells in testicular cell spreads. (DOC 29 kb)
Table S4
RNA FISH of specific genes in late zygotene–early pachytene and in mid–late pachytene spermatocytes (DOC 32 kb)
Table S5
Probes for RNA FISH (DOC 36 kb)
Table S6
Summary of primers used for qRT-PCR (DOC 37 kb)
Figure S1
Substages of prophase I and round spermatids characterized by immunofluorescence microscopy of SYCP3 and H1t. The spread nuclei of testicular cells were immunostained by antibodies against SYCP3 and H1t, a testis-specific variant of histone linker H1. Based on the characteristic patterns of SYCP3, the substages of prophase I were determined for each spermatocyte and counted according to following criteria: leptotene, zygotene, and early pachynema are H1t negative; mid-pachytene stage shows mild H1t stain; and late pachytene spermatocytes show the most intense H1t stain. Diplotene spermatocytes and round spermatids show intense staining too. (DOC 30 kb)
Figure S2
Number of unrepaired double-strand breaks and their sub-nuclear location in mid–late pachytene spermatocytes differ between fertile and sterile males. a The spread nuclei were immunostained with antibodies against SYCP3 and Rad51, followed by DAPI staining. Number and location of individual Rad51 foci was analyzed in 25 mid–late pachytene spermatocyte nuclei for all three genotypes: T/t 12, T+/+T43H (T/T43H), and t 12+/+T43H (t 12/T43H). b Graphical representation shows average total counts of Rad51 foci per nucleus (n = 25) for each genotype and is subdivided into three groups: the foci associated with sex chromosomes (blue), with the translocated quadrivalent (green), and with autosomes (red). (DOC 42 kb)
Figure S3
qRT-PCR for the selected Chr17 genes in isolated pre–mid-pachytene spermatocytes. Downregulation of most of the selected Chr 17 genes in T+/+T43H males accompanies the asynapsis around the translocation breakpoint. Even stronger silencing is apparent in t 12+/+T43H males, reflecting the higher proportion of cells with asynapsed Chr 17. qRT-PCR was performed on three biological replicates of isolated populations. Values were normalized to expression of B6 fertile controls. Standard deviations are plotted. (DOC 266 kb)
Figure S4
Transcription silencing within γH2AX domains is less efficient in late pachytene spermatocytes of the T+/+T43H (T/T43H) and t 12+/+T43H (t 12/T43H) sterile males than in fertile control males. The overall transcription activity in late pachytene spermatocytes is analyzed by cot1 RNA FISH (upper panel, green; middle panel, gray scale) together with immunostaining directed against γH2AX (upper panel, red) and testes-specific histone linker H1t (lower panel, red), the nuclei are stained by DAPI (blue). The late pachytene spermatocytes were identified by typical intense H1t staining, and the unsynapsed chromatin is disclosed by γH2AX stain. The chromatin of sex bodies in B6 and t 12/+ fertile controls is separated from the rest of nuclei and devoid of nascent transcripts as detected by γH2AX staining and cot1 RNA FISH. In the contrast, the T+/+T43H and t 12+/+T43H spermatocytes show γH2AX-positive chromatin possessing more disperse character with traces of nascent transcripts. (DOC 160 kb)
Figure S5
Translocated chromosome of T+/+T43H females remains asynapsed during oogenesis in pachynema. The spread nuclei were immunostained with antibodies against SYCP3 (green) and centromere (red), followed by DAPI staining. The partial asynapsis of translocated chromosome (arrow) is demonstrated by faint SYCP3 stain. (DOC 132 kb)
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Homolka, D., Jansa, P. & Forejt, J. Genetically enhanced asynapsis of autosomal chromatin promotes transcriptional dysregulation and meiotic failure. Chromosoma 121, 91–104 (2012). https://doi.org/10.1007/s00412-011-0346-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-011-0346-5