Abstract
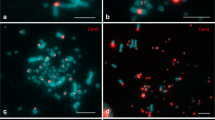
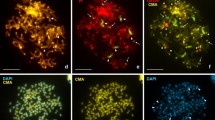
In situ hybridizations of single-copy GC-rich, gene-rich and GC-poor, gene-poor chicken DNA allowed us to localize the gene-rich and the gene-poor chromosomal regions in interphase nuclei of cold-blooded vertebrates. Our results showed that the gene-rich regions from amphibians (Rana esculenta) and reptiles (Podarcis sicula) occupy the more internal part of the nuclei, whereas the gene-poor regions occupy the periphery. This finding is similar to that previously reported in warm-blooded vertebrates, in spite of the lower GC levels of the gene-rich regions of cold-blooded vertebrates. This suggests that this similarity extends to chromatin structure, which is more open in the gene-rich regions of both mammals and birds and more compact in the gene-poor regions. In turn, this may explain why the compositional transition undergone by the genome at the emergence of homeothermy did not involve the entire ancestral genome but only a small part of it, and why it involved both coding and noncoding sequences. Indeed, the GC level increased only in that part of the genome that needed a thermodynamic stabilization, namely in the more open gene-rich chromatin of the nuclear interior, whereas the gene-poor chromatin of the periphery was stabilized by its own compact structure.






Similar content being viewed by others
References
Andreozzi L, Federico C, Motta S, Saccone S, Sazanova A, Sazanov A, Smirnov A, Galkina SA, Lukina NA, Rodionov AV, Carels N, Bernardi G (2001) Compositional mapping of chicken chromosomes and identification of the gene-richest regions. Chromosome Res 9:521–532
Bernardi G (1989) The isochore organization of the human genome. Annu Rev Genet 23:637–661
Bernardi G (2004) Structural and evolutionary genomics. Natural selection in genome evolution. Elsevier, Amsterdam
Bernardi G, Olofsson B, Filipski J, Zerial M, Salinas J, Cuny G, Meunier-Rotival M, Rodier F (1985) The mosaic genome of warm-blooded vertebrates. Science 228:953–957
Bolzer A, Kreth G, Solovei I, Koehler D, Saracoglu K, Fauth C, Müller S, Eils R, Cremer C, Speicher MR, Cremer T (2005) Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol 3(5):e157, pp 826–842
Cacciò S, Perani P, Saccone S, Kadi F, Bernardi G (1994) Single-copy sequence homology among the GC-richest isochores of the genomes from warm-blooded vertebrates. J Mol Evol 39:331–339
Capriglione T, Cardone A, Odierna G, Olmo E (1991) Evolution of a centromeric satellite DNA and phylogeny of a lacertid lizards. Comp Biochem Physiol 100:641–645
Corneo G, Ginelli E, Soave C, Bernardi G (1968) Isolation and characterization of mouse and guinea pig satellite deoxyribonucleic acids. Biochemistry 7:4373–4379
Cortadas J, Olofsson B, Meunier-Rotival M, Macaya G, Bernardi G (1979) The DNA components of the chicken genome. Eur J Biochem 99:179–186
Cuny G, Soriano P, Macaya G, Bernardi G (1981) The major components of the mouse and human genomes. 1. Preparation, basic properties and compositional heterogeneity. Eur J Biochem 115:227–233
Federico C, Saccone S, Bernardi G (1998) The gene-richest bands of human chromosomes replicate at the onset of the S-phase. Cytogenet Cell Genet 80:83–88
Federico C, Andreozzi L, Saccone S, Bernardi G (2000) Gene density in the Giemsa bands of human chromosomes. Chromosome Res 8:737–746
Federico C, Saccone S, Andreozzi L, Motta S, Russo V, Carels N, Bernardi G (2004) The pig genome: compositional analysis and identification of the gene-richest regions in chromosomes and nuclei. Gene 343:245–251
Ferreira J, Paolella G, Ramos C, Lamond AI (1997) Spatial organization of large-scale chromatin domains in the nucleus: a magnified view of single chromosome territories. J Cell Biol 139:1597–1610
Filipski J, Thiery JP, Bernardi G (1973) An analysis of the bovine genome by Cs2SO4-Ag+ density gradient centrifugation. J Mol Biol 80:177–197
Gilbert N, Boyle S, Fiegler H, Woodfine K, Carter NP, Bickmore WA (2004) Chromatin architecture of the human genome: gene-rich domains are enriched in open chromatin fibers. Cell 118:555–566
Habermann FA, Cremer M, Walter J, Kreth G, von Hase J, Bauer K, Wienberg J, Cremer C, Cremer T, Solovei I (2001) Arrangements of macro- and microchromosomes in chicken cells. Chromosome Res 9:569–584
Jabbari K, Bernardi G (2004) Cytosine methylation and CpG, TpG (CpA) and TpA frequencies. Gene 333:143–149
Kadi F, Mouchiroud D, Sabeur G, Bernardi G (1993) The compositional patterns of the avian genomes and their evolutionary implications. J Mol Evol 37:544–551
Macaya G, Thiery JP, Bernardi G (1976) An approach to the organization of eukaryotic genomes at a macromolecular level. J Mol Biol 108:237–254
McQueen HA, Fantes J, Cross SH, Clark VH, Archibald AL, Bird AP (1996) CpG islands of chicken are concentrated on microchromosomes. Nat Genet 12:321–324
Mouchiroud D, D'Onofrio G, Aïssani B, Macaya G, Gautier C, Bernardi G (1991) The distribution of genes in the human genome. Gene 100:181–187
Olmo E, Odierna G, Cobron O (1986) C-band variability and phylogeny of Lacertidae. Genetica 71:63–74
Olofsson B, Bernardi G (1983) Organization of nucleotide sequences in the chicken genome. Eur J Biochem 130:241–245
Saccone S, De Sario A, Della Valle G, Bernardi G (1992) The highest gene concentrations in the human genome are in T-bands of metaphase chromosomes. Proc Natl Acad Sci U S A 89:4913–4917
Saccone S, Pavlicek A, Federico C, Paces J, Bernardi G (2001) Genes, isochores and bands in human chromosomes 21 and 22. Chromosome Res 9:533–539
Saccone S, Federico C, Bernardi G (2002) Localization of the gene-richest and the gene-poorest isochores in the interphase nuclei of mammals and birds. Gene 300:169–178
Sadoni N, Langer S, Fauth C, Bernardi G, Cremer T, Turner BM, Zink D (1999) Nuclear organization of mammalian genomes: polar chromosome territories build up functionally distinct higher order compartments. J Cell Biol 146:1211–1226
Schempp W, Schmid M (1981) Chromosome banding in Amphibia. VI. BrdU-Replication patterns in anura and demonstration of XX/XY sex chromosomes in Rana esculenta. Chromosoma 83:697–710
Smith J, Bruley CK, Paton IR, Dunn I, Jones CT, Windsor D, Morrice DR, Law AS, Masabanda J, Sazanov A, Waddington D, Fries R, Burt DW (2000) Differences in gene density on chicken macrochromosomes and microchromosomes. Anim Genet 31:96–103
Strouboulis J, Wolffe AP (1996) Functional compartmentalization of the nucleus. J Cell Sci 109:1991–2000
Thiery JP, Macaya G, Bernardi G (1976) An analysis of eukaryotic genomes by density gradient centrifugation. J Mol Biol 108:219–235
Yonenaga-Yassuda Y, Kasahara S, Chu TH, Rodrigues MT (1988) High-resolution RGB-banding pattern in the genus Tropidurus (Sauria, Iguanidae). Cytogenet Cell Genet 48:68–71
Zoubak S, Clay O, Bernardi G (1996) The gene distribution of the human genome. Gene 174:95–102
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E.A. Nigg
Rights and permissions
About this article
Cite this article
Federico, C., Scavo, C., Cantarella, C.D. et al. Gene-rich and gene-poor chromosomal regions have different locations in the interphase nuclei of cold-blooded vertebrates. Chromosoma 115, 123–128 (2006). https://doi.org/10.1007/s00412-005-0039-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-005-0039-z