Abstract
Purpose
Cervical carcinoma is the second most prevalent and the fifth most deadly malignancy seen in women worldwide. Dysregulated activation of EGF ErbB system has been implicated in diverse types of human cancer; however, it is elusive how it is regulated in human cervical cancer cells. We herein aimed to explore the mechanisms of cervical carcinoma response to epidermal growth factor (EGF), with a view of the pathways activated by EGF.
Methods
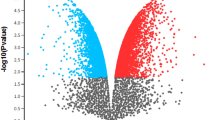
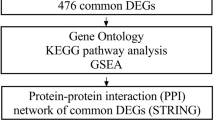
Using the GSE6783 affymetrix microarray data accessible from gene expression omnibus database, we first identified the differentially expressed genes between EGF-stimulated and -unstimulated samples. Then we constructed a regulation network and identified the network motifs. We also performed biological process and pathway enrichment analyses to functionally classify the genes in the regulation network.
Results
A total of 11 network motifs were identified in the regulation network. EGF treatment could increase the risk of cancer via dysregulation of cancer-related pathways and immune response pathways.
Conclusions
Network motif analysis is useful in mining the useful information underlying the network. We hope our work could serve as a basis for further experimentation.



Similar content being viewed by others
References
Armstrong EP (2010) Prophylaxis of cervical cancer and related cervical disease: a review of the cost-effectiveness of vaccination against oncogenic HPV types. J Manag Care Pharm 16(3):217–230
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ (2009) Cancer statistics. CA Cancer J Clin 59(4):225–249. doi:10.3322/caac.20006
Einstein MH, Schiller JT, Viscidi RP, Strickler HD, Coursaget P, Tan T, Halsey N, Jenkins D (2009) Clinician’s guide to human papillomavirus immunology: knowns and unknowns. Lancet Infect Dis 9(6):347–356. doi:10.1016/S1473-3099(09)70108-2
Brake T, Lambert PF (2005) Estrogen contributes to the onset, persistence, and malignant progression of cervical cancer in a human papillomavirus-transgenic mouse model. Proc Natl Acad Sci USA 102(7):2490–2495. doi:10.1073/pnas.0409883102
Herbst RS (2004) Review of epidermal growth factor receptor biology. Int J Radiat Oncol Biol Phys 59(2):21–26. doi:10.1016/j.ijrobp.2003.11.041
Levin ER (2003) Bidirectional signaling between the estrogen receptor and the epidermal growth factor receptor. Mol Endocrinol 17(3):309–317. doi:10.1210/me.2002-0368me.2002-0368
Vignon F, Bouton MM, Rochefort H (1987) Antiestrogens inhibit the mitogenic effect of growth factors on breast cancer cells in the total absence of estrogens. Biochem Biophys Res Commun 146(3):1502–1508, pii: 0006-291X(87)90819-9
Yarden Y, Sliwkowski MX (2001) Untangling the ErbB signalling network. Nat Rev Mol Cell Biol 2(2):127–137. doi:10.1038/35052073
Hynes NE, Lane HA (2005) ERBB receptors and cancer: the complexity of targeted inhibitors. Nat Rev Cancer 5(5):341–354. doi:10.1038/nrc1609
Mangan S, Alon U (2003) Structure and function of the feed-forward loop network motif. Proc Natl Acad Sci USA 100(21):11980–11985. doi:10.1073/pnas.21338411002133841100
Milo R, Shen-Orr S, Itzkovitz S, Kashtan N, Chklovskii D, Alon U (2002) Network motifs: simple building blocks of complex networks. Science 298(5594):824–827. doi:10.1126/science.298.5594.824298/5594/824
Han JD, Bertin N, Hao T, Goldberg DS, Berriz GF, Zhang LV, Dupuy D, Walhout AJ, Cusick ME, Roth FP, Vidal M (2004) Evidence for dynamically organized modularity in the yeast protein–protein interaction network. Nature 430(6995):88–93. doi:10.1038/nature02555nature02555
Jeong H, Mason SP, Barabasi AL, Oltvai ZN (2001) Lethality and centrality in protein networks. Nature 411(6833):41–42. doi:10.1038/3507513835075138
Jaimovich A, Elidan G, Margalit H, Friedman N (2006) Towards an integrated protein–protein interaction network: a relational Markov network approach. J Comput Biol 13(2):145–164. doi:10.1089/cmb.2006.13.145
Jeong H, Tombor B, Albert R, Oltvai ZN, Barabasi AL (2000) The large-scale organization of metabolic networks. Nature 407(6804):651–654. doi:10.1038/35036627
Amit I, Citri A, Shay T, Lu Y, Katz M, Zhang F, Tarcic G, Siwak D, Lahad J, Jacob-Hirsch J, Amariglio N, Vaisman N, Segal E, Rechavi G, Alon U, Mills GB, Domany E, Yarden Y (2007) A module of negative feedback regulators defines growth factor signaling. Nat Genet 39(4):503–512. doi:10.1038/ng1987
Matys V, Fricke E, Geffers R, Gossling E, Haubrock M, Hehl R, Hornischer K, Karas D, Kel AE, Kel-Margoulis OV, Kloos DU, Land S, Lewicki-Potapov B, Michael H, Munch R, Reuter I, Rotert S, Saxel H, Scheer M, Thiele S, Wingender E (2003) TRANSFAC: transcriptional regulation, from patterns to profiles. Nucl Acids Res 31(1):374–378
Jiang C, Xuan Z, Zhao F, Zhang MQ (2007) TRED: a transcriptional regulatory element database, new entries and other development. Nucl Acids Res 35:D137–D140. doi:10.1093/nar/gkl1041
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3 (3). doi:10.2202/1544-6115.1027
Derrick TR, Bates BT, Dufek JS (1994) Evaluation of time-series data sets using the Pearson product-moment correlation coefficient. Med Sci Sports Exerc 26(7):919–928
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. doi:10.1101/gr.123930313/11/2498
Wernicke S, Rasche F (2006) Fanmod: a tool for fast network motif detection. Bioinformatics 22(9):1152–1153. doi:10.1093/bioinformatics/btl038
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. doi:10.1038/nprot.2008.211
Brake T, Connor JP, Petereit DG, Lambert PF (2003) Comparative analysis of cervical cancer in women and in a human papillomavirus-transgenic mouse model: identification of minichromosome maintenance protein 7 as an informative biomarker for human cervical cancer. Cancer Res 63(23):8173–8180
Mangan S, Zaslaver A, Alon U (2003) The coherent feedforward loop serves as a sign-sensitive delay element in transcription networks. J Mol Biol 334(2):197–204, pii: S0022283603012038
Zaslaver A, Mayo AE, Rosenberg R, Bashkin P, Sberro H, Tsalyuk M, Surette MG, Alon U (2004) Just-in-time transcription program in metabolic pathways. Nat Genet 36(5):486–491. doi:10.1038/ng1348ng1348
Lo RS, Wotton D, Massague J (2001) Epidermal growth factor signaling via ras controls the smad transcriptional co-repressor TGIF. EMBO J 20(1–2):128–136. doi:10.1093/emboj/20.1.128
Derynck R, Zhang YE (2003) Smad-dependent and smad-independent pathways in TGF-beta family signalling. Nature 425(6958):577–584. doi:10.1038/nature02006
Lee HP, Seo SS (2002) The application of human papillomavirus testing to cervical cancer screening. Yonsei Med J 43(6):763–768
Li L, He S, Sun JM, Davie JR (2004) Gene regulation by Sp1 and Sp3. Biochem Cell Biol 82(4):460–471. doi:10.1139/o04-045o04-045
Wang LW, Li Q, Hua ZL, Zhou F, Keping X, Daoyan W, Yao J, Ajani J (2007) Expression of transcription factor Sp1 in human gastric cancer tissue and its correlation with prognosis. Zhonghua Zhong Liu Za Zhi 29(2):107–111
Wang F, Li Y, Zhou J, Xu J, Peng C, Ye F, Shen Y, Lu W, Wan X, Xie X (2011) miR-375 is down-regulated in squamous cervical cancer and inhibits cell migration and invasion via targeting transcription factor SP1. Am J Pathol 179(5):2580–2588. doi:10.1016/j.ajpath.2011.07.037
Cho S, Cinghu S, Yu JR, Park WY (2011) Helicase-like transcription factor confers radiation resistance in cervical cancer through enhancing the DNA damage repair capacity. J Cancer Res Clin Oncol 137(4):629–637. doi:10.1007/s00432-010-0925-5
Leslie MC, Bar-Eli M (2005) Regulation of gene expression in melanoma: new approaches for treatment. J Cell Biochem 94(1):25–38. doi:10.1002/jcb.20296
Papassava P, Gorgoulis VG, Papaevangeliou D, Vlahopoulos S, van Dam H, Zoumpourlis V (2004) Overexpression of activating transcription factor-2 is required for tumor growth and progression in mouse skin tumors. Cancer Res 64(23):8573–8584. doi:10.1158/0008-5472.CAN-03-0955
Vlahopoulos SA, Logotheti S, Mikas D, Giarika A, Gorgoulis V, Zoumpourlis V (2008) The role of ATF-2 in oncogenesis. Bioessays 30(4):314–327. doi:10.1002/bies.20734
Acknowledgments
This study was funded (in part) by Shanghai Science and Technology Committee Foundation, grant number [09411962500]. This study was funded (in part) by Shanghai Municipal Health Bureau Foundation, grant number [054019]. This study was funded (in part) by Foundation from The first People’s Hospital, Shanghai Jiaotong University, grant number [06B27].
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, S.F., Qian, W.Y., Zhang, J.W. et al. Network motifs in the transcriptional regulation network of cervical carcinoma cells respond to EGF. Arch Gynecol Obstet 287, 771–777 (2013). https://doi.org/10.1007/s00404-012-2608-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00404-012-2608-8