Abstract
The deposition of pathologic misfolded proteins in neurodegenerative disorders such as Alzheimer’s disease, Parkinson’s disease, frontotemporal dementia and amyotrophic lateral sclerosis is hypothesized to burden protein homeostatic (proteostatic) machinery, potentially leading to insufficient capacity to maintain the proteome. This hypothesis has been supported by previous work in our laboratory, as evidenced by the perturbation of cytosolic protein solubility in response to amyloid plaques in a mouse model of Alzheimer’s amyloidosis. In the current study, we demonstrate changes in proteome solubility are a common pathology to mouse models of neurodegenerative disease. Pathological accumulations of misfolded tau, α-synuclein and mutant superoxide dismutase 1 in CNS tissues of transgenic mice were associated with changes in the solubility of hundreds of CNS proteins in each model. We observed that changes in proteome solubility were progressive and, using the rTg4510 model of inducible tau pathology, demonstrated that these changes were dependent upon sustained expression of the primary pathologic protein. In all of the models examined, changes in proteome solubility were robust, easily detected, and provided a sensitive indicator of proteostatic disruption. Interestingly, a subset of the proteins that display a shift towards insolubility were common between these different models, suggesting that a specific subset of the proteome is vulnerable to proteostatic disruption. Overall, our data suggest that neurodegenerative proteinopathies modeled in mice impose a burden on the proteostatic network that diminishes the ability of neural cells to prevent aberrant conformational changes that alter the solubility of hundreds of abundant cellular proteins.








Similar content being viewed by others
References
Ayyadevara S, Balasubramaniam M, Parcon PA, Barger SW, Griffin WST, Alla R, Tackett AJ, Mackintosh SG, Petricoin E, Zhou W, Shmookler Reis RJ (2016) Proteins that mediate protein aggregation and cytotoxicity distinguish Alzheimer’s hippocampus from normal controls. Aging Cell 15:924–939. https://doi.org/10.1111/acel.12501
Babu JR, Geetha T, Wooten MW (2005) Sequestosome 1/p62 shuttles polyubiquitinated tau for proteasomal degradation. J Neurochem 94:192–203. https://doi.org/10.1111/j.1471-4159.2005.03181.x
Bai B, Hales CM, Chen P, Gozal Y, Dammer EB, Fritz JJ (2013) U1 small nuclear ribonucleoprotein complex and RNA splicing alterations in Alzheimer’ s disease. Proc Natl Acad Sci USA 110:16562–16567. https://doi.org/10.1073/pnas.1310249110. http://www.pnas.org/content/suppl/2013/09/10/1310249110.DCSupplemental
Bailey RM, Covy JP, Melrose HL, Rousseau L, Watkinson R, Knight J, Miles S, Farrer MJ, Dickson DW, Giasson BI, Lewis J (2013) LRRK2 phosphorylates novel tau epitopes and promotes tauopathy. Acta Neuropathol 126:809–827. https://doi.org/10.1007/s00401-013-1188-4
Balch WE, Morimoto RI, Dillin A, Kelly Jeffery W (2008) Adapting proteostasis for disease intervention. Science 319(80):916–919. https://doi.org/10.1126/science.1141448
Basso M, Massignan T, Samengo G, Cheroni C, De Biasi S, Salmona M, Bendotti C, Bonetto V (2006) Insoluble mutant SOD1 is partly oligoubiquitinated in amyotrophic lateral sclerosis mice. J Biol Chem 281:33325–33335. https://doi.org/10.1074/jbc.M603489200
Basso M, Samengo G, Nardo G, Massignan T, D’Alessandro G, Tartari S, Cantoni L, Marino M, Cheroni C, de Biasi S, Giordana MT, Strong MJ, Estevez AG, Salmona M, Bendotti C, Bonetto V (2009) Characterization of detergent-insoluble proteins in ALS indicates a causal link between nitrative stress and aggregation in pathogenesis. PLoS One 4:e8130. https://doi.org/10.1371/journal.pone.0008130
Bosco DA, LaVoie MJ, Petsko GA, Ringe D (2011) Proteostasis and movement disorders: Parkinson’s disease and amyotrophic lateral sclerosis. Cold Spring Harb Perspect Biol 3:a007500. https://doi.org/10.1101/cshperspect.a007500
Carrell RW, Lomas DA (1997) Conformational disease. Lancet 350:134–138. https://doi.org/10.1016/S0140-6736(97)02073-4
Cheroni C, Peviani M, Cascio P, DeBiasi S, Monti C, Bendotti C (2005) Accumulation of human SOD1 and ubiquitinated deposits in the spinal cord of SOD1G93A mice during motor neuron disease progression correlates with a decrease of proteasome. Neurobiol Dis 18:509–522. https://doi.org/10.1016/j.nbd.2004.12.007
Choi H, Larsen B, Lin Z-Y, Breitkreutz A, Mellacheruvu D, Fermin D, Qin ZS, Tyers M, Gingras A-C, Nesvizhskii AI (2011) SAINT: probabilistic scoring of affinity purification-mass spectrometry data. Nat Methods 8:70–73. https://doi.org/10.1038/nmeth.1541
Clippinger AK, D’Alton S, Lin WL, Gendron TF, Howard J, Borchelt DR, Cannon A, Carlomagno Y, Chakrabarty P, Cook C, Golde TE, Levites Y, Ranum L, Schultheis PJ, Xu G, Petrucelli L, Sahara N, Dickson DW, Giasson B, Lewis J (2013) Robust cytoplasmic accumulation of phosphorylated TDP-43 in transgenic models of tauopathy. Acta Neuropathol 126:39–50. https://doi.org/10.1007/s00401-013-1123-8
Colom-Cadena M, Gelpi E, Charif S, Belbin O, Blesa R, Marti MJ, Clarimon J, Lleo A (2013) Confluence of alpha-synuclein, tau, and beta-amyloid pathologies in dementia with Lewy bodies. J Neuropathol Exp Neurol 72:1203–1212. https://doi.org/10.1097/NEN.0000000000000018
Crippa V, Sau D, Rusmini P, Boncoraglio A, Onesto E, Bolzoni E, Galbiati M, Fontana E, Marino M, Carra S, Bendotti C, de Biasi S, Poletti A (2010) The small heat shock protein B8 (HspB8) promotes autophagic removal of misfolded proteins involved in amyotrophic lateral sclerosis (ALS). Hum Mol Genet 19:3440–3456. https://doi.org/10.1093/hmg/ddq257
Cuanalo-Contreras K, Mukherjee A, Soto C (2013) Role of protein misfolding and proteostasis deficiency in protein misfolding diseases and aging. Int J Cell Biol. https://doi.org/10.1155/2013/638083
Cuervo AM, Stefanis L, Fredenburg R, Lansbury PT, Sulzer D (2004) Impaired degradation of mutant-synuclein by chaperone-mediated autophagy. Science 305(80):1292–1295. https://doi.org/10.1126/science.1101738
David DC, Hauptmann S, Scherping I, Schuessel K, Keil U, Rizzu P, Ravid R, Dröse S, Brandt U, Müller WE, Eckert A, Götz J (2005) Proteomic and functional analyses reveal a mitochondrial dysfunction in P301L tau transgenic mice. J Biol Chem 280:23802–23814. https://doi.org/10.1074/jbc.M500356200
David DC, Layfield R, Serpell L, Narain Y, Goedert M, Spillantini MG (2002) Proteasomal degradation of tau protein. J Neurochem 83:176–185. https://doi.org/10.1046/j.1471-4159.2002.01137.x
Dickey CA, Koren J, Zhang Y-J, Xu Y-F, Jinwal UK, Birnbaum MJ, Monks B, Sun M, Cheng JQ, Patterson C, Bailey RM, Dunmore J, Soresh S, Leon C, Morgan D, Petrucelli L (2008) Akt and CHIP coregulate tau degradation through coordinated interactions. Proc Natl Acad Sci USA 105:3622–3627. https://doi.org/10.1073/pnas.0709180105
Dickey CA, Yue M, Lin W-L, Dickson DW, Dunmore JH, Lee WC, Zehr C, West G, Cao S, Clark AMK, Caldwell GA, Caldwell KA, Eckman C, Patterson C, Hutton M, Petrucelli L (2006) Deletion of the ubiquitin ligase CHIP leads to the accumulation, but not the aggregation, of both endogenous phospho- and caspase-3-cleaved tau species. J Neurosci 26:6985–6996. https://doi.org/10.1523/JNEUROSCI.0746-06.2006
Dickson DW, Bergeron C, Chin SS, Duyckaerts C, Horoupian D, Ikeda K, Jellinger K, Lantos PL, Lippa CF, Mirra SS, Tabaton M, Vonsattel JP, Wakabayashi K, Litvan I, Office of Rare Diseases of the National Institutes of Health (2002) Office of Rare Diseases neuropathologic criteria for corticobasal degeneration. J Neuropathol Exp Neurol 61:935–946
Dobson CM (1999) Protein misfolding, evolution and disease. Trends Biochem Sci 24:329–332. https://doi.org/10.1016/S0968-0004(99)01445-0
Dou F, Netzer WJ, Tanemura K, Li F, Hartl FU, Takashima A, Gouras GK, Greengard P, Xu H (2003) Chaperones increase association of tau protein with microtubules. Proc Natl Acad Sci USA 100:721–726. https://doi.org/10.1073/pnas.242720499
Douglas PM, Dillin A (2010) Protein homeostasis and aging in neurodegeneration. J Cell Biol 190:719–729. https://doi.org/10.1083/jcb.201005144
Drummond E, Nayak S, Faustin A, Pires G, Hickman RA, Askenazi M, Cohen M, Haldiman T, Kim C, Han X, Shao Y, Safar JG, Ueberheide B, Wisniewski T (2017) Proteomic differences in amyloid plaques in rapidly progressive and sporadic Alzheimer’s disease. Acta Neuropathol 133:933–954. https://doi.org/10.1007/s00401-017-1691-0
Dyllick-Brenzinger M, D’Souza CA, Dahlmann B, Kloetzel P-M, Tandon A (2010) Reciprocal effects of α-synuclein overexpression and proteasome inhibition in neuronal cells and tissue. Neurotox Res 17:215–227. https://doi.org/10.1007/s12640-009-9094-1
Elliott E, Tsvetkov P, Ginzburg I (2007) BAG-1 associates with Hsc70Tau complex and regulates the proteasomal degradation of Tau protein. J Biol Chem 282:37276–37284. https://doi.org/10.1074/jbc.M706379200
Emmanouilidou E, Stefanis L, Vekrellis K (2010) Cell-produced α-synuclein oligomers are targeted to, and impair, the 26S proteasome. Neurobiol Aging 31:953–968. https://doi.org/10.1016/j.neurobiolaging.2008.07.008
Flach K, Ramminger E, Hilbrich I, Arsalan-Werner A, Albrecht F, Herrmann L, Goedert M, Arendt T, Holzer M (2014) Axotrophin/MARCH7 acts as an E3 ubiquitin ligase and ubiquitinates tau protein in vitro impairing microtubule binding. Biochim Biophys Acta-Mol Basis Dis 1842:1527–1538. https://doi.org/10.1016/j.bbadis.2014.05.029
Frost B, Jacks RL, Diamond MI (2009) Propagation of Tau misfolding from the outside to the inside of a cell. J Biol Chem 284:12845–12852. https://doi.org/10.1074/jbc.M808759200
Giasson BI, Duda JE, Quinn SM, Zhang B, Trojanowski JQ, Lee VMY (2002) Neuronal α-synucleinopathy with severe movement disorder in mice expressing A53T human α-synuclein. Neuron 34:521–533. https://doi.org/10.1016/S0896-6273(02)00682-7
Giasson BI, Uryu K, Trojanowski JQ, Lee VMY (1999) Mutant and wild type human α-synucleins assemble into elongated filaments with distinct morphologies in vitro. J Biol Chem 274:7619–7622. https://doi.org/10.1074/jbc.274.12.7619
Gidalevitz T, Ben-Zvi A, Ho KH, Brignull HR, Morimoto RI (2006) Progressive disruption of cellular protein folding in models of polyglutamine diseases. Science 311(80):1471–1474. https://doi.org/10.1126/science.1124514
Gidalevitz T, Krupinski T, Garcia S, Morimoto RI (2009) Destabilizing protein polymorphisms in the genetic background direct phenotypic expression of mutant SOD1 toxicity. PLoS Genet. https://doi.org/10.1371/journal.pgen.1000399
Gozal YM, Gozal DM, Gearing M, Cheng D, Hanfelt JJ, Funderburk C, Peng J, Lah JJ, Levey AI (2009) Proteomics analysis reveals novel components in the detergent-insoluble subproteome in Alzheimer’ s disease research articles. J Proteome Res 8:5069–5079
Guo JL, Lee VMY (2011) Seeding of normal tau by pathological tau conformers drives pathogenesis of Alzheimer-like tangles. J Biol Chem 286:15317–15331. https://doi.org/10.1074/jbc.M110.209296
Gurney ME, Pu H, Chiu AY, Dal Canto MC, Polchow CY, Alexander DD, Caliendo J, Hentati A, Kwon YW, Deng H-X, Chen W, Zhai P, Sufit RL, Siddique T (1994) Motor neuron degeneration in mice that express a human Cu. Zn superoxide dismutase mutation. Science 264(80):1772–1775. https://doi.org/10.1126/science.8209258
Hales CM, Dammer EB, Deng Q, Duong DM, Gearing M, Troncoso JC, Thambisetty M, Lah JJ, Shulman JM, Levey AI, Seyfried NT (2016) Changes in the detergent-insoluble brain proteome linked to amyloid and tau in Alzheimer’s Disease progression. Proteomics. https://doi.org/10.1002/pmic.201600057
Hamano T, Gendron TF, Causevic E, Yen SH, Lin WL, Isidoro C, Deture M, Ko LW (2008) Autophagic-lysosomal perturbation enhances tau aggregation in transfectants with induced wild-type tau expression. Eur J Neurosci 27:1119–1130. https://doi.org/10.1111/j.1460-9568.2008.06084.x
Han DH, Na H-K, Choi WH, Lee JH, Kim YK, Won C, Lee S-H, Kim KP, Kuret J, Min D-H, Lee MJ (2014) Direct cellular delivery of human proteasomes to delay tau aggregation. Nat Commun 5:5633. https://doi.org/10.1038/ncomms6633
Hauw JJ, Daniel SE, Dickson D, Horoupian DS, Jellinger K, Lantos PL, McKee A, Tabaton M, Litvan I (1994) Preliminary NINDS neuropathologic criteria for Steele–Richardson–Olszewski syndrome (progressive supranuclear palsy). Neurology 44:2015–2019
Higashi S, Iseki E, Yamamoto R, Minegishi M, Hino H, Fujisawa K, Togo T, Katsuse O, Uchikado H, Furukawa Y, Kosaka K, Arai H (2007) Concurrence of TDP-43, tau and alpha-synuclein pathology in brains of Alzheimer’s disease and dementia with Lewy bodies. Brain Res 1184:284–294. https://doi.org/10.1016/j.brainres.2007.09.048
Higgs RE, Knierman MD, Freeman AB, Gelbert LM, Patil ST, Hale JE (2007) Estimating the statistical significance of peptide identifications from shotgun proteomics experiments. J Proteome Res 6:1758–1767. https://doi.org/10.1021/pr0605320
Hipp MS, Park SH, Hartl UU (2014) Proteostasis impairment in protein-misfolding and -aggregation diseases. Trends Cell Biol 24:506–514. https://doi.org/10.1016/j.tcb.2014.05.003
Hoover BR, Reed MN, Su J, Penrod RD, Kotilinek LA, Grant MK, Pitstick R, Carlson GA, Lanier LM, Yuan LL, Ashe KH, Liao D (2010) Tau mislocalization to dendritic spines mediates synaptic dysfunction independently of neurodegeneration. Neuron 68:1067–1081. https://doi.org/10.1016/j.neuron.2010.11.030
James BD, Wilson RS, Boyle PA, Trojanowski JQ, Bennett DA, Schneider JA (2016) TDP-43 stage, mixed pathologies, and clinical Alzheimer’s-type dementia. Brain 139:2983–2993. https://doi.org/10.1093/brain/aww224
Jinwal UK, O’Leary JC, Borysov SI, Jones JR, Li Q, Koren J, Abisambra JF, Vestal GD, Lawson LY, Johnson AG, Blair LJ, Jin Y, Miyata Y, Gestwicki JE, Dickey CA (2010) Hsc70 rapidly engages tau after microtubule destabilization. J Biol Chem 285:16798–16805. https://doi.org/10.1074/jbc.M110.113753
Johnston JA, Dalton MJ, Gurney ME, Kopito RR (2000) Formation of high molecular weight complexes of mutant Cu, Zn-superoxide dismutase in a mouse model for familial amyotrophic lateral sclerosis. Proc Natl Acad Sci USA 97:12571–12576. https://doi.org/10.1073/pnas.220417997
Kabashi E, Agar JN, Hong Y, Taylor DM, Minotti S, Figlewicz DA, Durham HD (2008) Proteasomes remain intact, but show early focal alteration in their composition in a mouse model of amyotrophic lateral sclerosis. J Neurochem 105:2353–2366. https://doi.org/10.1111/j.1471-4159.2008.05317.x
Kabashi E, Agar JN, Strong MJ, Durham HD (2012) Impaired proteasome function in sporadic amyotrophic lateral sclerosis. Amyotroph Lateral Scler 13:367–371. https://doi.org/10.3109/17482968.2012.686511
Kabashi E, Agar JN, Taylor DM, Minotti S, Durham HD (2004) Focal dysfunction of the proteasome: a pathogenic factor in a mouse model of amyotrophic lateral sclerosis. J Neurochem 89:1325–1335. https://doi.org/10.1111/j.1471-4159.2004.02453.x
Karagö GE, Duarte AMS, Akoury E, Ippel H, Biernat J, Moran Luengo T, Radli M, Didenko T, Nordhues BA, Veprintsev DB, Dickey CA, Mandelkow E, Zweckstetter M, Boelens R, Madl T, Rudiger SGD (2014) Hsp90-tau complex reveals molecular basis for specificity in chaperone action. Cell 156:963–974. https://doi.org/10.1016/j.cell.2014.01.037
Karch CM, Prudencio M, Winkler DD, Hart PJ, Borchelt DR (2009) Role of mutant SOD1 disulfide oxidation and aggregation in the pathogenesis of familial ALS. Proc Natl Acad Sci USA 106:7774–7779. https://doi.org/10.1073/pnas.0902505106
Kawarabayashi T, Younkin L (2001) Age-dependent changes in brain, CSF, and plasma amyloid β protein in the Tg2576 transgenic mouse model of Alzheimer’s disease. J Neurosci 21:372–381
Keck S, Nitsch R, Grune T, Ullrich O (2003) Proteasome inhibition by paired helical filament-tau in brains of patients with Alzheimer’s disease. J Neurochem 85:115–122. https://doi.org/10.1046/j.1471-4159.2003.01642.x
Labbadia J, Morimoto RI (2015) The biology of proteostasis in aging and disease. Annu Rev Biochem. https://doi.org/10.1146/annurev-biochem-060614-033955
Lee MJ, Lee JH, Rubinsztein DC (2013) Tau degradation: the ubiquitin–proteasome system versus the autophagy–lysosome system. Prog Neurobiol 105:49–59
Lewis J, Dickson DW, Lin W-L, Chisholm L, Corral A, Jones G, Yen S, Sahara N, Skipper L, Yager D, Eckman C, Hardy J, Hutton M (2001) Enhanced neurofibrillary degeneration in transgenic mice expressing mutant tau and APP. Science 293(80):1487–1491. https://doi.org/10.1126/science.1058189
Lewis J, McGowan E, Rockwood J, Melrose H, Nacharaju P, Van Slegtenhorst M, Gwinn-Hardy K, Paul Murphy M, Baker M, Yu X, Duff K, Hardy J, Corral A, Lin WL, Yen SH, Dickson DW, Davies P, Hutton M (2000) Neurofibrillary tangles, amyotrophy and progressive motor disturbance in mice expressing mutant (P301L) tau protein. Nat Genet 25:402–405. https://doi.org/10.1038/79109
Liang TW, Forman MS, Duda JE, McCluskey L, Trojanowski JQ, Siderowf A (2005) Multiple pathologies in a patient with a progressive neurodegenerative syndrome. J Neurol Neurosurg Psychiatry 76:252–255. https://doi.org/10.1136/jnnp.2004.039479
Liao L, Cheng D, Wang J, Duong DM, Losik TG, Gearing M, Rees HD, Lah JJ, Levey AI, Peng J (2004) Proteomic characterization of postmortem amyloid plaques isolated by laser capture microdissection. J Biol Chem 279:37061–37068. https://doi.org/10.1074/jbc.M403672200
Lindberg I, Shorter J, Wiseman RL, Chiti F, Dickey CA, McLean PJ (2015) Chaperones in Neurodegeneration. J Neurosci 35:13853–13859. https://doi.org/10.1523/JNEUROSCI.2600-15.2015
Love S, Saitoh T, Quijada S, Cole GM, Terry RD (1988) Alz-50, ubiquitin and tau immunoreactivity of neurofibrillary tangles, Pick bodies and Lewy bodies. J Neuropathol Exp Neurol 47:393–405
McKinley MP, Meyer RK, Kenaga L, Rahbar F, Cotter R, Serban A, Prusiner SB (1991) Scrapie prion rod formation in vitro requires both detergent extraction and limited proteolysis. J Virol 65:1340–1351
Mi H, Huang X, Muruganujan A, Tang H, Mills C, Kang D, Thomas PD (2017) PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res 45:D183–D189. https://doi.org/10.1093/nar/gkw1138
Mi H, Muruganujan A, Casagrande JT, Thomas PD (2013) Large-scale gene function analysis with the PANTHER classification system. Nat Protoc 8:1551–1566. https://doi.org/10.1038/nprot.2013.092
Miyata Y, Koren J, Kiray J, Dickey CA, Gestwicki JE (2011) Molecular chaperones and regulation of tau quality control: strategies for drug discovery in tauopathies. Future Med Chem 3:1523–1537. https://doi.org/10.4155/fmc.11.88
Morimoto RI, Cuervo AM (2014) Proteostasis and the aging proteome in health and disease. J Gerontol Ser A Biol Sci Med Sci 69:S33–S38. https://doi.org/10.1093/gerona/glu049
Myeku N, Clelland CL, Emrani S, Kukushkin NV, Yu WH, Goldberg AL, Duff KE (2015) Tau-driven 26S proteasome impairment and cognitive dysfunction can be prevented early in disease by activating cAMP-PKA signaling. Nat Med 6:1–11. https://doi.org/10.1038/nm.4011
Old WM (2005) Comparison of label-free methods for quantifying human proteins by shotgun proteomics. Mol Cell Proteomics 4:1487–1502. https://doi.org/10.1074/mcp.M500084-MCP200
Olzscha H, Schermann SM, Woerner AC, Pinkert S, Hecht MH, Tartaglia GG, Vendruscolo M, Hayer-Hartl M, Hartl FU, Vabulas RM (2011) Amyloid-like aggregates sequester numerous metastable proteins with essential cellular functions. Cell 144:67–78. https://doi.org/10.1016/j.cell.2010.11.050
Petrucelli L, Dickson D, Kehoe K, Taylor J, Snyder H, Grover A, De Lucia M, McGowan E, Lewis J, Prihar G, Kim J, Dillmann WH, Browne SE, Hall A, Voellmy R, Tsuboi Y, Dawson TM, Wolozin B, Hardy J, Hutton M (2004) CHIP and Hsp70 regulate tau ubiquitination, degradation and aggregation. Hum Mol Genet 13:703–714. https://doi.org/10.1093/hmg/ddh083
Piras A, Collin L, Grüninger F, Graff C, Rönnbäck A (2016) Autophagic and lysosomal defects in human tauopathies: analysis of post-mortem brain from patients with familial Alzheimer disease, corticobasal degeneration and progressive supranuclear palsy. Acta Neuropathol Commun 4:22. https://doi.org/10.1186/s40478-016-0292-9
Prokai L, Stevens SM, Rauniyar N, Nguyen V (2009) Rapid label-free identification of estrogen-induced differential protein expression in vivo from mouse brain and uterine tissue. J Proteome Res 8:3862–3871. https://doi.org/10.1021/pr900083v
Ramsden M, Kotilinek L, Forster C, Paulson J, McGowan E, SantaCruz K, Guimaraes A, Yue M, Lewis J, Carlson G, Hutton M, Ashe KH (2005) Age-dependent neurofibrillary tangle formation, neuron loss, and memory impairment in a mouse model of human tauopathy (P301L). J Neurosci 25:10637–10647. https://doi.org/10.1523/JNEUROSCI.3279-05.2005
Sacino AN, Brooks M, Thomas MA, McKinney AB, Lee S, Regenhardt RW, McGarvey NH, Ayers JI, Notterpek L, Borchelt DR, Golde TE, Giasson BI (2014) Intramuscular injection of α-synuclein induces CNS α-synuclein pathology and a rapid-onset motor phenotype in transgenic mice. Proc Natl Acad Sci USA 111:1–6. https://doi.org/10.1073/pnas.1321785111
SantaCruz K, Lewis J, Spires-Jones TL, Paulson J, Kotilinek L, Ingelsson M, Guimaraes A, DeTure M, Ramsden M, McGowan E, Forster C, Yue M, Orne J, Janus C, Mariash A, Kuskowski M, Hyman B, Hutton M, Ashe KH (2005) Tau suppression in a neurodegenerative mouse model improves memory function. Science 309(80):476–481. https://doi.org/10.1126/science.1113694
Sarkar M, Kuret J, Lee G (2008) Two motifs within the tau microtubule-binding domain mediate its association with the hsc70 molecular chaperone. J Neurosci Res 86:2763–2773. https://doi.org/10.1002/jnr.21721
Sau D, De Biasi S, Vitellaro-Zuccarello L, Riso P, Guarnieri S, Porrini M, Simeoni S, Crippa V, Onesto E, Palazzolo I, Rusmini P, Bolzoni E, Bendotti C, Poletti A (2007) Mutation of SOD1 in ALS: a gain of a loss of function. Hum Mol Genet 16:1604–1618. https://doi.org/10.1093/hmg/ddm110
Scherzinger E, Sittler A, Schweiger K, Heiser V, Lurz R, Hasenbank R, Bates GP, Lehrach H, Wanker EE (1999) Self-assembly of polyglutamine-containing huntingtin fragments into amyloid-like fibrils: implications for Huntington’s disease pathology. Proc Natl Acad Sci USA 96:4604–4609. https://doi.org/10.1073/pnas.96.8.4604
Seyfried NT, Gozal YM, Donovan LE, Herskowitz JH, Dammer EB, Xia Q, Ku L, Chang J, Duong DM, Rees HD, Cooper DS, Glass JD, Gearing M, Tansey MG, Lah JJ, Feng Y, Levey AI, Peng J (2012) Quantitative analysis of the detergent-insoluble brain proteome in frontotemporal lobar degeneration using SILAC internal standards. J Proteome Res 11:2721–2738. https://doi.org/10.1021/pr2010814
Shaw BF, Lelie HL, Durazo A, Nersissian AM, Xu G, Chan PK, Gralla EB, Tiwari A, Hayward LJ, Borchelt DR, Valentine JS, Whitelegge JP (2008) Detergent-insoluble aggregates associated with amyotrophic lateral sclerosis in transgenic mice contain primarily full-length, unmodified superoxide dismutase-1. J Biol Chem 283:8340–8350. https://doi.org/10.1074/jbc.M707751200
Shimura H, Schwartz D, Gygi SP, Kosik KS (2004) CHIP-Hsc70 complex ubiquitinates phosphorylated tau and enhances cell survival. J Biol Chem 279:4869–4876. https://doi.org/10.1074/jbc.M305838200
Sokal RR, Rohlf JF (1995) Biometry, the principles and practice of statistics in biological research. Freeman, W.H
Soto C (2003) Unfolding the role of protein misfolding in neurodegenerative diseases. Nat Rev Neurosci 4:49–60. https://doi.org/10.1038/nrn1007
Spillantini MG, Goedert M, Crowther RA, Murrell JR, Farlow MR, Ghetti B (1997) Familial multiple system tauopathy with presenile dementia: a disease with abundant neuronal and glial tau filaments. Proc Natl Acad Sci USA 94:4113–4118. https://doi.org/10.1073/pnas.94.8.4113
Spires-Jones TL, Attems J, Thal DR (2017) Interactions of pathological proteins in neurodegenerative diseases. Acta Neuropathol. https://doi.org/10.1007/s00401-017-1709-7
Strang KH, Croft CL, Sorrentino ZA, Chakrabarty P, Golde TE, Giasson BI (2018) Distinct differences in prion-like seeding and aggregation between tau protein variants provide mechanistic insights into tauopathies. J Biol Chem 293:2408–2421. https://doi.org/10.1074/jbc.M117.815357
Tai HC, Serrano-Pozo A, Hashimoto T, Frosch MP, Spires-Jones TL, Hyman BT (2012) The synaptic accumulation of hyperphosphorylated tau oligomers in Alzheimer disease is associated with dysfunction of the ubiquitin–proteasome system. Am J Pathol 181:1426–1435. https://doi.org/10.1016/j.ajpath.2012.06.033
Taylor RC, Dillin A (2011) Aging as an event of proteostasis collapse. Cold Spring Harb Perspect Biol 3:1–17. https://doi.org/10.1101/cshperspect.a004440
Trojanowski JQ, Wang J, Lee VM (1999) Purification of paired helical filament tau and normal tau from human brain tissue. Methods Enzymol 309:81–89
Walker L, Kirsty K, Mcaleese E, Thomas AJ, Johnson M, Martin-ruiz C, Parker C, Colloby SJ, Jellinger K, Attems J (2015) Neuropathologically mixed Alzheimer’s and Lewy body disease: burden of pathological protein aggregates differs between clinical phenotypes. Acta Neuropathol 129:729–748
Wang J, Slunt H, Gonzales V, Fromholt D, Coonfield M, Copeland NG, Jenkins NA, Borchelt DR (2003) Copper-binding-site-null SOD1 causes ALS in transgenic mice: aggregates of non-native SOD1 delineate a common feature. Hum Mol Genet 12:2753–2764. https://doi.org/10.1093/hmg/ddg312
Wang J, Xu G, Gonzales V, Coonfield M, Fromholt D, Copeland NG, Jenkins NA, Borchelt DR (2002) Fibrillar inclusions and motor neuron degeneration in transgenic mice expressing superoxide dismutase 1 with a disrupted copper-binding site. Neurobiol Dis 10:128–138. https://doi.org/10.1006/nbdi.2002.0498
Wang P, Joberty G, Buist A, Vanoosthuyse A, Stancu I-C, Vasconcelos B, Pierrot N, Faelth-Savitski M, Kienlen-Campard P, Octave J-N, Bantscheff M, Drewes G, Moechars D, Dewachter I (2017) Tau interactome mapping based identification of Otub1 as tau deubiquitinase involved in accumulation of pathological tau forms in vitro and in vivo. Acta Neuropathol 133:731–749. https://doi.org/10.1007/s00401-016-1663-9
Wang Q, Woltjer RL, Cimino PJ, Pan C, Montine KS, Zhang J, Montine TJ (2005) Proteomic analysis of neurofibrillary tangles in Alzheimer disease identifies GAPDH as a detergent-insoluble paired helical filament tau binding protein. FASEB J 19:869–871. https://doi.org/10.1096/fj.04-3210fje
Wang Y, Martinez-Vicente M, Krüger U, Kaushik S, Wong E, Mandelkow E-M, Cuervo AM, Mandelkow E (2010) Synergy and antagonism of macroautophagy and chaperone-mediated autophagy in a cell model of pathological tau aggregation. Autophagy 6:182–183
Woltjer RL, Cimino PJ, Boutte AM, Schantz AM, Montine KS, Larson EB, Bird T, Quinn JF, Zhang J, Montine TJ (2005) Proteomic determination of widespread detergent insolubility, including A but not tau, early in the pathogenesis of Alzheimer’s disease. FASEB J 22:1–23. https://doi.org/10.1096/fj.05-4263fje
Xia Q, Liao L, Cheng D, Duong DM, Gearing M, Lah JJ, Levey AI, Peng J (2008) Proteomic identification of novel proteins associated with Lewy bodies. Front Biosci 13:3850–3856. https://doi.org/10.2741/2973
Xu G, Pattamatta A, Hildago R, Pace MC, Brown H, Borchelt DR (2016) Vulnerability of newly synthesized proteins to proteostasis stress. J Cell Sci 129:1892–1901. https://doi.org/10.1242/jcs.176479
Xu G, Stevens SM, Kobiessy F, Brown H, McClung S, Gold MS, Borchelt DR (2012) Identification of proteins sensitive to thermal stress in human neuroblastoma and glioma cell lines. PLoS ONE 7:1–13. https://doi.org/10.1371/journal.pone.0049021
Xu G, Stevens SM, Moore BD, McClung S, Borchelt DR (2013) Cytosolic proteins lose solubility as amyloid deposits in a transgenic mouse model of Alzheimer-type amyloidosis. Hum Mol Genet 22:2765–2774. https://doi.org/10.1093/hmg/ddt121
Yokota O, Davidson Y, Bigio EH, Ishizu H, Terada S, Arai T, Hasegawa M, Akiyama H, Sikkink S, Pickering-Brown S, Mann DMA (2010) Phosphorylated TDP-43 pathology and hippocampal sclerosis in progressive supranuclear palsy. Acta Neuropathol 120:55–66. https://doi.org/10.1007/s00401-010-0702-1
Zhang NY, Tang Z, Liu CW (2008) α-synuclein protofibrils inhibit 26 S proteasome-mediated protein degradation: understanding the cytotoxicity of protein protofibrils in neurodegenerative disease pathogenesis. J Biol Chem 283:20288–20298. https://doi.org/10.1074/jbc.M710560200
Acknowledgements
We thank the Interdisciplinary Center for Biotechnology Research (ICBR), specifically the Proteomics and Mass Spectrometry Core for assistance in processing LC–MS/MS samples. We also thank personnel within the University of Florida Animal Care Services for assistance with animal care for the mice used in this study. We also acknowledge the generous contribution of wild-type tau K18 fibril preparations from Kevin Strang within the laboratory of Dr. Benoit Giasson.
Funding
This work was supported by a grants from the National Institute of Neurological Disorders and Stroke (R21NS083006 to D.R.B. and J.L.; R01NS089622 to B.I.G.; T32NS082128 supported M.P.), the National Institute on Aging (P50AG047266; R01AG049456 to D.R.B. and J.L.), and the BrightFocus Foundation (A20141085) to G.X.; and by funding from the Santa Fe HealthCare Alzheimer’s Disease Research Center.
Author information
Authors and Affiliations
Contributions
Conceptualization, DRB, JL, and GX; methodology, DRB, JL, GX, and BIG; formal analysis, MCP and GX; investigation, MCP, GX, KWC, SF, and JH; resources, DRB, JL, and BIG; writing—original draft, MCP; writing—reviewing and editing, MCP, DRB, JL, GX, and BIG.; supervision, DRB, JL, and GX; funding acquisition, DRB, JL, and GX.
Corresponding authors
Ethics declarations
Ethical approval
All applicable international, national and/or institutional guidelines for the care and use of animals were followed. All procedures performed in studies involving animals were in accordance with the ethical standards of the University of Florida Institutional Animal Care and Use Committee (IACUC). This article does not contain any studies with human participants performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
401_2018_1895_MOESM1_ESM.docx
Table S1. Statistical information for different proteinopathy animal groups analyzed via LC–MS/MS. Supplemental Materials and Methods. Supplementary material 1 (DOCX 46 kb)
401_2018_1895_MOESM2_ESM.xlsx
Table S13 Overlapping protein identifications between SDS-insoluble fractions from rTg4510 mice and previous proteomic studies of disease-associated pathological features. Table S2. Spectral counts exported from Scaffold (version Scaffold_4.7.3, Proteome Software Inc., Portland, OR) used for analysis of rTg4510 mice (with and without doxycycline treatment) and rTg21221 mice. Table S3. Comprehensive lists of proteins that were aberrantly detected in SDS-insoluble fractions in 7.0 and 9.5 month-old based upon fold-change and G-test criteria. Analysis was conducted on rTg4510 7.0-month-old (n = 10) and 9.5-month-old (n = 3) mice. For 7.0-month-old mice, any given protein must have reached our criterion in 7 out of 10 analyzed mice. Table S4. Comprehensive list of proteins that met SAINT score thresholds of 0.9 as over-represented in SDS-insoluble fractions in all of the proteinopathy models. Analysis was conducted on JNPL3 (n = 7), rTg4510 7-month (n = 10), rTg4510 9.5 month (n = 3), M83 (n = 6), M83 seeded (n = 4), APPswe/PS1dE9 20-month (n = 3) and G93A SOD1 (n = 4) mice. Models of spinal proteinopathy were harvested at end-stage phenotype (paresis in one or more hind limbs). Proteins were accepted if they reached a SAINT score of ≥ 0.9 between control (NTg) mice and each corresponding transgenic model of neurodegenerative proteinopathy. Table S5. Comparison of proteins in SDS-insoluble fractions across cortical models (rTg4510 & L85) and spinal models (JNPL3, M83, M83 seeded, and G93A SOD1). Table S6. Spectral count data from proteins analyzed by immunoblotting Fig. 2a. Table S7. List of proteins that are most highly over-represented in SDS-insoluble fractions from spinal cords of end-stage G93A SOD1 mice. Individual animal spectral counts are separated by a comma in the SDS-insoluble columns. All spectral count comparisons exhibited a G-test value of < 0.01 (n = 4). nTg = nontransgenic, Tg = transgenic, PBS-S = PBS soluble. Table S8. List of the 91 common proteins that lose solubility in the G93A SOD1, APPswe/PS1dE9, and rTg4510 models of neurodegenerative proteinopathy. The protein list was compiled based upon the lists of proteins generated in Online Resource 2 Table S4. Table S9 List of proteins common to SDS-insoluble fractions across all spinal proteinopathy models (JNPL3, M83, M83 seeded, and G93A SOD1). The protein list was compiled based upon the lists of proteins generated in Online Resource, Table S4. Table S10 Compiled lists of proteins that lose solubility in rTg4510 mice that either overlap (Column A) or do not overlap (Column B) with previous studies identifying proteins that may be interacting with tau (see Online Resource 1, Table S13). Table S11 Compiled lists of proteins that lose solubility in rTg4510 mice that either overlap (Column A) or do not overlap (Column B) with previous studies identifying proteins that may interact with pathologic features of AD (see Online Resource 1, Table S13). Table S12 Complete lists of overlapping proteins between rTg4510 SDS-insoluble forebrains and the two first studies listed in Online Resource 1, Table S13. Supplementary material 2 (XLSX 1107 kb)
401_2018_1895_MOESM3_ESM.tif
Two-way clustering of spectral count data from rTg4510 mice. Clustering is based upon detergent-insoluble peptide spectra for 206 total proteins identified as affected in rTg4510 mice of any age (with and without DOX treatment). Red is indicative of the highest number of peptide spectra for a given protein relative to nontransgenic control mice, while blue is indicative of an absence of the peptide in SDS-insoluble fractions, or absence of a difference between transgenic and nontransgenic samples. Figure generated using JMP Pro Statistical Discovery from SAS (version 13.0, Cary, NC, USA). Supplementary material 3 (TIFF 1165 kb)
401_2018_1895_MOESM4_ESM.tif
Venn diagram of common proteins identified in APPswe/PS1dE9 (L85) across different analysis timepoints. Newly analyzed L85 mice were compared to previously analyzed animals. Supplementary material 4 (TIFF 3643 kb)
401_2018_1895_MOESM5_ESM.xlsx
Spectral counts exported from Scaffold (version Scaffold_4.7.3, Proteome Software Inc., Portland, OR) used for analysis of the APPswe/PS1E9 model of amyloidosis, the JNPL3, M83, and G93A models of spinal proteinopathy, and the seeded N2a cells. Supplementary material 5 (XLSX 428 kb)
401_2018_1895_MOESM6_ESM.tif
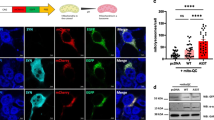
Immunoreactivity for Hspa4 protein is not highly co-localized with neurofibrillary tangle pathology of rTg4510 mice. Nontransgenic mice (a & b) exhibit minimal positive staining of Hspa4 puncta (green). rTg4510 (c & d) exhibit dramatic increases in Hspa4 puncta (green) that do not directly co-localize with neurofibrillary tangles (stained using the MC1 antibody to misfolded human tau, red). rTg4510 mice that received doxycycline to suppress mutant tau expression from 4.5 – 7.0 months of age (e & f) exhibited reduced Hspa4 immunostaining compared to rTg4510 mice that did not receive doxycycline. The graph shows the number of spectral counts for Hspa4 in SDS-insoluble fractions from the forebrains of NTg, 7-month-old rTg4510 mice, and 7-month-old rTg4510 that began DOX treatment at 4.5 months of age (g). Supplementary material 6 (TIFF 24679 kb)
401_2018_1895_MOESM7_ESM.tif
Two-way clustering of SDS-insoluble spectra for rTg4510, M83, M83 seeded, JNPL3, and G93A SOD1 models. Clustering is based upon detergent-insoluble peptide spectra for 310 total proteins identified as affected in any spinal proteinopathy model. Red is indicative of the highest number of peptide spectra for a given protein relative to nontransgenic control mice, while blue is indicative of an absence of the peptide in SDS-insoluble fractions, or absence of a difference between transgenic and nontransgenic samples. Figure generated using JMP Pro Statistical Discovery from SAS (version 13.0, Cary, NC, USA). Supplementary material 7 (TIFF 6224 kb)
401_2018_1895_MOESM8_ESM.tif
Bioinformatic analysis of protein classes that are statistically over-represented in SDS-insoluble fractions rTg4510 mice. Pie chart of protein classes uniquely affected in rTg4510 mice. The protein list was compiled based upon the lists of proteins generated in Online Resource 2, Table S4. Supplementary material 8 (TIFF 4701 kb)
401_2018_1895_MOESM9_ESM.tif
Combined Venn diagram representative of both diagrams from Fig. 8, encompassing the numbers of overlapping proteins from all types of methodologies used in previous literature (IP, Detergent-Insoluble, LCM, and Other). Supplementary material 9 (TIFF 25481 kb)
Rights and permissions
About this article
Cite this article
Pace, M.C., Xu, G., Fromholt, S. et al. Changes in proteome solubility indicate widespread proteostatic disruption in mouse models of neurodegenerative disease. Acta Neuropathol 136, 919–938 (2018). https://doi.org/10.1007/s00401-018-1895-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-018-1895-y