Abstract
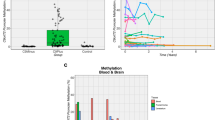
C9orf72 promoter hypermethylation inhibits the accumulation of pathologies which have been postulated to be neurotoxic. We tested here whether C9orf72 hypermethylation is associated with prolonged disease in C9orf72 mutation carriers. C9orf72 methylation was quantified from brain or blood using methylation-sensitive restriction enzyme digest-qPCR in a cross-sectional cohort of 118 C9orf72 repeat expansion carriers and 19 non-carrier family members. Multivariate regression models were used to determine whether C9orf72 hypermethylation was associated with age at onset, disease duration, age at death, or hexanucleotide repeat expansion size. Permutation analysis was performed to determine whether C9orf72 methylation is heritable. We observed a high correlation between C9orf72 methylation across tissues including cerebellum, frontal cortex, spinal cord and peripheral blood. While C9orf72 methylation was not significantly different between ALS and FTD and did not predict age at onset, brain and blood C9orf72 hypermethylation was associated with later age at death in FTD (brain: β = 0.18, p = 0.006; blood: β = 0.15, p < 0.001), and blood C9orf72 hypermethylation was associated with longer disease duration in FTD (β = 0.03, p = 0.007). Furthermore, C9orf72 hypermethylation was associated with smaller hexanucleotide repeat length (β = −16.69, p = 0.033). Finally, analysis of pedigrees with multiple mutation carriers demonstrated a significant association between C9orf72 methylation and family relatedness (p < 0.0001). C9orf72 hypermethylation is associated with prolonged disease in C9orf72 repeat expansion carriers with FTD. The attenuated clinical phenotype associated with C9orf72 hypermethylation suggests that slower clinical progression in FTD is associated with reduced expression of mutant C9orf72. These results support the hypothesis that expression of the hexanucleotide repeat expansion is associated with a toxic gain of function.




Similar content being viewed by others
References
Ash PE, Bieniek KF, Gendron TF et al (2013) Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron 77:639–646. doi:10.1016/j.neuron.2013.02.004
Beck J, Poulter M, Hensman D et al (2013) Large C9orf72 hexanucleotide repeat expansions are seen in multiple neurodegenerative syndromes and are more frequent than expected in the UK population. Am J Hum Genet 92:345–353. doi:10.1016/j.ajhg.2013.01.011
Bell JT, Pai AA, Pickrell JK et al (2011) DNA methylation patterns associate with genetic and gene expression variation in HapMap cell lines. Genome Biol 12:R10. doi:10.1186/gb-2011-12-1-r10
Belzil VV, Bauer PO, Prudencio M et al (2013) Reduced C9orf72 gene expression in c9FTD/ALS is caused by histone trimethylation, an epigenetic event detectable in blood. Acta Neuropathol 126:895–905. doi:10.1007/s00401-013-1199-1
Belzil VV, Bauer PO, Gendron TF, Murray ME, Dickson D, Petrucelli L (2014) Characterization of DNA hypermethylation in the cerebellum of c9FTD/ALS patients. Brain Res. doi:10.1016/j.brainres.2014.02.015
Boeve BF, Boylan KB, Graff-Radford NR et al (2012) Characterization of frontotemporal dementia and/or amyotrophic lateral sclerosis associated with the GGGGCC repeat expansion in C9ORF72. Brain 135:765–783. doi:10.1093/brain/aws004
Brettschneider J, Van Deerlin VM, Robinson JL et al (2012) Pattern of ubiquilin pathology in ALS and FTLD indicates presence of C9ORF72 hexanucleotide expansion. Acta Neuropathol 123:825–839. doi:10.1007/s00401-012-0970-z
Byrne S, Elamin M, Bede P et al (2012) Cognitive and clinical characteristics of patients with amyotrophic lateral sclerosis carrying a C9orf72 repeat expansion: a population-based cohort study. Lancet Neurol 11:232–240. doi:10.1016/S1474-4422(12)70014-5
Castaldo I, Pinelli M, Monticelli A et al (2008) DNA methylation in intron 1 of the frataxin gene is related to GAA repeat length and age of onset in Friedreich ataxia patients. J Med Genet 45:808–812. doi:10.1136/jmg.2008.058594
Chio A, Borghero G, Restagno G et al (2012) Clinical characteristics of patients with familial amyotrophic lateral sclerosis carrying the pathogenic GGGGCC hexanucleotide repeat expansion of C9ORF72. Brain 135:784–793. doi:10.1093/brain/awr366
Ciura S, Lattante S, Le Ber I et al (2013) Loss of function of C9orf72 causes motor deficits in a zebrafish model of Amyotrophic Lateral Sclerosis. Ann Neurol. doi:10.1002/ana.23946
Cleary JD, Tome S, Lopez Castel A et al (2010) Tissue- and age-specific DNA replication patterns at the CTG/CAG-expanded human myotonic dystrophy type 1 locus. Nat Struct Mol Biol 17:1079–1087. doi:10.1038/nsmb.1876
Colak D, Zaninovic N, Cohen MS et al (2014) Promoter-bound trinucleotide repeat mRNA drives epigenetic silencing in fragile X syndrome. Science 343:1002–1005. doi:10.1126/science.1245831
Cooper-Knock J, Hewitt C, Highley JR et al (2012) Clinico-pathological features in amyotrophic lateral sclerosis with expansions in C9ORF72. Brain 135:751–764. doi:10.1093/brain/awr365
DeJesus-Hernandez M, Mackenzie IR, Boeve BF et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72:245–256. doi:10.1016/j.neuron.2011.09.011
Devys D, Biancalana V, Rousseau F, Boue J, Mandel JL, Oberle I (1992) Analysis of full fragile X mutations in fetal tissues and monozygotic twins indicate that abnormal methylation and somatic heterogeneity are established early in development. Am J Med Genet 43:208–216
Dion V, Lin Y, Hubert L Jr, Waterland RA, Wilson JH (2008) Dnmt1 deficiency promotes CAG repeat expansion in the mouse germline. Hum Mol Genet 17:1306–1317. doi:10.1093/hmg/ddn019
Dion V, Wilson JH (2009) Instability and chromatin structure of expanded trinucleotide repeats. Trends Genet 25:288–297. doi:10.1016/j.tig.2009.04.007
Dobson-Stone C, Hallupp M, Loy CT et al (2013) C9ORF72 repeat expansion in Australian and Spanish frontotemporal dementia patients. PLoS One 8:e56899. doi:10.1371/journal.pone.0056899
Dols-Icardo O, Garcia-Redondo A, Rojas-Garcia R et al (2014) Characterization of the repeat expansion size in C9orf72 in amyotrophic lateral sclerosis and frontotemporal dementia. Hum Mol Genet 23:749–754. doi:10.1093/hmg/ddt460
Donnelly CJ, Zhang PW, Pham JT et al (2013) RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron 80:415–428. doi:10.1016/j.neuron.2013.10.015
Evans-Galea MV, Carrodus N, Rowley SM et al (2012) FXN methylation predicts expression and clinical outcome in Friedreich ataxia. Ann Neurol 71:487–497. doi:10.1002/ana.22671
Evans-Galea MV, Hannan AJ, Carrodus N, Delatycki MB, Saffery R (2013) Epigenetic modifications in trinucleotide repeat diseases. Trends Mol Med 19:655–663. doi:10.1016/j.molmed.2013.07.007
Farg MA, Sundaramoorthy V, Sultana JM et al (2014) C9ORF72, implicated in amytrophic lateral sclerosis and frontotemporal dementia, regulates endosomal trafficking. Hum Mol Genet. doi:10.1093/hmg/ddu068
Felle M, Hoffmeister H, Rothammer J, Fuchs A, Exler JH, Langst G (2011) Nucleosomes protect DNA from DNA methylation in vivo and in vitro. Nucleic Acids Res 39:6956–6969. doi:10.1093/nar/gkr263
Gallagher MD, Suh E, Grossman M et al (2014) TMEM106B is a genetic modifier of frontotemporal lobar degeneration with C9orf72 hexanucleotide repeat expansions. Acta Neuropathol 127:407–418. doi:10.1007/s00401-013-1239-x
Gibbs JR, van der Brug MP, Hernandez DG et al (2010) Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet 6:e1000952. doi:10.1371/journal.pgen.1000952
Gijselinck I, Van Langenhove T, van der Zee J et al (2012) A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol 11:54–65. doi:10.1016/S1474-4422(11)70261-7
Goldman JS, Quinzii C, Dunning-Broadbent J et al (2014) Multiple system atrophy and amyotrophic lateral sclerosis in a family with hexanucleotide repeat expansions in C9orf72. JAMA Neurol. doi:10.1001/jamaneurol.2013.5762
Gorno-Tempini ML, Hillis AE, Weintraub S et al (2011) Classification of primary progressive aphasia and its variants. Neurology 76:1006–1014. doi:10.1212/WNL.0b013e31821103e6
Goula AV, Stys A, Chan JP, Trottier Y, Festenstein R, Merienne K (2012) Transcription elongation and tissue-specific somatic CAG instability. PLoS Genet 8:e1003051. doi:10.1371/journal.pgen.1003051
Haeusler AR, Donnelly CJ, Periz G et al (2014) C9orf72 nucleotide repeat structures initiate molecular cascades of disease. Nature 507:195–200. doi:10.1038/nature13124
Hagerman RJ, Hull CE, Safanda JF et al (1994) High functioning fragile X males: demonstration of an unmethylated fully expanded FMR-1 mutation associated with protein expression. Am J Med Genet 51:298–308. doi:10.1002/ajmg.1320510404
Harms M, Benitez BA, Cairns N et al (2013) C9orf72 hexanucleotide repeat expansions in clinical Alzheimer disease. JAMA Neurol 70:736–741. doi:10.1001/2013.jamaneurol.537
Hensman Moss DJ, Poulter M, Beck J et al (2014) C9orf72 expansions are the most common genetic cause of Huntington disease phenocopies. Neurology 82:292–299. doi:10.1212/WNL.0000000000000061
Hsiung GY, DeJesus-Hernandez M, Feldman HH et al (2012) Clinical and pathological features of familial frontotemporal dementia caused by C9ORF72 mutation on chromosome 9p. Brain 135:709–722. doi:10.1093/brain/awr354
Hubers A, Marroquin N, Schmoll B et al (2014) Polymerase chain reaction and Southern blot-based analysis of the C9orf72 hexanucleotide repeat in different motor neuron diseases. Neurobiol Aging 35(1214):e1211–e1216. doi:10.1016/j.neurobiolaging.2013.11.034
Irwin DJ, McMillan CT, Brettschneider J et al (2013) Cognitive decline and reduced survival in C9orf72 expansion frontotemporal degeneration and amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry 84:163–169. doi:10.1136/jnnp-2012-303507
Kerkel K, Spadola A, Yuan E et al (2008) Genomic surveys by methylation-sensitive SNP analysis identify sequence-dependent allele-specific DNA methylation. Nat Genet 40:904–908. doi:10.1038/ng.174
Khan BK, Yokoyama JS, Takada LT et al (2012) Atypical, slowly progressive behavioural variant frontotemporal dementia associated with C9ORF72 hexanucleotide expansion. J Neurol Neurosurg Psychiatry 83:358–364. doi:10.1136/jnnp-2011-301883
Kwon I, Xiang S, Kato M et al (2014) Poly-dipeptides encoded by the C9ORF72 repeats bind nucleoli, impede RNA biogenesis, and kill cells. Science. doi:10.1126/science.1254917
Lagier-Tourenne C, Baughn M, Rigo F et al (2013) Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proc Natl Acad Sci 110:E4530–E4539. doi:10.1073/pnas.1318835110
Laird NM, Ware JH (1982) Random-effects models for longitudinal data. Biometrics 38:963–974
Lee YB, Chen HJ, Peres JN et al (2013) Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Rep 5:1178–1186. doi:10.1016/j.celrep.2013.10.049
Lesage S, Le Ber I, Condroyer C et al (2013) C9orf72 repeat expansions are a rare genetic cause of parkinsonism. Brain 136:385–391. doi:10.1093/brain/aws357
Levine TP, Daniels RD, Gatta AT, Wong LH, Hayes MJ (2013) The product of C9orf72, a gene strongly implicated in neurodegeneration, is structurally related to DENN Rab-GEFs. Bioinformatics 29:499–503. doi:10.1093/bioinformatics/bts725
Libby RT, Hagerman KA, Pineda VV et al (2008) CTCF cis-regulates trinucleotide repeat instability in an epigenetic manner: a novel basis for mutational hot spot determination. PLoS Genet 4:e1000257. doi:10.1371/journal.pgen.1000257
Liu EY, Russ J, Wu K et al (2014) C9orf72 hypermethylation protects against repeat expansion-associated pathology in ALS/FTD. Acta Neuropathol. doi:10.1007/s00401-014-1286-y
Mahoney CJ, Beck J, Rohrer JD et al (2012) Frontotemporal dementia with the C9ORF72 hexanucleotide repeat expansion: clinical, neuroanatomical and neuropathological features. Brain 135:736–750. doi:10.1093/brain/awr361
May S, Hornburg D, Schludi MH et al (2014) C9orf72 FTLD/ALS-associated Gly-Ala dipeptide repeat proteins cause neuronal toxicity and Unc119 sequestration. Acta Neuropathol. doi:10.1007/s00401-014-1329-4
McConkie-Rosell A, Lachiewicz AM, Spiridigliozzi GA et al (1993) Evidence that methylation of the FMR-I locus is responsible for variable phenotypic expression of the fragile X syndrome. Am J Hum Genet 53:800–809
Merenstein SA, Sobesky WE, Taylor AK, Riddle JE, Tran HX, Hagerman RJ (1996) Molecular-clinical correlations in males with an expanded FMR1 mutation. Am J Med Genet 64:388–394. doi:10.1002/(SICI)1096-8628(19960809)64:2<388:AID-AJMG31>3.0.CO;2-9
Mizielinska S, Gronke S, Niccoli T et al (2014) C9orf72 repeat expansions cause neurodegeneration in Drosophila through arginine-rich proteins. Science. doi:10.1126/science.1256800
Mori K, Lammich S, Mackenzie IR et al (2013) hnRNP A3 binds to GGGGCC repeats and is a constituent of p62-positive/TDP43-negative inclusions in the hippocampus of patients with C9orf72 mutations. Acta Neuropathol 125:413–423. doi:10.1007/s00401-013-1088-7
Mori K, Weng SM, Arzberger T et al (2013) The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339:1335–1338. doi:10.1126/science.1232927
Murray ME, DeJesus-Hernandez M, Rutherford NJ et al (2011) Clinical and neuropathologic heterogeneity of c9FTD/ALS associated with hexanucleotide repeat expansion in C9ORF72. Acta Neuropathol 122:673–690. doi:10.1007/s00401-011-0907-y
Murray ME, Bieniek KF, Banks Greenberg M et al (2013) Progressive amnestic dementia, hippocampal sclerosis, and mutation in C9ORF72. Acta Neuropathol 126:545–554. doi:10.1007/s00401-013-1161-2
Rascovsky K, Hodges JR, Knopman D et al (2011) Sensitivity of revised diagnostic criteria for the behavioural variant of frontotemporal dementia. Brain 134:2456–2477. doi:10.1093/brain/awr179
Renton AE, Majounie E, Waite A et al (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72:257–268. doi:10.1016/j.neuron.2011.09.010
Sareen D, O’Rourke JG, Meera P et al (2013) Targeting RNA foci in iPSC-derived motor neurons from ALS patients with a C9ORF72 repeat expansion. Sci Transl Med 5:208ra149. doi:10.1126/scitranslmed.3007529
Simon-Sanchez J, Dopper EG, Cohn-Hokke PE et al (2012) The clinical and pathological phenotype of C9ORF72 hexanucleotide repeat expansions. Brain 135:723–735. doi:10.1093/brain/awr353
Snowden JS, Rollinson S, Thompson JC et al (2012) Distinct clinical and pathological characteristics of frontotemporal dementia associated with C9ORF72 mutations. Brain 135:693–708. doi:10.1093/brain/awr355
Strong MJ, Grace GM, Freedman M et al (2009) Consensus criteria for the diagnosis of frontotemporal cognitive and behavioural syndromes in amyotrophic lateral sclerosis. Amyotroph Lateral Scler 10:131–146
Toledo JB, Van Deerlin VM, Lee EB et al (2013) A platform for discovery: the University of Pennsylvania integrated neurodegenerative disease biobank. Alzheimers Dement. doi:10.1016/j.jalz.2013.06.003
van Blitterswijk M, DeJesus-Hernandez M, Niemantsverdriet E et al (2013) Association between repeat sizes and clinical and pathological characteristics in carriers of C9ORF72 repeat expansions (Xpansize-72): a cross-sectional cohort study. Lancet Neurol 12:978–988. doi:10.1016/S1474-4422(13)70210-2
van Blitterswijk M, Mullen B, Nicholson AM et al (2014) TMEM106B protects C9ORF72 expansion carriers against frontotemporal dementia. Acta Neuropathol 127:397–406. doi:10.1007/s00401-013-1240-4
Waite AJ, Baumer D, East S et al (2014) Reduced C9orf72 protein levels in frontal cortex of amyotrophic lateral sclerosis and frontotemporal degeneration brain with the C9ORF72 hexanucleotide repeat expansion. Neurobiol Aging 35:1779 e1775–1779 e1713. doi:10.1016/j.neurobiolaging.2014.01.016
Xi Z, Zinman L, Moreno D et al (2013) Hypermethylation of the CpG island near the G4C2 repeat in ALS with a C9orf72 expansion. Am J Hum Genet 92:981–989. doi:10.1016/j.ajhg.2013.04.017
Xi Z, Rainero I et al (2014) Hypermethylation of the CpG-island near the C9orf72 G4C2-repeat expansion in FTLD patients. Hum Mol Genet. doi:10.1093/hmg/ddu279
Xie SX, Baek Y, Grossman M et al (2011) Building an integrated neurodegenerative disease database at an academic health center. Alzheimers Dement 7(4):e84–93. doi:10.1016/j.jalz.2010.08.233
Xu Z, Poidevin M, Li X et al (2013) Expanded GGGGCC repeat RNA associated with amyotrophic lateral sclerosis and frontotemporal dementia causes neurodegeneration. Proc Natl Acad Sci 110:7778–7783. doi:10.1073/pnas.1219643110
Zhang D, Iyer LM, He F, Aravind L (2012) Discovery of novel DENN proteins: implications for the evolution of eukaryotic intracellular membrane structures and human disease. Front Genet 3:283. doi:10.3389/fgene.2012.00283
Acknowledgments
We thank the patients and patients’ families who made this research possible. We acknowledge Drs. J. Q. Trojanowski and V. M.-Y. Lee and the Center for Neurodegenerative Disease Research for their support. This study was supported in part by a grant from the Judith & Jean Pape Adams Foundation and by the National Institutes of Health (K08AG039510, P30AG10124, P01AG017586, P01AG032953, P50AG005681). EBL is supported by a Doris Duke Charitable Foundation Clinical Scientist Development Award.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Russ, J., Liu, E.Y., Wu, K. et al. Hypermethylation of repeat expanded C9orf72 is a clinical and molecular disease modifier. Acta Neuropathol 129, 39–52 (2015). https://doi.org/10.1007/s00401-014-1365-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-014-1365-0