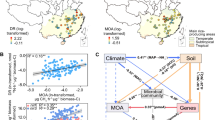
Abstract
Conventional aerobic methanotrophs oxidize methane (CH4) and covert CH4-derived carbon (C) into biomass at the oxic-anoxic interface of inundated rice paddy fields, playing indispensable role in mitigating greenhouse gas emissions and loss of organic C from methanogenesis. Two phylogenetically distinct groups of methanotrophs, type I (γ-proteobacteria) and type II (α-proteobacteria) methanotrophs, often co-exist in rice paddy soil and compete for CH4 biotransformation. Since these two methanotrophic groups also possess differential kinetics of CH4 oxidation and pathways of C assimilation, the consequence of their niche differentiation and metabolic differences in soil is expected to affect the CH4 oxidation rate and C conversion efficiency. Here, we examined the microbiology, chemistry, and CH4 metabolism in 24 geographically different paddy soils, covering four climate zones of eastern China. High-throughput sequencing of pmoA gene displayed a clear separation of in situ methanotrophic compositions between temperate (warm and mid-temperate) and warmer (subtropics and tropics) climate zones, likely driven by soil pH. Both methanotrophic groups were detected in soils but proportions of type I methanotrophs increased in temperate soils of higher pH (accounting for 76.1 ± 12.4% and 44.1 ± 14.8% in warm temperate and mid-temperate, respectively). Type II methanotrophs prevailed in warmer zones (accounting for 66.2 ± 21.6% and 70.5 ± 12.1% in tropics and subtropics, respectively) where soils were more acidic. Higher incorporation of 13C for synthesis in C14+C16 PLFAs (63.1–93.4% of total production of 13C-PLFAs) was found based on microcosm incubation, reflecting type I methanotrophs dominated the CH4 assimilation in paddy soils. Particularly, temperate soils with increased proportions of type I methanotrophs showed higher CH4 oxidation rate and C conversion efficiency. Collectively, this study depicts a continental-scale disparity of methanotrophic dynamics that tightly associates with consequence of niche differentiation of different types of methanotrophs and highlights the importance of microbiological control to maximize the rate and efficiency of methanotrophy.






Similar content being viewed by others
Data availability
Data are available from the corresponding author on request.
Change history
28 December 2023
A Correction to this paper has been published: https://doi.org/10.1007/s00374-023-01784-8
References
Bodelier PL, Bär Gillisen MJ, Hordijk K, Sinninghe Damsté JS, Rijpstra WIC, Geenevasen JA, Dunfield PF (2009) A reanalysis of phospholipid fatty acids as ecological biomarkers for methanotrophic bacteria. ISME J 3:606–617. https://doi.org/10.1038/ismej.2009.6
Bodelier PLE, Meima-Franke M, Hordijk CA, Steenbergh AK, Hefting MM, Bodrossy L, von Bergen M, Seifert J (2013) Microbial minorities modulate methane consumption through niche partitioning. ISME J 7:2214–2228. https://doi.org/10.1038/ismej.2013.99
Boschker HTS, Vasquez-Cardenas D, Bolhuis H, Moerdijk-Poortvliet TWC, Moodley L (2014) Chemoautotrophic carbon fixation rates and active bacterial communities in intertidal marine sediments. PLoS ONE 9:e101443. https://doi.org/10.1371/journal.pone.0101443
Cai Y, Zheng Y, Bodelier PLE, Conrad R, Jia Z (2016) Conventional methanotrophs are responsible for atmospheric methane oxidation in paddy soils. Nat Commun 7:1–10. https://doi.org/10.1038/ncomms11728
Chen X, Hu Y, Xia Y, Zheng S, Ma C, Rui Y, He H, Huang D, Zhang Z, Ge T, Wu J, Guggenberger G, Su Kuzyakov Y, Y, (2021) Contrasting pathways of carbon sequestration in paddy and upland soils. Glob Change Biol 27:2478–2490. https://doi.org/10.1111/gcb.15595
Chen Y, Dumont MG, McNamara NP, Chamberlain PM, Bodrossy L, Stralis-Pavese N, Murrell JC (2008) Diversity of the active methanotrophic community in acidic peatlands assessed by mRNA and SIP-PLFA analyses. Environ Microbiol 10:446–459. https://doi.org/10.1111/j.1462-2920.2007.01466.x
Conrad R, Rothfuss F (1991) Methane oxidation in the soil surface layer of a flooded rice field and the effect of ammonium. Biol Fert Soils 12:28–32. https://doi.org/10.1007/bf00369384
Costello AM, Auman AJ, Macalady JL, Scow KM, Lidstrom ME (2002) Estimation of methanotroph abundance in a freshwater lake sediment. Environ Microbiol 4:443–450. https://doi.org/10.1046/j.1462-2920.2002.00318.x
Degelmann DM, Borken W, Drake HL, Kolb S (2010) Different atmospheric methane-oxidizing communities in European beech and Norway spruce soils. Appl Environ Microbiol 76:3228–3235. https://doi.org/10.1128/aem.02730-09
Dumont MG, Lüke C, Deng Y, Frenzel P (2014) Classification of pmoA amplicon pyrosequences using BLAST and the lowest common ancestor method in MEGAN. Front Microbiol 5:77–79. https://doi.org/10.3389/fmicb.2014.00034
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194. https://doi.org/10.1093/bioinformatics/btr381
Eller G, Frenzel P (2001) Changes in activity and community structure of methane-oxidizing bacteria over the growth period of rice. Appl Environ Microb 67:2395–2403. https://doi.org/10.1128/aem.67.6.2395-2403.2001
Eller G, Krüger M, Frenzel P (2005) Comparing field and microcosm experiments: a case study on methano- and methylo-trophic bacteria in paddy soil. FEMS Microbiol Ecol 51:279–291. https://doi.org/10.1016/j.femsec.2004.09.007
Evershed RP, Crossman ZM, Bull ID, Mottram H, Dungait JAJ, Maxfield PJ, Brennand EL (2006) 13C-labelling of lipids to investigate microbial communities in the environment. Curr Opin Biotechnol 17:72–82. https://doi.org/10.1016/j.copbio.2006.01.003
Fish JA, Chai B, Wang Q, Sun Y, Titus BC, Tiedje JM (2013) FunGene: the functional gene pipeline and repository. Front Microbiol 4:291. https://doi.org/10.3389/fmicb.2013.00291
Frenzel P, Rothfuss F, Conrad R (1992) Oxygen profiles and methane turnover in a flooded rice microcosm. Biol Fert Soils 14:84–89. https://doi.org/10.1007/bf00336255
Frostegård Å, Bååth E, Tunlio A (1993) Shifts in the structure of soil microbial communities in limed forests as revealed by phospholipid fatty acid analysis. Soil Biol Biochem 25:723–730. https://doi.org/10.1016/0038-0717(93)90113-p
Gao D, Sheng R, Whiteley AS, Moreira-Grez B, Wei W (2020) Effect of phosphorus amendments on rice rhizospheric methanogens and methanotrophs in a phosphorus deficient soil. Geoderma 368:114312. https://doi.org/10.1016/j.geoderma.2020.114312
Hall EK, Bernhardt ES, Bier RL, Bradford MA, Boot CM, Cotner JB, del Giorgio PA, Evans SE, Graham EB, Jones SE, Lennon JT, Locey KJ, Nemergut D, Osborne BB, Rocca JD, Schimel JP, Waldrop MP, Wallensteinet MD (2018) Understanding how microbiomes influence the systems they inhabit. Nat Microbiol 3:977–982. https://doi.org/10.1038/s41564-018-0201-z
Hanson RS, Hanson TE (1996) Methanotrophic bacteria. Microbiol Rev 2:439–471. https://doi.org/10.1128/mr.60.2.439-471.1996
He R, Wooller MJ, Pohlman JW, Tiedje JM, Leigh MB (2015) Methane-derived carbon flow through microbial communities in arctic lake sediments. Environ Microbiol 17:3233–3250. https://doi.org/10.1111/1462-2920.12773
Hedlund K (2002) Soil microbial community structure in relation to vegetation management on former agricultural land. Soil Biol Biochem 34:1299–1307. https://doi.org/10.1016/s0038-0717(02)00073-1
Henneberger R, Chiri E, Bodelier PLE, Frenzel P, Lüke C, Schroth MH (2015) Field-scale tracking of active methane-oxidizing communities in a landfill-cover soil reveals spatial and seasonal variability. Environ Microbiol 17:1721–1737. https://doi.org/10.1111/1462-2920.12617
Ho A, Kerckhof FM, Luke C, Reim A, Krause S, Boon N, Bodelier PLE (2013) Conceptualizing functional traits and ecological characteristics of methane-oxidizing bacteria as life strategies. Environ Microbiol Rep 5:335–345. https://doi.org/10.1111/j.1758-2229.2012.00370.x
Ho A, Lee HJ, Reumer M, Meima-Franke M, Raaijmakers C, Zweers H, de Boer W, Van der Puttend WH, Bodelier PLE (2018) Unexpected role of canonical aerobic methanotrophs in upland agricultural soils. Soil Biol Biochem 131:1–8. https://doi.org/10.1016/j.soilbio.2018.12.020
Ho A, Lüke C, Frenzel P (2011) Recovery of methanotrophs from disturbance: population dynamics evenness and functioning. ISME J 5:750–758. https://doi.org/10.1038/ismej.2010.163
IPCC (2013). In: Stocker TF (ed) Contribution of Working Group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change. Press, Cambridge, UK, pp 465–570. https://doi.org/10.1017/cbo9781107415324.014
Joergensen RG (2022) Phospholipid fatty acids in soil—drawbacks and future prospects. Biol Fertil Soils 58:1–6. https://doi.org/10.1007/s00374-021-01613-w
Kalyuzhnaya MG, Yang S, Rozova ON, Smalley NE, Clubb J, Lamb A, Nagana Gowda GA, Raftery D, Fu Y, Bringel F, Vuilleumier S, Beck DAC, Trotsenko YA, Khmelenina VN, Lidstrom ME (2013) Highly efficient methane biocatalysis revealed in a methanotrophic bacterium. Nat Commun 4:2785. https://doi.org/10.1038/ncomms3785
Knief C (2015) Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front Microbiol 6:1–38. https://doi.org/10.3389/fmicb.2015.01346
Kögel-Knabner I, Amelung W, Cao Z, Fiedler S, Frenzel P, Jahn R, Kalbitz K, Kölbl A, Schloter M (2010) Biogeochemistry of paddy soils. Geoderma 157:1–14. https://doi.org/10.1016/j.geoderma.2010.03.009
Kou Y, Li J, Wang Y, Li C, Tu B, Yao M, Li X (2017) Scale-dependent key drivers controlling methane oxidation potential in Chinese grassland soils. Soil Biol Biochem 111:104–114. https://doi.org/10.1016/j.soilbio.2017.04.005
Kou Y, Wei K, Li C, Wang Y, Yao M (2020) Deterministic processes dominate soil methanotrophic community assembly in grassland soils. Geoderma 359:114004. https://doi.org/10.1016/j.geoderma.2019.114004
Krause S, Luke C, Frenzel P (2012) Methane source strength and energy flow shape methanotrophic communities in oxygen-methane counter-gradients. Environ Microbiol Rep 4:203–208. https://doi.org/10.1111/j.1758-2229.2011.00322.x
Lehmann J, Hansel CM, Kaiser C, Kleber M, Maher K, Manzoni S, Nunan N, Reichstein M, Schimel JP, Torn MS, Wieder WR, Kögel-Knabner I (2020) Persistence of soil organic carbon caused by functional complexity. Nat Geosci 13:529–534. https://doi.org/10.1038/s41561-020-0612-3
Lüke C, Frenzel P (2011) Potential of pmoA amplicon pyrosequencing for methanotroph diversity studies. Appl Environ Microb 77:6305–6309. https://doi.org/10.1128/aem.05355-11
Mohanty SR, Bodelier PLE, Floris V, Conrad R (2006) Differential effects of nitrogenous fertilizers on methane-consuming microbes in rice field and forest soils. Appl Environ Microbiol 72:1346–1354. https://doi.org/10.1128/aem.72.2.1346-1354.2006
Nouchi I, Hosono T, Aoki K, Minami K (1994) Seasonal variation in methane flux from rice paddies associated with methane concentration in soil water rice biomass and temperature and its modeling. Plant Soil 161:195–208. https://doi.org/10.1007/BF00046390
Price MN, Dehal PS, Arkin AP (2010) FastTree 2 — approximately maximum-likelihood trees for large alignments. PLoS ONE 5:e9490. https://doi.org/10.1371/journal.pone.0009490
Qiu QF, Noll M, Abraham WR, Lu YH, Conrad R (2008) Applying stable isotope probing of phospholipid fatty acids and rRNA in a Chinese rice field to study activity and composition of the methanotrophic bacterial communities in situ. ISME J 2:602–614. https://doi.org/10.1038/ismej.2008.34
Scheutz C, Kjeldsen P, Bogner JE, De Visscher A, Gebert J, Hilger HA, Huber-Humer M, Spokas K (2009) Microbial methane oxidation processes and technologies for mitigation of landfill gas emissions. Waste Manage Res 27:409–455. https://doi.org/10.1177/0734242x09339325
Shrestha M, Shrestha PM, Frenzel P, Conrad R (2010) Effect of nitrogen fertilization on methane oxidation abundance community structure and gene expression of methanotrophs in the rice rhizosphere. ISME J 4:1545–1556. https://doi.org/10.1038/ismej.2010.89
Sullivan B, Selmants PC, Hart SC (2013) Does dissolved organic carbon regulate biological methane oxidation in semiarid soils? Glob Change Biol 19:2149–2157. https://doi.org/10.1111/gcb.12201
Sultana N, Zhao J, Cai Y, Rahman GKMM, Alam MS, Faheem M, Ho A, Jia Z (2022) Methanotrophy-driven accumulation of organic carbon in four paddy soils of Bangladesh. Pedosphere 32:348–358. https://doi.org/10.1016/s1002-0160(20)60030-3
Sultana N, Zhao J, Zheng Y, Cai Y, Faheem M, Peng X, Wang W, Jia Z (2019) Stable isotope probing of active methane oxidizers in rice field soils from cold regions. Biol Fertil Soils 55:243–250. https://doi.org/10.1007/s00374-018-01334-7
Trimmer M, Shelley FC, Purdy KJ, Maanoja ST, Chronopoulou PM, Grey J (2015) Riverbed methanotrophy sustained by high carbon conversion efficiency. ISME J 9:2304–2314. https://doi.org/10.1038/ismej.2015.98
Trotsenko YA, Murrell JC (2008) Metabolic aspects of aerobic obligate methanotrophy. Adv Appl Microbiol 63:183–229. https://doi.org/10.1016/s0065-2164(07)00005-6
Wang Q, Quensen JF, Fish JA, Lee TK, Sun Y, Tiedje JM, Cole JR (2013) Ecological patterns of nifH genes in four terrestrial climatic zones explored with targeted metagenomics using FrameBot a new informatics tool. mBio 4:00592–00513. https://doi.org/10.1128/mbio.00592-13
Xia Y, Chen X, Hu Y, Zheng S, Ning Z, Guggenberger G, He H, Wu J, Su Y (2019) Contrasting contribution of fungal and bacterial residues to organic carbon accumulation in paddy soils across eastern China. Biol Fertil Soils 55:767–776. https://doi.org/10.1007/s00374-019-01390-7
Zhang L, Adams JM, Dumont MG, Li Y, Shi Y, He D, He J-S, Chu H (2019) Distinct methanotrophic communities exist in habitats with different soil water contents. Soil Biol Biochem 132:143–152. https://doi.org/10.1016/j.soilbio.2019.02.007
Zhang L, Dumont MG, Bodelier PLE, Adams JM, Chu H (2020) DNA stable-isotope probing highlights the effects of temperature on functionally active methanotrophs in natural wetlands. Soil Biol Biochem 149:107954. https://doi.org/10.1016/j.soilbio.2020.107954
Zhao J, Cai Y, Jia Z (2020) The pH-based ecological coherence of active canonical methanotrophs in paddy soils. Biogeosciences 17:1451–1462. https://doi.org/10.5194/bg-17-1451-2020
Zheng S, Xia Y, Hu Y, Chen X, Rui Y, Gunina A, He X, Ge T, Wu J, Su Y, Kuzyakov Y (2021) Stoichiometry of carbon nitrogen and phosphorus in soil: effects of agricultural land use and climate at a continental scale. Soil Tillage Res 209:104903. https://doi.org/10.1016/j.still.2020.104903
Zhu Y, Purdy KJ, Eyice Ö, Shen L, Harpenslager SF, Yvon-Durocher G, Dumbrell AJ, Trimmer M (2020) Disproportionate increase in freshwater methane emissions induced by experimental warming. Nat Clim Change 10:1–6. https://doi.org/10.1038/s41558-020-0824-y
Funding
This work was financially supported by the National Key Research Program (2021YFD1901204, 2021YFD1901203), National Natural Science Foundation of China (grant numbers 42377348, 42177295), Natural Science Foundation of Fujian Province (2022J05263), Nanping City Science and Technology Plan Project (NP2021KTS02), Talent Introduction Project of Wuyi University (YJ202117), and Open Foundation of State Key Laboratory of Microbial Technology in Shandong University (M2022-05).
Author information
Authors and Affiliations
Contributions
J.S.W., Y.R.S., and X.B.C. conceived and designed the experiments. S.M.Z., S.H.D., C.M., Y.H.X., and H.Q. performed the experiments and collected the data. S.M.Z. and S.H.D. analyzed the data. S.M.Z. illustrated the figures and wrote the first draft of the manuscript. J.Z., W.G., Q.T., Y.M.Z., Y.C.R., J.S.W., Y.R.S., and X.B.C. revised the manuscript. All authors contributed to the final draft of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: The article title has been corrected.
Supplementary information
ESM 1
(DOCX 1006 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zheng, S., Deng, S., Ma, C. et al. Type I methanotrophs dominated methane oxidation and assimilation in rice paddy fields by the consequence of niche differentiation. Biol Fertil Soils 60, 153–165 (2024). https://doi.org/10.1007/s00374-023-01773-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-023-01773-x