Abstract
Introduction
The genomic revolution has transformed our understanding of urinary tract infection. There has been a paradigm shift from the dogmatic statement that urine is sterile in healthy people, as we are becoming forever more familiar with the knowledge that bacterial communities exist within the urinary tracts of healthy people. Metagenomics can investigate the broad populations of microbial communities, analysing all the DNA present within a sample, providing comprehensive data regarding the state of the microenvironment of a patient’s urinary tract. This permits medical practitioners to more accurately target organisms that may be responsible for disease—a form of ‘precision medicine’.
Methods and Results
This paper is derived from an extensive review and analysis of the available literature on the topic of metagenomic sequencing in urological science, using the PubMed search engine. The search yielded a total of 406 results, and manual selection of appropriate papers was subsequently performed. Only one randomised clinical trial comparing metagenomic sequencing to standard culture and sensitivity in the arena of urinary tract infection was found.
Conclusion
Out of this process, this paper explores the limitations of traditional methods of culture and sensitivity and delves into the recent studies involving new high-throughput genomic technologies in urological basic and clinical research, demonstrating the advances made in the urinary microbiome in its entire spectrum of pathogens and the first attempts of clinical implementation in several areas of urology. Finally, this paper discusses the challenges that must be overcome for such technology to become widely used in clinical practice.

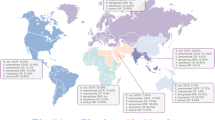
Adapted from Knief C [4]
Similar content being viewed by others
References
Mouraviev V, McDonald MW (2018) An implementation of next generation sequencing for prevention and diagnosis of urinary tract infection in urology. Can J Urol 25:9349–9356
Forbes JD, Knox NC, Ronholm J et al (2017) Metagenomics: the next culture-independent game changer. Front Microbiol 8:1069
American Society for Microbiology (2016) Applications of clinical microbial next-generation sequencing: report on an American Academy of Microbiology Colloquium held in Washington, DC, in April 2015. American Academy of Microbiology https://www.ncbi.nlm.nih.gov/books/NBK513764/ Accessed 1 Dec 2018
Knief C (2014) Analysis of plant microbe interactions in the era of next generation sequencing technologies. Front Plant Sci 5:206
Rizzo JM, Buck MJ (2012) Key principles and clinical applications of “Next-Generation” DNA sequencing. Cancer Prev Res 5:887–900
Wellinghausen N, Kochem AJ, Disqué C et al (2009) Diagnosis of bacteremia in whole-blood samples by use of a commercial universal 16S rRNA gene-based PCR and sequence analysis. J Clin Microbiol 47:2759–2765
Prachayangprecha S, Schapendonk CM, Koopmans MP et al (2014) Exploring the potential of next-generation sequencing in detection of respiratory viruses. J Clin Microbiol 52:3722–3730
McDonald MW, Kameh D, Johnson ME et al (2017) A head-to-head comparative phase ii study of standard urine culture and sensitivity versus DNA next-generation sequencing testing for urinary tract infections. Rev Urol 19:213–220
Wagenlehner FM, Abramov-Sommariva D, Höller M et al (2018) Non-Antibiotic Herbal Therapy (BNO 1045) versus antibiotic therapy (fosfomycin trometamol) for the treatment of acute lower uncomplicated urinary tract infections in women: a double-blind, parallel-group, randomized, multicentre, non-inferiority phase III trial. Urol Int 101:327–336
Elshal AM, Atwa AM, El-Nahas AR et al (2018) Chemoprophylaxis during transrectal prostate needle biopsy: critical analysis through randomized clinical trial. World J Urol 36:1845–1852
Mouraviev V, McDonald MW, Skinner C et al (2018) MP15-14 the value of next generation dna sequencing testing in rectal swabs before transrectal prostate biopsy for individual and targeted prophylaxis of urinary tract infection. J Urol 199(4):e194
Sfanos KS, Isaacs WB, De Marzo AM (2013) Infections and inflammation in prostate cancer. Am J Clin Exp Urol 1:3–11
Shrestha E, White JR, Yu SH et al (2017) Profiling the urinary microbiome in men with positive versus negative biopsies for prostate cancer. J Urol 199:161–171
Frugé AD, Ptacek T, Tsuruta Y et al (2018) Dietary changes impact the gut microbe composition in overweight and obese men with prostate cancer undergoing radical prostatectomy. J Acad Nutr Diet 118:714–714.e1
Cavarretta I, Ferrarese R, Cazzaniga W et al (2017) The microbiome of the prostate tumor microenvironment. Eur Urol 72:625–631
Krieger JN, Nyberg L Jr, Nickel JC (1999) NIH consensus definition and classification of prostatitis. JAMA 282:236–237
Shoskes DA, Wang H, Polackwich AS et al (2016) Analysis of gut microbiome reveals significant differences between men with chronic prostatitis/chronic pelvic pain syndrome and controls. J Urol 196:435–441
Karstens L, Asquith M, Davin S et al (2016) Does the urinary microbiome play a role in urgency urinary incontinence and its severity? Front Cell Infect Microbiol 6:78
Pearce MM, Zilliox MJ, Rosenfeld AB et al (2015) The female urinary microbiome in urgency urinary incontinence. Am J Obstet Gynecol 213:347.e1–347.e11
Brubaker L, Nager CW, Richter HE et al (2014) Urinary bacteria in adult women with urgency urinary incontinence. Int Urogynaecol J 25:1179–1184
Pearce MM, Hilt EE, Rosenfeld AB et al (2014) The female urinary microbiome: a comparison of women with and without urgency urinary incontinence. mBio 5:e01283-14
Fouts DE, Pieper R, Szpakowski S et al (2012) Integrated next-generation sequencing of 16S rDNA and metaproteomics differentiate the healthy urine microbiome from asymptomatic bacteriuria in neuropathic bladder associated with spinal cord injury. J Transl Med 10:174
Mouraviev V, McDonald MW (2018) MP23-06 An utilization of next generation sequencing of urine samples for monitoring of urinary tract infection in patients with neurogenic bladder. J Urol 199:e283
Smelov V, Naber K, Bjerklund Johansen TE (2016) Improved classification of urinary tract infection: future consideration. Eur Urol Suppl 15:71–80
Hiergeist A, Gessner A (2017) Clinical implications of microbiome in urinary tract diseases. Curr Opin Urol 27:93–98
Pak TR, Kasarkis A (2015) How Next-Generation Sequencing and multiscale data analysis will transform infection disease management. Clin Infect Dis 61:1695–1702
Smelov V, Naber K, Bjerklund Johansen TE (2016) Letter to the editor: DIAGNOSTIC criteria in urological diseases do not always match with findings by extended culture techniques and metagenomics sequencing of 16S rDNA. Open Microbiol J 10:23–26
Davenport M, Mach KE, Shortliffe LMD et al (2017) New and developing diagnostic technologies for urinary tract infection. Nat Rev Urol 14:296–310
Onsongo G, Erdmann J, Spears MD et al (2014) Implementation of cloud based next generation sequencing data analysis in a clinical laboratory. BMC Res Notes 7:314
Aziz N, Zhao Q, Bry L et al (2015) College of American Pathologists’ laboratory standards for next-generation sequencing clinical tests. Arch Pathol Lab Med 139:481–493
Acknowledgements
The authors would like to thank Jennifer White, M.S., MB(ASCP)cm for her excellent technical assistance and thorough description of metagenomics and Next Generation Sequence technologies.
Author information
Authors and Affiliations
Contributions
M Dixon: manuscript writing/editing. M Stefil: manuscript writing/editing. M McDonald: principal investigator of clinical trial; interpretation of NGS results; manuscript writing/editing. TE Bjerklund Johansen: diagnostic results analysis; manuscript writing/editing. K Naber: diagnostic results analysis; manuscript writing/editing. F Wagenlehner: diagnostic results analysis; manuscript writing/editing. V Mouraviev: project development; interpretation of NGS results; manuscript writing/editing.
Corresponding author
Ethics declarations
Conflicts of interest
Dr Mouraviev is a consultant for MicroGenDX.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dixon, M., Stefil, M., McDonald, M. et al. Metagenomics in diagnosis and improved targeted treatment of UTI. World J Urol 38, 35–43 (2020). https://doi.org/10.1007/s00345-019-02731-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00345-019-02731-9