Abstract
Indole-3-acetic acid (IAA) amidosynthetases catalyzing the ATP-dependent conjugation of IAA and amino acids play an important role in the maintenance of auxin homeostasis in plant cells. A new amidosynthetase, indole-3-acetic acid:l-aspartic acid ligase (IAA-Asp synthetase) involved in IAA-amino acid biosynthesis, was isolated via a biochemical approach from immature seeds of the pea (Pisum sativum L). The enzyme was purified to homogeneity by a three-step procedure, involving PEG 6000 fractionation, DEAE-Sephacel anion-exchange chromatography, and preparative PAGE, and characterized as a 70-kDa monomeric protein by analytical gel filtration and SDS-PAGE. Rabbit antiserum against recombinant AtGH3.5 cross-reacted with the pea IAA-Asp synthetase, and a single immunoreactive polypeptide band was observed at 70 kDa. The purified enzyme had an apparent isoelectric point at pH 4.7, the highest activity at pH 8.2, preferred Mg2+ as a cofactor, and was strongly activated by reducing agents. Similar to known recombinant GH3 enzymes, an IAA-Asp synthetase from pea catalyzes the conjugation of phytohormone acyl substrates to amino acids. The enzyme had the highest synthesizing activity on IAA, followed by 1-NAA, SA, 2,4-D, and IBA, whereas activities on l-Trp, IPA, PAA, (±)JA, and 2-NAA were not significant or not detected. Of 14 amino acids tested, the enzyme had the highest activity on Asp and lower activity on Ala and Lys. Glutamate was found to be a very poor substrate and no conjugating activity was observed on the rest of the amino acids. Steady-state kinetic analysis indicated that IAA and aspartate were preferred substrates for the pea IAA-Asp synthetase. The enzyme exhibited both higher affinities for IAA and Asp (K m = 0.2 and 2.5 mM, respectively) and catalytic efficiencies (k cat/K m = 682,608.7 and 5080 s−1 M−1, respectively) compared with other auxins and amino acids examined. This study describes the first amidosynthetase isolated and purified from plant tissue and provides the foundation for future genetic approaches to explain the role of IAA-Asp in Pisum sativum physiology.
Similar content being viewed by others
Introduction
Indole-3-acetic acid (IAA) conjugation is known to be essential for maintenance of auxin homeostasis and, thus, plant growth and development (Woodward and Bartel 2005). Most of the IAA in plants is found as ester or amide conjugates, which are putative storage or inactive forms of the hormone (Bandurski and others 1995). IAA conjugation is widespread in the plant kingdom, and IAA conjugates have been found in both seeds and vegetative tissues (Bandurski and Schulze 1977; Cohen and Bandurski 1982; Sztein and others 1995). However, IAA conjugation patterns are distinct in different plant species, and IAA conjugate profiles can be regulated by tissue-specific as well as developmental processes (reviewed in Woodward and Bartel 2005).
Because of the potential involvement of IAA conjugates in hormone homeostasis, numerous studies have focused on conjugate synthesis and hydrolysis during plant growth and development. The enzymes and genes involved in metabolism of IAA-ester conjugates have been isolated and characterized in some species (Kowalczyk and Bandurski 1991; Jakubowska and others 1993; Szerszen and others 1994; Jackson and others 2001; Kowalczyk and others 2003; Jakubowska and Kowalczyk 2004, 2005). A relatively large amount of information is now available on the hydrolysis of the auxin-amino acid conjugates in plants (Bartel and Fink 1995; Ludwig-Müller and others 1996; Davies and others 1999; LeClere and others 2002, Campanella and others 2003a, b, 2004; Rampey and others 2004; Schuller and Ludwig-Müller 2006; Campanella and others 2008, Savić and others 2009) and some plant pathogens (Chou and others 1996, 1998; Chou and Huang 2005).
By contrast, much less information is available on the formation of the amide-linked conjugates. IAA-amino acid conjugate biosynthesis by an in vitro enzymatic system has been impossible to demonstrate for many years. Bacterial IAA-ε-lysine synthetase was the only characterized enzyme and its encoding DNA sequence was identified (Roberto and others 1990). The identification of the Arabidopsis thaliana JAR1 and several closely related genes that are involved in conjugation of different phytohormones such as jasmonic acid (JA), indole-3-acetic acid (IAA), or salicylic acid (SA) to amino acids has greatly advanced our understanding of the formation of IAA-amide conjugates (Staswick and others 2002, 2005). These genes are members of the AtGH3 multigene family and are related to the auxin-induced GH3 gene of soybean (Hagen and Guilfoyle 1985; Wang and others 2008). Besides being identified in Arabidopsis, the GH3 gene family members have been identified in many other plant species and in bacteria (Terol and others 2006; Wang and others 2008). Plant GH3 proteins belong to the firefly luciferase superfamily that consists of acyl-adenylate/thioester-forming enzymes, some of which were shown to possess adenylation activity on phytohormones such as IAA, JA, SA, or IBA (indole-3-butyric acid) (Staswick and others 2002, 2005; Ludwig-Müller and others 2009). They catalyze the ATP-dependent formation of hormone conjugates in a two-step reaction proceeding via an acyl-adenylate intermediate (Staswick and others 2002, 2005). The GH3 protein family has been well studied in Arabidopsis but less so in other species. Of the 19 Arabidopsis GH3 proteins, six (GH3.2, GH3.3, GH3.4, GH3.5, GH3.6, GH3.17) form IAA-amino acid conjugates (Staswick and others 2002, 2005). One (JAR1/GH3.11) conjugates only JA, not IAA (Staswick and others 2002); one (GH3.5) is active on both IAA and SA (Staswick and others 2005; Zhang and others 2007); and one (GH3.12) conjugates 4-substituted benzoates (Okrent and others 2009). Thus, these proteins are synthetases catalyzing the formation of amide-linked conjugates that are involved in hormone homeostasis. Many of these conjugates are inactive or degraded, but in the case of JA-Ile conjugation by JAR1, conjugation is necessary to activate the hormone (Staswick and others 2002). Recently, tryptophan conjugates of IAA and JA were also found to have biological activity, although it is not clear yet if GH3 proteins are involved (Staswick 2009).
Besides A. thaliana, two moss (Physcomitrella patens) GH3 proteins were described (Ludwig-Müller and others 2009). PpGH3-1 and PpGH3-2 convert IAA, JA, and IBA to amide conjugates but with different preferences for the attached amino acid. Recently, two rice (Oryza sativa) proteins, OsGH3.8 and TLD1/OsGH3.13, were shown to conjugate amino acids to IAA (Chen and others 2009; Zhang and others 2009). A tomato enzyme SlJAR1 conjugating JA has just been reported (Suza and others 2010).
To date, enzyme activity of these proteins has been limited to analysis of recombinant proteins expressed and purified from Escherichia coli, whereas purification of the enzyme from plant tissue via a biochemical approach has not succeeded. In our previous study we demonstrated in vitro enzymatic synthesis of IAA-amino acid conjugates by crude extract from immature seeds of pea (Ostrowski and Jakubowska 2008). Here we present a purification procedure yielding the electrophoretically homogeneous protein of IAA-aspartate synthetase from pea immature seeds and detailed biochemical and kinetic analyses of this enzyme. To our knowledge, this is the first characterization of an enzyme conjugating IAA to amino acid isolated from plant tissue.
Materials and Methods
Plant Material and Sample Preparation
Pea (P. sativum) plants were grown under field conditions during the summer of 2007. The pods with maturating seeds were harvested at different stages of seed development. Dry seeds were purchased from a local market. Immature seeds were selected for analysis according to their diameter and were used directly. Dry seeds were imbibed for 4 h in water before analysis. For each experiment, 3 g of seeds were ground using a mortar and pestle in one volume of 50 mM Tris-HCl, pH 7.6, containing 2 mM EDTA and 5 mM 2-mercaptoethanol. The homogenate was filtered through a nylon cloth, and the filtrate was used for assay of IAA-aspartate synthetase activity. For enzyme purification, immature seeds 5 mm in diameter were selected, frozen, and stored in freezer bags at −20°C.
Purification of the IAA-Asp Synthetase
All steps were performed at 0–4°C. A 200-g portion of immature frozen pea seeds was homogenized with 200 ml of 50 mM Tris-HCl, pH 7.6, containing 2 mM EDTA and 5 mM 2-mercaptoethanol using a mortar and pestle and then a Polytron homogenizer. The homogenate was filtered through a nylon cloth, and after adjusting the pH to 7.6, it was centrifuged at 2000g for 20 min. The resulting supernatant was used in the fractionation with poly(ethylene glycol) 6000 (PEG 6000).
Step I. Fractionation with PEG 6000
Solid PEG 6000 was added slowly to the supernatant fluid to obtain a 20% (w/v) final concentration. The solution was gently stirred for 2 h and then centrifuged at 22,000g for 45 min.
Step II. DEAE-Sephacel Anion-Exchange Chromatography
After PEG 6000 fractionation, the cleared supernatant was applied to a DEAE-Sephacel column (2.5 × 20 cm) equilibrated with 25 mM Tris-HCl, pH 7.6, containing 5 mM 2-mercaptoethanol. The anionite was washed with the same buffer and then with 150 mM NaCl in this buffer until the A280 decreased to baseline. The bound proteins were eluted with 250 mM NaCl in the buffer at the flow rate of 0.9 ml min−1 and 8-ml fractions were collected. The fractions containing the most activity of IAA-Asp synthetase were pooled and concentrated by ultrafiltration using an Amicon Diaflo YM-10 filter (Millipore, Billerica, MA, USA).
Step III. Preparative Polyacrylamide Gel Electrophoresis
The concentrated fractions from step II were dialyzed against 25 mM Tris-HCl, pH 7.6, containing 5 mM 2-mercaptoethanol overnight and subjected to a preparative PAGE on 12% (w/v) running gel and 6% (w/v) stacking gel, according to the procedure of Ogita and Markert (1979). Proteins were separated in a 20-cm × 19-cm × 1-mm gel using a vertical slab electrophoresis apparatus STANDARD (Kucharczyk Electrophoretical Techniques, Poland). The sample was applied to a 40-mm-wide well, and after the electrophoresis, the gel was cut into three strips according to the direction of resolving. The outside strips were stained briefly with Coomassie Brilliant Blue G-250 in 3.5% (v/v) perchloric acid and 20% (v/v) methanol (Blakesly and Boezi 1977) and then washed with distilled water. The appropriate 3-mm-wide gel segment from the middle strip (not stained) was excised based on the position of the protein bands stained on the outside strips. This segment of the gel was transferred into a dialysis tube and placed in a horizontal electrophoresis apparatus. The proteins were electroeluted with the 12.5 mM Tris-glycine buffer, pH 8.3, at 20 mA for 12 h. Enzyme activity and proteins were determined in fractions obtained after electroelution. The fractions containing the most activity of IAA-Asp synthetase were pooled and concentrated by centrifugation at 1000g for 30 min through a Millipore filter.
Isoelectric Focusing
Isoelectric focusing (IEF) was performed with 2% (v/v) Ampholyte, pH 3–10 (Fluka), in 5% (w/v) polyacrylamide gel (10 cm × 10 cm × 1 mm) using a MiniProtean II apparatus (Bio-Rad, Hercules, CA, USA) according to the method of Bollag and Edelstein (1991). IEF was run at 200 V for 24 h at 4°C in 20 mM NaOH as catholite and 10 mM H3PO4 as anolite. IEF-MIX 3.6–9.3 (Sigma-Aldrich, Germany), containing amyloglucosidase (pI 3.6), trypsin inhibitor from soybean (pI 4.6), β-lactoglobulin A (pI 5.1), carbonic anhydrase II (pI 5.9), carbonic anhydrase I (pI 6.8), myoglobin (pI 7.2) and lectin from Lenus culinaris (pI 8.3, 8.6, 8.8), was used as a marker.
Enzyme Activity Assay
The standard reaction mixture contained 2 mM IAA, 10 mM l-Asp, 5 mM ATP, 5 mM MgCl2, and 2 mM DTT in 50 mM Tris-HCl, pH 8.2, and the enzyme preparation in a final volume of 8 μl. For radioactive assays, 0.05 μCi [14C]IAA (50 mCi mmol−1) or 0.05 μCi [14C]l-Asp (50 mCi mmol−1) (Hartmann Analytic GmBH, Germany) was added. To test the effect of metal cofactors on IAA-Asp synthesis, MgCl2, MnCl2, CuCl2, and Zn (CH3COO)2 were added to a 5 mM final concentration. The effect of the reducing agents 2-mercaptoethanol (ME), dithiotreitol (DTT), 1,4-dithioerytritol (DTE), glutathione oxidized (GSSG), glutathione reduced (GSH), and l-cysteine in a final concentration of 2 mM was also tested. After a 60-min incubation at 30°C, the time course of product accumulation in the reaction was analyzed on a Silica Gel F260 plate (Merck, Germany) developed in chloroform/ethyl acetate/formic acid (33/55/10 by volume) (Staswick and others 2005). Indole-ring compounds were visualized by staining the plate in Van Urk-Salkovski reagent (Ehmann 1977). When quantitative measurements of enzyme activity were necessary, the reaction mixture containing radioactive IAA or IAA-Asp was used. The time course of the reaction was monitored under short-wavelength UV. The UV absorbing region, identified as IAA-Asp, was excised and placed into a vial with 2 ml EcoLite (+) scintillation fluid (ICN, USA). Radioactivity was measured in a Wallac 1409 liquid scintillation counter. In the case of the compounds without an indole ring (substrate specificity assays), anion-exchange chromatography was used to separate the reaction products according to the method described by Zelená (2000). The reaction was stopped by adding of 500 μl cold 50% (v/v) ethanol. Then the mixture was centrifuged and 500 μl of the clear supernatant was applied on a 2-ml bed volume DEAE-Sephadex A-25 column in acetate form in 50% (v/v) ethanol. The anionite was washed with one volume of 50% (v/v) ethanol. Free IAA was eluted with 5% (v/v) acetic acid in 50% (v/v) ethanol, and amide conjugates were eluted with 3% (v/v) trifluoroacetic acid (TFA) in 50% (v/v) ethanol. Two milliliters of EcoLite (+) scintillation fluid was added to the 500 μl of eluted fractions for radioactivity measurement.
Substrate Specificity and Kinetic Studies
The partially purified enzyme preparation (after DEAE-Sephacel chromatography) was used for determination of the substrate specificity. The standard reaction mixture contained 2 mM IAA [IBA, IPA (indole-3-pyruvic acid), l-Trp, 1-NAA (1-naphthalacetic acid), 2-NAA, 2,4-D (2,4-dichlorophenoxyacetic acid), PAA (phenylacetic acid), SA, or (±)JA], 10 mM l-Asp, and 0.05 μCi [14C]l-Asp (50 mCi mmol−1) for determination of the activity toward auxins, or 10 mM l-Asp (l-Glu, l-Ala, l-Leu, l-Lys, l-Met, l-Phe, l-Tyr, l-Trp, l-Gly, l-Asn, l-Gln, l-Ile, l-Val), 2 mM IAA, and 0.05 μCi [14C]IAA (50 mCi mmol−1) for determination of the activity toward amino acids. The enzyme activity was measured as described above.
Steady-state kinetic analysis was performed using the purified IAA-Asp synthetase preparation and standard radioactive assay conditions but with one substrate concentration varied and two substrate concentrations fixed as follows: varied IAA, IPA, l-Trp, (±)JA (0.1–2 mM) or 1-NAA, 2,4-D, PAA (0.1–10 mM), fixed Asp (10 mM), and ATP (5 mM); varied amino acids (0.5–10 mM), fixed IAA (2 mM), and ATP (5 mM); varied ATP (0.1–8 mM), fixed IAA (2 mM), and Asp (10 mM). The enzyme activity was assayed as described above. The V max and K m values were calculated using a Lineweaver-Burk plot (double-reciprocal plot) and the Michaelis-Menten equation, respectively. The k cat values were calculated by dividing the V max values by the concentration of IAA-Asp synthetase present in the reaction mixture.
Determination of the Molecular Mass by Gel Filtration
The molecular mass of the native IAA-Asp synthetase was determined after a DEAE-Sephacel anion-exchange chromatography step by gel filtration using a Sephadex G-150 column (1.9 × 91 cm). Alcohol dehydrogenase (150 kDa), bovine serum albumin (67 kDa), ovalbumin (45 kDa), carbonic anhydrase (29 kDa), and cytochrome c (12.4 kDa) were used as molecular mass standards. Elution was done with 25 mM Tris-HCl, pH 7.6, containing 50 mM NaCl and 5 mM 2-mercaptoethanol at the flow rate of 0.3 ml min−1.
Protein A Immunoprecipitation
A total of 0.1 ml of homogenate, enzyme preparations from each purification step, and the purified enzyme were incubated with 5 μl of rabbit anti-recombinant GH3.5 polyclonal antibodies for 2 h at 0–2°C. Antigen-antibody complexes were immunoprecipitated by mixing them with 10 μl of a suspension of Protein A immobilized on 250-μm acrylic beads (0.16 mg/ml, Sigma-Aldrich, Germany). After washing, proteins were eluted by incubation with 25 mM glycine–HCl, pH 2.5, for 5 min and analyzed by 12% (w/v) SDS-PAGE followed by Western blot (Bollag and Edelstein 1991).
SDS-PAGE and Western Blot
SDS-PAGE was performed according to the method of Ogita and Markert (1979) in a Mini Protean II electrophoresis apparatus (Bio-Rad) using 12% (w/v) running gel and 6% (w/v) stacking gel. The proteins used as molecular mass standards were a 10-kDa Protein Ladder (Gibco, UK). SDS denatured proteins were stained with silver nitrate according to the modified method of Harlow and Lane (1988). The proteins separated by SDS-PAGE were transferred onto a nitrocellulose membrane (Whatman GmbH, Germany) in the transfer buffer containing 50 mM Tris, 380 mM glycine, 0.1% (w/v) SDS, and 20% (v/v) methanol. The position of the protein markers was visualized by staining with 0.2% (w/v) Ponceau S in 3% (w/v) sulfosalicylic acid and 3% (w/v) trichloroacetic acid. The blot was incubated with rabbit antiserum containing polyclonal anti-Arabidopsis thaliana GH3.5 recombinant protein antibodies (Zhang and others 2007). IAA-Asp synthetase protein was detected using goat anti-rabbit IgG antibodies conjugated to alkaline phosphatase (Harlow and Lane 1988).
Protein Determination
After ion-exchange chromatography and gel filtration, protein was measured spectrophotometrically at 280 nm. Protein concentration was determined by the Bradford method (1976) using bovine serum albumin as a standard.
Results
Activity of the IAA-Asp Synthetase During Maturation of Pea Seeds
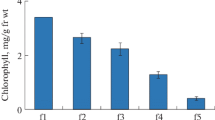
A study was performed to investigate IAA-Asp synthetase activity in developing seeds of pea. The stages of seed development were determined based on seed diameter. Figure 1 shows changes in enzyme activity during seed maturation. IAA-Asp synthetase activity was approximately 6 nmol h−1 g−1 FW in the youngest seeds (with a diameter of 3 mm) and then increased to 12 nmol h−1 g−1 FW (seeds with a diameter of 5 mm). Synthesis of IAA-aspartate clearly declined during later development to approximately 3 nmol h−1 g−1 FW in seeds with a diameter of 8 mm. Fully mature (dry) seeds revealed only a trace of IAA-Asp synthetase activity. Because the highest IAA-Asp synthetases activity was observed with seeds 5 mm in diameter, seeds at this stage were used for future enzyme isolation and purification.
Temporal changes in IAA-Asp synthetase activity during pea seed maturation. Assay conditions are described in Materials and Methods. Error bars indicate SD of the mean for three repetitions of each experiment
Purification of the IAA-Asp Synthetase
Because our previous study (Ostrowski and Jakubowska 2008) revealed that the enzyme precipitation with ammonium sulfate caused total loss of the activity, poly(ethylene glycol) PEG 6000 was used (step I). Initial fractionation of the crude homogenate with 20% (w/v) PEG 6000 enabled the removal of 65.4% of contaminating proteins with a 1.4-fold increase in total activity. DEAE-Sephacel anion-exchange chromatography provided an additional efficient fractionation step (step II). IAA-Asp synthetase activity was eluted with 250 mM NaCl in Tris-HCl, pH 7.6, as a single peak (data not shown). A 40.5-fold increase in the specific activity with a 24.3% loss of the total activity was noted after this step. The fraction at R f = 0.45 obtained after preparative PAGE was apparently homogeneous, yielding a single band as assessed by silver nitrate staining after SDS-PAGE (Fig. 2, lane 4). The results of each purification step are summarized in Table 1. The specific activity of the purified IAA-Asp synthetase preparation was 180 nmol min−1 mg−1 protein (approximately 1500-fold purification with a yield of 2.05% from the crude enzyme).
SDS-PAGE and Western blots of the crude (a) and purified (b) IAA-Asp synthetase. Lane 1, homogenate after SDS-PAGE; lane 2, immunoblot of homogenate; lane 3, immunoblot of the purified IAA-Asp synthetase; lane 4, the purified IAA-Asp synthetase after SDS-PAGE; lane 5, molecular mass standards. SDS-PAGE and Western blots were performed as described in Materials and Methods. Proteins after SDS-PAGE were visualized by the silver nitrate staining procedure or transferred onto nitrocellulose membrane. The polypeptide cross-reacting with anti-AtGH3.5 polyclonal antibodies was localized by the alkaline phosphatase-mediated immunostaining procedure
Characterization of the IAA-Asp Synthetase
The molecular mass of the native IAA-Asp synthetase estimated by gel filtration on a Sephadex G-150 column was 70.8 kDa, according to the elution volume of the purified enzyme and the linear equation (data not shown). When subjected to SDS-PAGE, the purified enzyme yielded a single silver-stained band with a molecular mass of 70 kDa (Fig. 2, lane 4). These similar results suggest that the IAA-Asp synthetase from pea seeds is a monomeric protein. Western blot analysis after Protein A immunoprecipitation demonstrated that rabbit anti-recombinant GH3.5 antiserum detected a single immunoreactive band at 70 kDa in pea seed crude extract and in the final enzyme preparation (Fig. 2, lane 3).
The isoelectric point of the enzyme was determined by electrofocusing to be 4.7 (data not shown). This was consistent with its behavior in DEAE-Sephacel anion-exchange chromatography because it bonded to DEAE-Sephacel and was eluted with 250 mM NaCl at pH 7.6. Analysis of the pH dependence of IAA-aspartate synthesis indicated that the enzyme was active within a range of pH 8.0–9.5, reaching optimum catalytic activity at pH 8.2 (data not shown). As shown in Fig. 3a, the enzyme-preferred cofactors were Mg2+ and Ca2+ (138 and 130% of control, respectively). Whereas Mn2+ supported 108% of the enzyme activity, lower rates of IAA-Asp synthesis were observed in the presence of Cu2+ (80%) or Zn2+ (70%) under test conditions. The effect of various reducing agents on IAA-Asp synthetase activity is shown in Fig. 3b. It was found that all tested thiol reagents activated the enzyme, but 2-mercaptoethanol (ME) and dithiotreitol (DTT) were the best among them. The enzyme activity was upregulated by ME and DTT about sixfold and fivefold, respectively. Surprisingly, oxidized glutathione (GSSG) was more effective than reduced glutathione (GSH).
Biochemical characterization of the pea IAA-Asp synthetase. a Effect of divalent cations on the IAA-Asp synthetase activity. Enzyme activity in the presence of cations (5 mM each) is expressed relative to the control set at 100. Assay conditions are described in Materials and Methods. Error bars indicate SD of the mean for three repetitions of each experiment. b Effect of reducing agents on the IAA-Asp synthetase activity. Enzyme activity in the presence of thiol reagents (5 mM each) is expressed relative to the control set at 100. Assay conditions are described in Materials and Methods. Error bars indicate SD of the mean for three repetitions of each experiment
Substrate Specificity and Kinetic Properties
To examine the substrate specificity and preference of the enzyme, a range of auxins (natural and synthetic), SA, and JA as well as amino acids were tested, with the results summarized in Table 2. The highest activity of the pea enzyme was observed on IAA (164.1 ± 5 nmol min−1 mg−1 protein), whereas moderate (24.1% of the relative activity) and low (6.4% of the relative activity) activity was observed on the other natural auxins IBA and IPA, respectively. The indole-containing amino acid tryptophan was a rather poor acyl substrate for conjugation as well (19.3% of the relative activity). Interestingly, the enzyme exhibited a significant activity toward two synthetic auxins: 1-NAA (about 55% of the relative activity) and 2,4-D (28% of the relative activity). By contrast, very little activity with PAA (3.6% of the relative activity) and no activity with the nonactive form of auxin, 2-NAA, was detected. Taking into account that AtGH3.5 also has adenylating activity on SA and AtGH3.11 (JAR1) conjugates JA to amino acids (Staswick and others 2002), the pea enzyme activity on these phytohormones was tested. Although SA supported more than 30% of the enzyme activity, compared to that with IAA, JA was almost ineffective as a substrate for conjugation to Asp (5.2% of the relative activity). A total of 14 different amino acids were examined in the IAA conjugation reaction catalyzed by the pea enzyme. We found that the enzyme was most active with Asp, but it also conjugated Ala and Lys (about 50% and more than 40% of the relative activity, respectively). Surprisingly, although Glu is structurally related to Asp, the enzyme displayed just under 10% of the catalytic activity compared to that with Asp. Finally, no activity was found when the rest of the amino acids were used.
To characterize further the substrate preferences of the pea IAA-Asp synthetase, its kinetic properties were determined using the indicated auxins and amino acids, with the results summarized in Table 2. The kinetic parameters for ATP (with IAA and Asp concentrations fixed) were also estimated to ensure that sufficient ATP was used in kinetic assays. We found that the purified enzyme follows the Michaelis-Menten model, with the K m = 0.5 mM for ATP and saturating concentration of 3 mM. Similarly, the enzyme exhibited standard Michaelis-Menten kinetics with all tested acyl substrates as well as amino acids. The V max and K m values for the acyl substrates IAA, IBA, 1-NAA, 2,4-D, and SA with saturating concentrations of Asp and ATP were determined. IAA-Asp synthetase had the highest affinity for IAA (K m = 0.23 mM), followed by IBA (K m = 1.5 mM), 1-NAA (K m = 3.6 mM), SA (K m = 3.8 mM), and 2,4-D (K m = 5.7 mM). As a result, the highest catalytic efficiency (k cat/K m value) was for IAA (682,608.7 M−1 min−1). Catalytic efficiencies for IBA, 1-NAA, 2,4-D, and SA were about 27-, 29-, or even 50-fold lower than that for IAA. Then, the V max and K m values of the pea enzyme for amino acid substrates Asp, Glu, Lys, and Ala with saturating concentrations of IAA and ATP were determined. The K m value was lowest for Asp (2.5 mM), whereas for Glu, Lys, and Ala it was 4.2, 4.5, and 6.0 mM, respectively. According to amino acid substrate affinity, the enzyme exhibited the highest catalytic efficiency for Asp (5080 s−1 M−1) with reduced catalytic efficiencies for other amino acids: 8.5 times (Glu), 11.4 times (Lys), and 9.6 times (Ala) lower compared with that for Asp.
Discussion
Most of the IAA present in pea seeds is found as IAA-amide conjugates, and IAA-aspartate is probably a major amide conjugate (Bandurski and Schulze 1977). We previously found that enzymatic activity catalyzes biosynthesis of several IAA-amino acid conjugates in immature pea seeds (Ostrowski and Jakubowska 2008). Because IAA-Asp was dominant among products synthesized by pea crude extract, we decided to isolate and characterize this activity, especially because there has been no report on IAA-amidosynthetase isolated from plant material. Analysis of the enzyme activity, when calculated per gram of fresh weight (FW), showed that IAA-Asp synthetase activity, which was relatively high at early maturation steps, declined during seed maturation. Dry seeds revealed only a trace of the enzyme activity. Comparison of these results with the data available in the literature is limited because only the changes of conjugated IAA levels and not the enzyme activity in developing seeds were reported (Sandberg and others 1987; Bialek and Cohen 1989). However, it is obvious that the conjugated IAA levels are related to enzyme activity which synthesizes these metabolites. Thus, based on our data on IAA-Asp synthetase activity and previous reports on the changes in the IAA-amide conjugate levels (Bialek and Cohen 1989, 1992), it seems that pea seeds reveal a similar developmental pattern of these compounds. Seeds with a diameter of 5 mm were used in isolation and purification of IAA-Asp synthetase because of the highest enzyme activity presented at this stage.
We developed a simple three-step purification method that yielded a homogeneous enzyme preparation. The enzyme could not be precipitated by ammonium sulfate because its activity was completely lost and so PEG 6000 was used. The highest activity of IAA-Asp synthetase remained in the supernatant after homogenate fractionation with 20% (w/v) concentration of PEG 6000. The failure of this enzyme to be precipitated by PEG can suggest a certain physical similarity to the enzyme catalyzing synthesis of the ester conjugate, 1-O-(indol-3-ylacetyl)-β-d-glucose (1-O-IAA-Glu) in maize kernels (Kowalczyk and Bandurski 1991) and legume seeds (Jakubowska and Kowalczyk 2004). Elution of the IAA-Asp synthetase activity at pH 7.6 required 250 mM NaCl concentration during DEAE-Sephacel anion-exchange chromatography, which points out a relatively low isoelectric point. Indeed, the isoelectric point of IAA-Asp synthetase was found to be 4.7. This value is lower than the values in the range 5.61–5.99 calculated for the three auxin-inducible GH3-like proteins correlating with adventitious root development in A. thaliana (Sorin and others 2006) and the theoretical value of 5.37 calculated for the predicted protein Pp-GH3.16 from Pinus pinaster (Reddy and others 2006). Preparative PAGE was found to be a very effective purification step. It allowed the practical removal of all remaining protein contamination and resulted in a 32-fold increase in specific activity. The IAA-Asp synthetase was finally purified about 1500-fold to a specific activity of 180 nmol min−1 mg−1. The rate of IAA-Asp synthesis by this enzyme is comparable to that found for recombinant AtGH3.6 protein (Staswick and others 2005). The IAA-Asp synthetase molecular mass of 70 kDa is within the range of those of plant GH3 proteins reported so far. For example, the masses of the two proteins from the A. thaliana adventitious roots proteome, AtGH3.5 and AtGH3.6, are 70.01 and 70.41 kDa, respectively (Sorin and others 2006). Similar molecular masses were also estimated for the two proteins from Physcomitrella patens, PpGH3-1 (about 71 kDa) and PpGH3-2 (about 67 kDa) (Bierfreund and others 2004), for the CcGH3 from Capsicum chinese (67.4 kDa) (Liu and others 2005), for a putative GH3 protein from Pinus pinaster roots, designated as Pp-GH3.16 (69.9 kDa) (Reddy and others 2006), and for the OsGH3.8 from Oryza sativa (66.9 kDa) (Chen and others 2009). By contrast, the only known bacterial amidosynthetase conjugating IAA to lysine in Pseudomonas savastanoi is characterized by a substantially lower molecular mass (44 kDa) (Roberto and others 1990).
Like the enzymes from A. thaliana, the pea IAA-Asp synthetase activity was the highest at basic conditions, with the maximal rate of IAA-Asp formation at pH 8.2. This result may suggest how an enzyme interacts with its substrates as well as the potential cytoplasmic localization of the enzyme. At present, we have no information on the cellular localization of our enzyme, but previous studies have shown that Arabidopsis JAR1/FIN219 and moss PpGH3-1 are cytoplasmic proteins (Hsieh and others 2000; Ludwig-Müller and others 2009). Calculations based on the predicted open reading frame (ORF) for Capsicum chinese GH3 indicated that the CcGH3 also exists as a soluble cytoplasmic protein (Liu and others 2005). Enzymatic synthesis of IAA-Asp in pea seeds requires ATP and Mg2+, like the known GH3 protein forming acyl-adenylate intermediate (Staswick and others 2002, 2005; Chen and others 2009; Okrent and others 2009). The pea enzyme was strongly influenced by the reducing agents, especially by ME and DTT. We believe that the reducing agents protect –SH groups, which in a free state stabilize the enzyme structure. A similar effect of the thiol reagents was observed for maize synthase of 1-O-IAA-glucose (Leznicki and Bandurski 1988). However, another enzyme involved in the biosynthesis of IAA-ester conjugates in maize endosperm, IAA-inositol synthase, was found to be strongly inhibited by DTT and ME (Kesy and Bandurski 1990). The inhibition of enzyme activity by DTT was also reported for both plant (Ludwig-Müller and others 1996; Campanella and others 2008) and bacterial IAA-amidohydrolases (Chou and others 1996; Chou and Huang 2005). Analysis of amino acid sequences of putative amidosynthetases has shown that these enzymes, like the amidohydrolases, contain conserved cysteine residues (Roux and Perrot-Rechenmann 1997; Campanella and others 2008; Bitto and others 2009). These findings suggest that the redox status of the -SH groups is important in IAA conjugate synthesis and hydrolysis. One can suppose that a different influence on IAA-Asp synthetase and hydrolase activities by sulfhydryl compounds is connected with the protein structure of the enzymes. It would be interesting to compare the enzyme structures; however, only the X-ray structure of ILL2, an auxin-conjugate amidohydrolase from A. thaliana, is currently known (Bitto and others 2009). The catalytic domain of this protein contains Cys 137, which is one of the five amino acid residues likely involved in metal coordination.
Analysis of the substrate specificity revealed that the studied enzyme was active on several natural and synthetic auxins, similar to the six A. thaliana GH3 recombinant proteins (Staswick and others 2005). The enzyme was most active with IAA, whereas activities on other natural auxins, IBA and IPA, which were effective as acyl substrates for some Arabidopsis GH3 enzymes, were reduced relative to IAA by over 75 and 85%, respectively. However, the pea enzyme was able to conjugate the synthetic auxin 1-NAA (54.9% of the relative activity), like Arabidopsis IAA-conjugating enzymes. Although these enzymes had essentially no activity on 2,4-D, IAA-Asp synthetase from pea could conjugate 2,4-D, but the reaction rate was considerably lower compared with that of IAA (26.9% of the relative activity). However, the pea enzyme had very little activity with PAA (3.6% of the relative activity), unlike Arabidopsis IAA-amidosynthetases, for which adenylating activity was the same or even higher (GH3.3, GH3.5, GH3.6) than that with IAA. The pea enzyme was also active on SA (33.3% of the relative activity), like the AtGH3.5 protein (Staswick and others 2002). However, JA was quite an ineffective substrate for this synthetase, like the six Arabidopsis enzymes (Staswick and others 2005). More detailed kinetic analysis has shown that the IAA-Asp synthetase from immature seeds of pea clearly preferred IAA as acyl substrate, because the enzyme had the highest affinity for IAA (K m = 0.23 mM). The K m values for IBA, 1-NAA, and 2,4-D were lower by about 7-, 15-, and 25-fold, respectively. This resulted in the highest catalytic efficiency for IAA compared with those for other auxins. Comparison of these results with the data available in the literature is limited because kinetic parameters have been determined for only two recombinant proteins: GH3.8 from rice (Chen and others 2009) and JAR1 (GH3.11) which conjugates only JA in Arabidopsis and tomato (Suza and others 2010). The K m of 0.23 mM for the pea enzyme is about twofold higher than the K m of 0.123 mM for the rice enzyme (Chen and others 2009) and about fourfold to eightfold higher than the K m values for JAR1 (+)-7-iso-jasmonic acid substrate from Arabidopsis and tomato (0.06 and 0.03 mM, respectively) (Suza and others 2010). However, the K m of the pea enzyme is the same as the K m of 0.24 mM for a recombinant A. thaliana IAA glucosyltransferase (Jackson and others 2001) and lower than the K m of 0.8 mM for the maize enzyme (Kowalczyk and others 2002), which are involved in IAA-ester conjugate biosynthesis.
Of the 14 amino acids tested, only four were conjugated to IAA by the pea enzyme, unlike studies with Arabidopsis IAA-amidosynthetases that revealed a broad specificity for amino acids (Staswick and others 2005). Similar to some Arabidopsis GH3 proteins, the examined enzyme had the highest activity on aspartate and lower activity with alanine and lysine (49.4 and 43.4% of the relative activity, respectively). Surprisingly, only trace activity with glutamate and no activity with methionine, which were detectable in the crude extract by TLC in our previous study (Ostrowski and Jakubowska 2008), were determined with the purified enzyme. This may indicate that IAA-Asp synthetase is abundant in pea seeds and may confirm our earlier predictions that the activity in crude extract represents a mixture of different enzyme activities. In general, the K m values for amino acids used as the substrates for conjugation to IAA were relatively high (2.5–6.5 mM), but the data from kinetic analysis suggest that the pea enzyme prefers Asp is this reaction. The k cat for the enzyme with Asp (12.7 s−1) was four-, five-, and over sixfold higher than the k cat for Ala, Glu, and Lys, respectively. This resulted in the highest catalytic efficiency for Asp (5080 s−1 M−1) compared with those for other amino acids, which were lower by around one order of magnitude. Differences in the amino acid preferences of the pea enzyme could contribute to the accumulation of IAA-Asp, which is a major IAA-amide conjugate in pea seeds. The K m values of both Arabidopsis and tomato JAR1 for isoleucine were substantially lower than that for the pea enzyme: over 80-fold and 125-fold, respectively (0.03 and 0.02 mM) (Suza and Staswick 2008; Suza and others 2010). By contrast, the K m of the pea enzyme for Asp (2.5 mM) was similar to that for OsGH3.8 protein (1.58 mM) (Chen and others 2009). However, in the case of the rice enzyme, it is not possible to discuss its preferences to amino acid substrates because only aspartate was tested.
In conclusion, we have isolated and characterized the enzyme involved in IAA amino acid biosynthesis from plant material for the first time. Similar to characterized IAA-amidosynthetases, the IAA-Asp synthetase from immature pea seeds required basic pH, Mg2+ and ATP for IAA conjugation. Based on detailed kinetic analysis, the pea native enzyme seems to be a more specific enzyme compared to the bacterially expressed GH3 proteins characterized by a rather broad substrate specificity (Staswick and others 2005; Ludwig-Müller and others 2009). Although the purified IAA-Asp synthetase was able to use several acyl and amino acid substrates, the enzyme displayed a clear preference for synthesizing IAA-Asp. As Western blot analysis demonstrated that a 70-kDa protein band cross-reacts with anti-AtGH3.5 antibodies, one can suppose that the pea enzyme is a candidate for a new member of the GH3-like protein family. We are continuing our study to obtain partial amino acid sequences and compare them to those of previously obtained GH3 proteins. It could provide evidence that this enzyme is really a GH3-like protein. We also hope to clone the gene encoding the enzyme protein and to further investigate the IAA-Asp functions in plants.
References
Bandurski RS, Schulze A (1977) Concentration of indole-3-acetic acid and its derivatives in plants. Plant Physiol 60:211–213
Bandurski RS, Cohen JD, Slovin JP, Reinecke DM (1995) Auxin biosynthesis and metabolism. In: Davies PJ (ed) Plant hormones: physiology, biochemistry and molecular biology. Kluwer, Academic Publishers, Dordrecht, pp 39–65
Bartel B, Fink GR (1995) ILR1, an amidohydrolase that releases active indole-3-acetic acid from conjugates. Science 268:1745–1748
Bialek K, Cohen JD (1989) Free and conjugated indole-3-acetic acid in developing bean seeds. Plant Physiol 91:775–779
Bialek K, Cohen JD (1992) Amide-linked indoleacetic acid conjugates may control levels of indoleacetic acid in germinating seedlings of Phaseolus vulgaris. Plant Physiol 100:2002–2007
Bierfreund NM, Tintelnot S, Reski R, Decker EL (2004) Loss of GH3 function does not affect phytochrome-mediated development in a moss, Physcomitrella patens. J Plant Physiol 161:823–835
Bitto E, Bingman CA, Bittova L, Houston NL, Boston RS, Fox BG, Phillips GN Jr (2009) X-ray structure of ILL2, an auxin-conjugate amidohydrolase from Arabidopsis thaliana. Proteins 74:61–71
Blakesly RW, Boezi JA (1977) A new staining technique for proteins in polyacrylamide gels using Coomassie Brilliant Blue G250. Anal Biochem 82:580–582
Bollag DM, Edelstein SJ (1991) Protein methods. John Willey & Sons, New York, pp 181–191
Bradford MM (1976) A rapid and sensitive method for quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Campanella JJ, Bakllamaja V, Restieri T, Vomacka M, Herron J, Patterson M, Shahtaheri S (2003a) Isolation of an ILR1 auxin conjugate hydrolase homolog from Arabidopsis suecica. Plant Growth Regul 39:175–181
Campanella JJ, Ludwig-Müller J, Bakllamaja V, Sharma V, Cartier A (2003b) ILR1 and sILR1 IAA amidohydrolase homologs differ in expression pattern and substrate specificity. Plant Growth Regul 41:215–223
Campanella JJ, Olajide AF, Magnus V, Ludwig-Müller J (2004) A novel auxin conjugate hydrolase from wheat with substrate specificity for longer side-chain auxin amide conjugates. Plant Physiol 135:2230–2240
Campanella JJ, Smith SM, Leibu D, Wexler S, Ludwig-Müller J (2008) The auxin conjugate hydrolase family of Medicago truncatula and their expression during the interaction with two symbionts. J Plant Growth Regul 27:26–38
Chen Q, Zhang B, Hicks LM, Wang S, Jez JM (2009) A liquid chromatography-tandem mass spectrometry-based assay for indole-3-acetic acid-amido synthetase. Anal Biochem 390:149–154
Chou JC, Huang YB (2005) Induction and characterization of an indole-3-acetyl-l-alanine hydrolase from Arthrobacter ilicis. J Plant Growth Regul 24:11–18
Chou JC, Kuleck GA, Cohen JD, Mulbry WW (1996) Partial purification and characterization of an inducible indole-3-acetyl-l-aspartic acid hydrolase from Enterobacter agglomerans. Plant Physiol 112:1281–1287
Chou JC, Mulbry WW, Cohen JD (1998) The gene for indole-3-acetyl-l-aspartic acid hydrolase from Enterobacter agglomerans: molecular cloning, nucleotide sequence, and expression in Escherichia coli. Mol Gen Genet 259:172–178
Cohen JD, Bandurski RS (1982) Chemistry and physiology of the bound auxins. Ann Rev Plant Physiol 33:403–430
Davies RT, Goetz DH, Lasswell J, Anderson MN, Bartel B (1999) IAR3 encodes an auxin conjugate hydrolase from Arabidopsis. Plant Cell 11:365–376
Ehmann A (1977) The Van Urk-Salkowski reagent—a sensitive and specific chromogenic reagent for silica gel thin-layer chromatographic detection and identification of indole derivatives. J Chromatogr 132:267–276
Hagen G, Guilfoyle TJ (1985) Rapid induction of selective transcription by auxins. Mol Cell Biol 5:1197–1203
Harlow E, Lane D (1988) Antibodies. A laboratory manual. Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Hsieh HL, Okamoto H, Wang M, Ang LH, Matsui M, Goodman H, Deng XW (2000) FIN219, an auxin-regulated gene, defines a link between phytochrome A and the downstream regulator COP1 in light control of Arabidopsis development. Gen Dev 14:1958–1970
Jackson RG, Lim EK, Li Y, Kowalczyk M, Sandberg G, Hoggett J, Ashford DA, Bowles DJ (2001) Identification and biochemical characterization of an Arabidopsis indole-3-acetic acid glucosyltransferase. J Biol Chem 276:4350–4356
Jakubowska A, Kowalczyk S (2004) The auxin conjugate 1-O-indole-3-acetyl-β-d-glucose is synthesized in immature legume seeds by IAGlc synthase and may be used for modification of some high molecular weight compounds. J Exp Bot 55:791–801
Jakubowska A, Kowalczyk S (2005) A specific enzyme hydrolyzing 6-O(4-O)-indole-3-ylacetyl-β-d-glucose in immature kernels of Zea mays. J Plant Physiol 162:207–213
Jakubowska A, Kowalczyk S, Leźnicki AJ (1993) Enzymatic hydrolysis of 4-O and 6-O-indol-3-ylacetyl-β-d-glucose in plant tissues. J Plant Physiol 142:61–66
Kesy JM, Bandurski RS (1990) Partial purification and characterization of indole-3-ylacetylglucose: myo-inositol indol-3-ylacetyltransferase (indoleacetic acid-inositol synthase). Plant Physiol 94:1598–1604
Kowalczyk S, Bandurski RS (1991) Enzymic synthesis of 1-O-(indol-3-ylacetyl)-β-d-glucose. Biochem J 279:509–514
Kowalczyk S, Jakubowska A, Bandurski RS (2002) 1-Naphthalene acetic acid induces indole-3-ylacetylglucose synthase in Zea mays seedling tissue. Plant Growth Regul 38:127–134
Kowalczyk S, Jakubowska A, Zielińska E, Bandurski RS (2003) Bifunctional indole-3-acetyl transferase catalyses synthesis and hydrolysis of indole-3-acetyl-myo-inositol in immature endosperm of Zea mays. Physiol Plant 119:165–174
LeClere S, Tellez R, Rampey RA, Matsuda SPT, Bartel B (2002) Characterization of a family of IAA-amino acid conjugate hydrolases from Arabidopsis. J Biol Chem 277:20446–20452
Leznicki AJ, Bandurski RS (1988) Enzymic synthesis of indole-3-acetyl-1-O-β-d-glucose. Plant Physiol 88:1481–1485
Liu K, Kang BC, Jiang H, Moore SL, Li H, Watkins CB, Setter TL, Jahn MM (2005) A GH3-like gene, CcGH3, isolated from Capsicum chinese L. fruit is regulated by auxin and ethylene. Plant Mol Biol 58:447–464
Ludwig-Müller J, Epstein E, Hilgenberg W (1996) Auxin-conjugate hydrolysis in Chinese cabbage: characterization of an amidohydrolase and its role during infection with clubroot disease. Physiol Plant 97:627–634
Ludwig-Müller J, Jülke S, Bierfreund NM, Decker EL, Reski R (2009) Moss (Physcomitrella patens) GH3 proteins act in auxin homeostasis. New Phytol 181:323–338
Ogita ZI, Markert CL (1979) A miniaturized system for electrophoresis on polyacrylamide gels. Anal Biochem 99:233–241
Okrent RA, Brooks MD, Wildermuth MC (2009) Arabidopsis GH3.12 (PBS3) conjugates amino acids to 4-substituted benzoates and is inhibited by salicylate. J Biol Chem 284:9742–9754
Ostrowski M, Jakubowska A (2008) Identification of enzyme activity that conjugates indole-3-acetic acid to aspartate in immature seeds of pea (Pisum sativum). J Plant Physiol 165:564–569
Rampey RA, LeClere S, Kowalczyk M, Ljung K, Sanberg G, Bartel B (2004) A family of auxin-conjugate hydrolases that contributes to free indole-3-acetic acid levels during Arabidopsis germination. Plant Physiol 135:978–988
Reddy SM, Hitchin S, Melayah D, Pandey AK, Raffier C, Henderson J, Marmeisse R, Gay G (2006) The auxin-inducible GH3 homologue Pp-GH3.16 is downregulated in Pinus pinaster root system on ectomycorrhizal symbiosis establishment. New Phytol 170:391–400
Roberto FF, Klee H, White F, Nordeen R, Kosuge T (1990) Expression and fine structure of the gene encoding N ε-(indole-3-acetyl)-l-lysine synthetase from Pseudomonas savastanoi. Proc Natl Acad Sci USA 87:5797–5801
Roux C, Perrot-Rechenmann C (1997) Isolation by differential display and characterization of a tobacco auxin-responsive cDNA Nt-gh3 related to GH3. FEBS Lett 419:131–136
Sandberg G, Ernstsen A, Hamnede M (1987) Dynamics of indole-3-acetic acid and indole-3-ethanol during development and germination of Pinus sylvestris seeds. Physiol Plant 71:411–418
Savić B, Tomić S, Magnus V, Gruden K, Barla K, Grenković R, Ludwig-Müller J, Salopek-Sondi B (2009) Auxin amidohydrolases from Brassica rapa cleave the alanine conjugate of indolepropionic acid as a preferable substrate: a biochemical and modeling approach. Plant Cell Physiol 50:1587–1599
Schuller A, Ludwig-Müller J (2006) A family of auxin conjugate hydrolases from Brasica rapa: characterization and expression during clubroot disease. New Phytol 171:145–158
Sorin C, Negroni L, Balliau T, Corti H, Jacquemot MP, Davanture M, Sandberg G, Zivy M, Bellini C (2006) Proteomic analysis of different mutant genotypes of Arabidopsis led to the identification of 11 proteins correlating with adventitious root development. Plant Physiol 140:349–364
Staswick PE (2009) The tryptophan conjugates of jasmonic and indole-3-acetic acids are endogenous auxin inhibitors. Plant Physiol 150:1310–1321
Staswick PE, Tiryaki I, Rowe ML (2002) Jasmonate response locus JAR1 and several related Arabidopsis genes encode enzymes of the firefly luciferase superfamily that show activity on jasmonic, salicylic, and indole-3-acetic acid in an assay for adenylation. Plant Cell 14:1405–1415
Staswick PE, Serban B, Rowe M, Tiryaki I, Maldonado MT, Maldonado MC, Suza W (2005) Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17:616–627
Suza WP, Staswick PE (2008) The role of JAR1 in jasmonoyl-l-isoleucine production during Arabidopsis wound response. Planta 227:1221–1232
Suza WP, Rowe ML, Hamberg M, Staswick PE (2010) A tomato enzyme synthesizes (+)-7-iso-jasmonoyl-l-isoleucine in wounded leaves. Planta 321:717–728
Szerszen JB, Szczyglowski K, Bandurski RS (1994) iaglu, a gene from Zea mays involved in conjugation of growth hormone indole-3-acetic acid. Science 265:1699–1701
Sztein AE, Cohen JD, Slovin JP, Cooke TJ (1995) Auxin metabolism in representative land plants. Am J Bot 82:1514–1521
Terol J, Domingo C, Talón M (2006) The GH3 family in plants: genome wide analysis in rice and evolutionary history based on EST analysis. Gene 371:279–290
Wang H, Tian C, Duan J, Wu K (2008) Research progress on GH3s, one family of primary auxin-responsive genes. Plant Growth Regul 56:225–232
Woodward AW, Bartel B (2005) Auxin: regulation, action, and interaction. Ann Bot 95:707–735
Zelená E (2000) The effect of light on metabolism of IAA in maize seedlings. Plant Growth Regul 30:23–29
Zhang Z, Li Q, Li Z, Staswick PE, Wang M, Zhu Y, He Z (2007) Dual regulation role of GH3.5 in salicylic acid and auxin signaling during Arabidopsis—Pseudomonas syringae interaction. Plant Physiol 145:450–464
Zhang SW, ChH Li, Cao J, Zhang YC, Zhang SQ, Xia YF, Sun DY, Sun Y (2009) Altered architecture and enhanced drought tolerance in rice via the down-regulation of indole-3-acetic acid by TLD1/OsGH3.13 activation. Plant Physiol 151:1889–1901
Acknowledgments
We are very grateful to Professor Zuhua He, National Key Laboratory of Plant Molecular Genetics, Institute of Plant Physiology and Ecology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, for the generous gift of rabbit antiserum against AtGH3.5 protein. This work was supported by the EFS-ZPORR grant No. 69-UMK-09.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Ostrowski, M., Jakubowska, A. Purification and Biochemical Characterization of Indole-3-acetyl-aspartic Acid Synthetase from Immature Seeds of Pea (Pisum sativum). J Plant Growth Regul 30, 30–40 (2011). https://doi.org/10.1007/s00344-010-9164-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-010-9164-x