Abstract
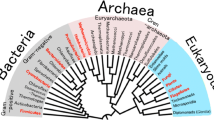
Antarctica is a region with harsh conditions which have become a focus of research for identifying novel extreme microorganisms in recent years. Here, we identified and characterized the KGI_MA19 isolate obtained by enriched culture and freeze–thaw stress treatment from the sediment samples of King George Island in Antarctica. Whole-genome sequencing, MLSA, and phylogenetic analyses revealed that this isolate was a novel strain of Pseudomonas mandelii sharing less similarity to the first Pseudomonas mandelii isolate 6A1 from Antarctica. The phenotypic and biochemical characterizations indicate that this psychrotolerant and halotolerant strain can use disaccharides as a carbon source, has denitrification ability, and has multiple antibiotic resistance. Moreover, comparative proteomic analyses showed that P. mandelii KGI_MA19 adapts to the cold environment by changing its membrane composition, excreting EPS, and producing major cold shock proteins, in addition to turning off its other stress response mechanisms.




Similar content being viewed by others
References
Abraham WP, Raghunandanan S, Gopinath V, Suryaletha K, Thomas S (2020) Deciphering the cold adaptive mechanisms in Pseudomonas psychrophila MTCC12324 isolated from the Arctic at 79 N. Curr Microbiol 77(9):2345–2355. https://doi.org/10.1007/s00284-020-02006-2
Alanjary M, Steinke K, Ziemert N (2019) AutoMLST: an automated web server for generating multi-locus species trees highlighting natural product potential. Nucleic Acids Res 47(W1):W276–W282. https://doi.org/10.1093/nar/gkz282
Arcarons N, Vendrell-Flotats M, Yeste M, Mercade E, Lopez-Bejar M, Mogas T (2019) Cryoprotectant role of exopolysaccharide of Pseudomonas sp. ID1 in the vitrification of IVM cow oocytes. Reprod Fertil Dev 31(9):1507–1519. https://doi.org/10.1071/RD18447
Ashburner A, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, MAtese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. Nat Genet 25(1):25–29. https://doi.org/10.1038/75556
Ashburner A, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, MAtese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2021) The gene ontology resource: enriching a GOld mine. Nucleic Acids Res 49(D1):D325–D334. https://doi.org/10.1093/nar/gkaa1113
Auch AF, von Jan M, Klenk HP, Göker M (2010) Digital DNA-DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genom Sci 2(1):117–134. https://doi.org/10.4056/sigs.531120
Biemer JJ (1973) Antimicrobial susceptibility testing by the Kirby-Bauer disc diffusion method. Ann Clin Lab Sci 3(2):135–140
Bozal N, Montes MJ, Mercadé E (2007) Pseudomonas guineae sp. nov., a novel psychrotolerant bacterium from an Antarctic environment. Int J Syst Evol Microbiol 57(Pt11):2609–2612. https://doi.org/10.1099/ijs.0.65141-0
Carrión O, Miñana-Galbis D, Montes MJ, Mercade E (2011) Pseudomonas deceptionensis sp. nov., a psychrotolerant bacterium from the Antarctic. Int J Syst Evol Microbiol 61(Pt10):2401–2405. https://doi.org/10.1099/ijs.0.024919-0
Catal T, Fan Y, Li K, Bermek H, Liu H (2011) Utilization of mixed monosaccharides for power generation in microbial fuel cells. J Chem Technol Biotechnol 86(4):570–574. https://doi.org/10.1002/jctb.2554
Coker JA (2016) Extremophiles and biotechnology: current uses and prospects. F1000Research. https://doi.org/10.12688/f1000research.7432.1
Cowan DA, Tow LA (2004) Endangered Antarctic Environments. Annual Rev of Microbiol 58:649–690. https://doi.org/10.1146/annurev.micro.57.030502.090811
Deming JW (2002) Psychrophiles and polar regions. Curr Opin Microbiol 5(3):301–309. https://doi.org/10.1016/s1369-5274(02)00329-6
Eng JK, McCormack AL, Yates JR (1994) An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J Am Soc Mass Spectrom 5:976–989. https://doi.org/10.1016/1044-0305(94)80016-2
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol 57(Pt 1):81–91. https://doi.org/10.1099/ijs.0.64483-0
Gunes Y, Balci N (2021) The catalytic effect of the heterotrophic bacterium Virgibacillus marismortui on basaltic rock dissolution under excess nutrient conditions. Geomicrobiol J 38(4):315–328. https://doi.org/10.1080/01490451.2020.1852453
Gunes Y, Balci N (2022) Sediment and water geochemistry record of water-rock interactions in King George Island Antarctic Peninsula. Arct Sci 34(1):58–78. https://doi.org/10.1017/S0954102021000560
Güven S, Demirel Zorba NN (2013) Genel Mikrobiyoloji ve Laboratuvar Kılavuzu, Nobel, Ankara
Higuera-Llantén S, Vásquez-Ponce F, Núñez-Gallegos M, Pavlov MS, Marshall S, Olivares-Pacheco J (2017) Phenotypic and genotypic characterization of a novel multi-antibiotic-resistant, alginate hyperproducing strain of Pseudomonas mandelii isolated in Antarctica. Polar Biol 41(3):469–480. https://doi.org/10.1007/s00300-017-2206-0
Hitit M, Özbek M, Ayaz-Guner S, Guner H, Oztug M, Bodu M, Kirbas M, Bulbul B, Bucak MN, Ataman MB, Memili E, Kaya A (2021) Proteomic fertility markers in ram sperm. Anim Reprod Sci. https://doi.org/10.1016/J.ANIREPROSCI.2021.106882
Jang SH, Kim J, Kim J, Hong S, Lee C (2012) Genome sequence of cold-adapted Pseudomonas mandelii strain JR-1. Anim Reprod Sci 194(12):3263. https://doi.org/10.1128/jb.00517-12
Kawahara H (2002) The structures and functions of ice crystal-controlling proteins from bacteria. J Biosci Bioeng 94(6):492–496. https://doi.org/10.1016/s1389-1723(02)80185-2
Keto-Timonen R, Hietala N, Palonen E, Hakakorpi A, Lindström M, Korkeala H (2016) Cold shock proteins: a minireview with special emphasis on Csp-family of enteropathogenic Yersinia. Front Microbiol 7:1151. https://doi.org/10.3389/fmicb.2016.01151
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64((Pt_2)):346–351. https://doi.org/10.1099/ijs.0.059774-0
Konstantinidis KT, Tiedje JM (2005) Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci USA 102(7):2567–2572. https://doi.org/10.1073/pnas.0409727102
Kosina M, Barták M, Mašlaňová I, Pascutti AV, Sedo O, Lexa M, Sedláček I (2013) Pseudomonas prosekii sp. nov., a novel psychrotrophic bacterium from Antarctica. Curr Microbiol 67(6):637–646. https://doi.org/10.1007/s00284-013-0406-6
Kulkarni J, Sharma S, Srivastava AK, Penna S (2020) Halotolerant microbes and their applications in sustainable agriculture. Physiological and biotechnological aspects of extremophiles, 5th edn. Elsevier, Amsterdam, pp 39–49
Kumar GS, Jagannadham MV, Ray MK (2002) Low-temperature-induced changes in composition and fluidity of lipopolysaccharides in the Antarctic psychrotrophic bacterium Pseudomonas syringae. J Bacteriol 184(23):6746–6749. https://doi.org/10.1128/JB.184.23.6746-6749.2002
Leclercq R (2002) Mechanisms of resistance to macrolides and lincosamides: nature of the resistance elements and their clinical implications. Clin Infect Dis 34(4):482–492. https://doi.org/10.1086/324626
Lefort V, Desper R, Gascuel O (2015) FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol Biol Evol 32(10):2798–2800. https://doi.org/10.1093/molbev/msv150
Li R, Jiang Y, Wang X, Yang J, Gao Y, Zi X, Zhang X, Gao H, Hu N (2013) Psychrotrophic Pseudomonas mandelii CBS-1 produces high levels of poly-b-hydroxybutyrate. Springerplus 2:335. https://doi.org/10.1186/2193-1801-2-335
López N, Pettinari MJ, Stackebrandt E, Tribelli PM, Potter M, Steinbüchel A, Méndez BS (2009) Pseudomonas extremaustralis sp. nov., a Poly(3-hydroxybutyrate) producer isolated from an Antarctic environment. Curr Microbiol 59(5):514–519. https://doi.org/10.1007/s00284-009-9469-9
Lopez B, Bastias J, Matus D, Jana R, Leppe M (2021) The geology of King George Island, South Shetland Islands: uniting local geological maps and stratigraphical columns, EGU General Assembly 2021, online, 19–30. https://doi.org/10.5194/egusphere-egu21-13145
Lorv JS, Rose DR, Glick BR (2014) Bacterial ice crystal controlling proteins. Scientifica. https://doi.org/10.1155/2014/976895
Lu L, Zhou L, Zhang H, Weng Y (2018) The effects of industrial energy consumption on energy-related carbon emissions at national and provincial levels in China. Energy Sci Eng 6(5):371–384. https://doi.org/10.1002/ese3.206
Martínez-Espinosa RM (2020) Microorganisms and their metabolic capabilities in the context of the biogeochemical nitrogen cycle at extreme environments. Int J Mol Sci 21(12):4228. https://doi.org/10.3390/ijms21124228
May JM, Owens TW, Mandler MD, Simpson BW, Lazarus MB, Sherman DJ, Davis RM, Okuda S, Massefski W, Ruiz N, Kahne D (2017) The antibiotic novobiocin binds and activates the ATPase that powers lipopolysaccharide transport. J Am Chem Soc 139(48):17221–17224. https://doi.org/10.1021/jacs.7b07736
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun 10(1):1–10. https://doi.org/10.1038/s41467-019-10210-3
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60. https://doi.org/10.1186/1471-2105-14-60
Melisi D, Curcio A, Luongo E, Morelli E, Rimoli MG (2011) D-galactose as a vector for prodrug design. Curr Top Med Chem 11(18):2288–2298. https://doi.org/10.2174/156802611797183258
Mykytczuk NCS, Lawrence JR, Omelon CR, Southam G, Whyte LG (2016) Microscopic characterization of the bacterial cell envelope of Planococcus halocryophilus Or1 during subzero growth at – 15 C. Polar Biol 39:701–712. https://doi.org/10.1007/s00300-015-1826-5
Núñez-Montero K, Barrientos L (2018) Advances in Antarctic research for antimicrobial discovery: a comprehensive narrative review of bacteria from antarctic environments as potential sources of novel antibiotic compounds against human pathogens and microorganisms of industrial importance. Antibiotics 7(4):90. https://doi.org/10.3390/antibiotics7040090
Ottoni JR, e Silva TR, de Oliveira M, Passarini MRZ (2019) Characterization of amylase produced by cold-adapted bacteria from Antarctic samples. Biocatal Agric Biotechnol 23:101452. https://doi.org/10.1016/j.bcab.2019.101452
Panizzon JP, Pilz Júnior HL, Knaak N, Ramos RC, Ziegler DR, Fiuza LM (2015) Microbial diversity: relevance and relationship between environmental conservation and human health. Braz Arch of Bio and Tech 58:137–145. https://doi.org/10.1590/S1516-8913201502821
Peterson E, Kaur P (2018) Antibiotic resistance mechanisms in bacteria: relationships between resistance determinants of antibiotic producers, environmental bacteria, and clinical pathogens. Front Microbiol 9:2928. https://doi.org/10.3389/fmicb.2018.02928
Rakusa-Suszczewski S (2002) King George Island-South Shetland Islands, Maritime Antarctic. Geoecology of Antarctic Ice-Free Coastal Landscapes, 1st edn. Springer, Heidelberg, pp 23–36
Reygaert WC (2018) An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol 4(3):482–501. https://doi.org/10.3934/microbiol.2018.3.482
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106(45):19126–19131. https://doi.org/10.1073/pnas.0906412106
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Prepr. https://doi.org/10.7287/peerj.preprints.1900v1
Saleh-Lakha S, Shannon KE, Henderson SL, Goyer C, Trevors JT, Zebarth BJ, Burton DL (2009) Effect of pH and temperature on denitrification gene expression and activity in Pseudomonas mandelii. Appl Environ Microbiol 75(12):3903–3911. https://doi.org/10.1128/AEM.00080-09
Schwidetzky R, Lukas M, Yazdan Yar A, Kunert AT, Pöschl U, Domke KF, Fröhlich-Nowoisky J, Bonn M, Koop T, Nagata Y, Meister K (2021) Specific Ion-Protein Interactions influence bacterial ice nucleation. Eur J Chem 27(26):7402–7407. https://doi.org/10.1002/chem.202004630
Scoma A, Garrido-Amador P, Nielsen SD, Røy H, Kjeldsen KU (2019) The polyextremophilic bacterium Clostridium paradoxum attains piezophilic traits by modulating its energy metabolism and cell membrane composition. Appl Environ Microbiol 85(15):e00802-e819. https://doi.org/10.1128/AEM.00802-19
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30(14):2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Seker F, Balci N (2022) Microbial acid sulfate weathering of basaltic rocks: implications for enzymatic rocks. Aquat Geochem 28:155–184. https://doi.org/10.1007/s10498-022-09407-8
Sherathiya VN, Schaid MD, Seiler JL, Lopez GC, Lerner TN (2021) GuPPy, a Python toolbox for the analysis of fiber photometry data. Sci Rep 11(1):1–9. https://doi.org/10.1038/s41598-021-03626-9
Shintani T (2019) Food industrial production of monosaccharides using microbial, enzymatic, and chemical methods. Fermentation 5(2):47. https://doi.org/10.3390/fermentation5020047
Silva TR, Duarte AWF, Passarini MRZ, Ruiz ALTG, Franco CH, Moraes CB, Soares de Melo I, Rodrigues RA, Fantinatti-Garboggini F, Oliveira VM (2018) Bacteria from Antarctic environments: diversity and detection of antimicrobial, antiproliferative, and antiparasitic activities. Polar Biol 41:1505–1519. https://doi.org/10.1007/s00300-018-2322-5
Singh R, Buttar HK, Manchanda G, Kaur R et al (2020) The multifaceted life of microbes survival varied environments. In: Manchanda G, Singh RP et al (eds) Microbial versatility varied environments. Springer, Singapore, pp 3–10
Stothard P, Wishart DS (2005) Circular genome visualization and exploration using CGView. Bioinformatics 21(4):537–539. https://doi.org/10.1093/bioinformatics/bti054
Sultan I, Rahman S, Jan AT, Siddiqui MT, Mondal AH, Haq QMR (2018) Antibiotics, resistome and resistance mechanisms: a bacterial perspective. Front Microbiol 9:2066. https://doi.org/10.3389/fmicb.2018.02066
Ting L, Williams TJ, Cowley MJ, Lauro FM, Guilhaus M, Raftery MJ, Cavicchioli R (2010) Cold adaptation in the marine bacterium, Sphingopyxis alaskensis, assessed using quantitative proteomics. Environ Microbiol 12(10):2658–2676. https://doi.org/10.1111/j.1462-2920.2010.02235.x
Tribelli PM, López NI (2018) Reporting key features in cold-adapted bacteria. Life 8(1):8. https://doi.org/10.3390/life8010008
Vásquez-Ponce F, Higuera-Llantén S, Pavlov MS, Marshall SH, Olivares-Pacheco J (2018) Phylogenetic MLSA and phenotypic analysis identification of three probable novel Pseudomonas species isolated on King George Island, South Shetland Antarctica. Braz J of Microbiol 49(4):695–702. https://doi.org/10.1016/j.bjm.2018.02.005
Verhille S, Baida N, Dabboussi F, Izard D, Leclerc H (1999) Taxonomic study of bacteria isolated from natural mineral waters: proposal of Pseudomonas jessenii sp. nov. and Pseudomonas mandelii sp. nov. Syst Appl Microbiol 22(1):45–58. https://doi.org/10.1016/S0723-2020(99)80027-7
Verwaaijen B, Cevahir Ö, Hitz F, Römmich J, Wulf D (2022) Complete genome sequence of Pseudomonas sp. Strain MM213, an Isolate from a brookside in Bielefeld. Germany Microbiol Resour Announc 11(1):86621. https://doi.org/10.1128/mra.00866-21
Wick RR, Judd LM, Gorrie CL, Holt KE (2017a) Completing bacterial genome assemblies with multiplex MinION sequencing. Microb Genom 3(10):e000132. https://doi.org/10.1099/mgen.0.000132
Wick RR, Judd LM, Gorrie CL, Holt KE (2017b) Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol 13(6):e1005595. https://doi.org/10.1371/journal.pcbi.1005595
Xu H, Griffith M, Patten CL, Glick BE (1998) Isolation and characterization of an antifreeze protein with ice nucleation activity from the plant growth promoting rhizobacterium Pseudomonas putida GR12-2. Can J Microbiol 44(1):64–73. https://doi.org/10.1139/w97-126
Acknowledgements
This study was carried under the auspices of Presidency of The Republic of Turkey, supported by the Ministry of Industry and Technology, and coordinated by Istanbul Technical University (ITU) Polar Research Center (PolReC). This study was in the frame of bilateral agreement with Polish Institute of Biochemistry and Biophysics (IBB) and PolReC. We would like to thank to 42nd expedition crew for their hospitality and help in Henryk Arctowski Polish Antarctic Station.
Funding
This project was supported by TUBITAK (the Scientific and Technological Research Council of Turkey) (Grant No. 119Y411) and ITU Scientific Research Projects Division (Grant Nos. 41729 and 42256).
Author information
Authors and Affiliations
Contributions
NGK and NB conceived the research. NB and YG collected the samples. HMT, ETSC, MMK, AC, and MOK conducted the laboratory work. HMT, IK, MOK, ETSC analyzed the data. HMT, ETSC, MMK, AC, YG, IK, and MOK wrote the original manuscript. All authors revised and approved the manuscript. NGK received funding from Istanbul Technical University, Scientific Research Projects Coordination Unit.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mumcu, H., Sarac Cebeci, E.T., Menekse Kılıc, M. et al. Identification of phenotypic and genotypic properties and cold adaptive mechanisms of novel freeze–thaw stress-resistant strain Pseudomonas mandelii from Antarctica. Polar Biol 46, 169–183 (2023). https://doi.org/10.1007/s00300-023-03114-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-023-03114-y