Abstract
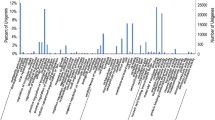
Unique physiological and metabolic properties of Arctic mosses are responsible for their acclimation to the inclement polar environment. To perform transcriptome analysis of an Arctic moss species adapted to polar conditions, we constructed a complementary DNA (cDNA) library using total high-quality RNA extracted from the moss species Aulacomnium turgidum. The library consisted of 1.81 × 106 of independent clones with 97.41% of recombinants. A total of 509 cDNA clones were sequenced. After eliminating poor quality sequences, vector trimming and clustering, 360 unigenes consisting of 33 contigs and 327 singletons were identified. Basic Local Alignment Search Tool X searches generated 245 significant hits (E value <10−5). For further Gene Ontology analysis, 158 unigenes were annotated and classified with terms for molecular function, biological process and cellular component. Among the expressed sequence tags, seven genes were selected based on their putative roles in stress response, and they showed enhanced transcripts level under various abiotic stresses such as low temperature, heat and high-salinity. Also, two rare-cold-inducible genes showed different expression patterns under low temperature and UV-B treatment, indicating their distinct roles in adaptation to Arctic environment. Although experiments have been conducted on a limited scale, this study provides useful information for better understanding the mechanism of stress acclimation of polar mosses and material basis for potential genomic modification for higher plants to increase stress tolerance.





Similar content being viewed by others
References
Achkor H, Díaz M, Fernández MR, Biosca JA, Parés X, Martínez MC (2003) Enhanced formaldehyde detoxification by overexpression of glutathione-dependent formaldehyde dehydrogenase from Arabidopsis. Plant Physiol 132:2248–2255. doi:10.1104/pp.103.022277
Ashton NW, Cove DJ (1977) The isolation and preliminary characterisation of auxotrophic and analogue resistant mutants of the moss, Physcomitrella patens. Mol Gen Genet 154:87–95. doi:10.1007/BF00265581
Ayres E, van der Wal R, Sommerkorn M, Bardgett RD (2006) Direct uptake of soil nitrogen by mosses. Biol Lett 2:286–288. doi:10.1098/rsbl.2006.0455
Barber J, De Las Rivas J (1993) A functional model for the role of cytochrome b559 in the protection against donor and acceptor side photoinhibition. Proc Natl Acad Sci USA 90:10942–10946. doi:10.1073/pnas.90.23.10942
Billings WD, Mooney HA (1968) The ecology of arctic and alpine plants. Biol Rev 43:481–529. doi:10.1111/j.1469-185X.1968.tb00968.x
Birks HJB, Jones VJ, Rose NL (2004) Recent environmental change and atmospheric contamination on Svalbard as recorded in lake sediments—an introduction. J Paleolimnol 31:403–410. doi:10.1023/B:JOPL.0000022542.87722.a6
Birnboim HC (1992) Extraction of high molecular weight RNA and DNA from cultured mammalian cells. Methods Enzymol 216:154–160
Brown AP, Dunn MA, Goddard NJ, Hughes MA (2001) Identification of a novel low-temperature-response element in the promoter of the barley (Hordeum vulgare L) gene blt101.1. Planta 213:770–780. doi:10.1007/s004250100549
Capel J, Jarillo JA, Salinas J, Martínez-Zapater JM (1997) Two homologous low-temperature-inducible genes from Arabidopsis encode highly hydrophobic proteins. Plant Physiol 115:569–576. doi:10.1104/pp.115.2.569
Flatberg KI, Thingsgaard K (2003) Taxonomy and geography of Sphagnum tundrae with a description of S. mirum, sp. nov. (Sphagnaceae, sect. Squarrosa). Bryol 106:501–515. doi:10.1639/0007-2745(2003)106[501:TAGOST]2.0.CO;2
Frishman D (2007) Protein annotation at genomic scale: the current status. Chem Rev 107:3448–3466. doi:10.1021/cr068303k
Fu XH, Geng SL, Su GH, Zeng QL, Shi SH (2004) Isolating high-quality RNA from mangroves without liquid nitrogen. Plant Mol Biol Report 22:197a–197e. doi:10.1007/BF02772728
Goddard NJ, Dunn MA, Zhang AJ, White PL, Jack PL, Hughes MA (1993) Molecular analysis and spatial expression pattern of a low-temperature-specific barley gene, blt 101. Plant Mol Biol 23:871–879
Gornall JL, Jónsdóttir IS, Woodin SJ, Van der Wal R (2007) Arctic mosses govern below-ground environment and ecosystem processes. Oecologia 153:931–941. doi:10.1007/s00442-007-0785-0
Greilhuber J, Såstad SM, Flatberg KI (2003) Ploidy determination in Sphagnum samples from Svalbard, Arctic Norway, by DNA image cytometry. J Bryol 25:235–239. doi:10.1179/037366803225013083
Harris MA, Clark J, Ireland A et al (2004) The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res 32:D258–D261. doi:10.1093/nar/gkh036
Jones C, Brown A, Baumann U (2007) Estimating the annotation error rate of curated GO database sequence annotations. BMC Bioinformatics 8:170. doi:10.1186/1471-2105-8-170
Lee H, Cho HH, Kim IC, Yim JH, Lee HK, Lee YK (2008) Expressed sequence tag analysis of Antarctic hairgrass Deschampsia antarctica from King George Island, Antarctica. Mol Cells 25:258–264
Longton RE (1997) The role of bryophytes and lichens in polar ecosystems. In: Woodin SJ, Marquiss M (eds) Ecology of Arctic environments. Blackwell, Oxford, pp 69–96
Lovelock CE, Jackson AE, Melick DR, Seppelt RD (1995) Reversible photoinhibition in Antarctic moss during freezing and thawing. Plant Physiol 109:955–961
Mathew A, Morimoto RI (1998) Role of the heat-shock response in the life and death of proteins. Ann N Y Acad Sci 851:99–111. doi:10.1111/j.1749-6632.1998.tb08982.x
Mathew A, Mathur SK, Morimoto RI (1998) Heat shock response and protein degradation: regulation of HSF2 by the ubiquitin–proteasome pathway. Mol Cell Biol 18:5091–5098
Medina J, Catalá R, Salinas J (2001) Developmental and stress regulation of RCI2A and RCI2B, two cold-inducible genes of Arabidopsis encoding highly conserved hydrophobic proteins. Plant Physiol 125:1655–1666. doi:10.1104/pp.125.4.1655
Morgan-Kiss RM, Priscu JC, Pocock T, Gudynaite-Savitch L, Huner NPA (2006) Adaptation and acclimation of photosynthetic microorganisms to permanently cold environments. Microbiol Mol Biol Rev 70:222–252. doi:10.1128/MMBR.70.1.222-252.2006
Rabbani MA, Maruyama K, Abe H, Khan MA, Katsura K, Ito Y, Yoshiwara K, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Monitoring expression profiles of rice genes under cold, drought, and high-salinity stresses and abscisic acid application using cDNA microarray and RNA gel-blot analyses. Plant Physiol 133:1755–1767. doi:10.1104/pp.103.025742
Reski R, Frank W (2005) Moss (Physcomitrella patens) functional genomics-gene discovery and tool development, with implications for crop plants and human health. Brief Funct Genomics Proteomics 4:48–57. doi:10.1093/bfgp/4.1.48
Rogers EE, Ausubel FM (1997) Arabidopsis enhanced disease susceptibility mutants exhibit enhanced susceptibility to severa1 bacterial pathogens and alterations in PR-I gene expression. Plant Cell 9:305–316. doi:10.1105/tpc.9.3.305
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Plainview
Skotnicki ML, Ninhami JA, Selkirk PM (2000) Genetic diversity, mutagenesis and dispersal of Antarctic mosses—a review of progress with molecular studies. Antarct Sci 12:363–373
Skotnicki ML, Mackenzie AM, Clements MA, Selkirk PM (2005) DNA sequencing and genetic diversity of the 18S–26S nuclear ribosomal internal transcribed spacers (ITS) in nine Antarctic moss species. Antarct Sci 17:377–384. doi:10.1017/S0954102005002816
Wang WX, Vinocur B, Shoseyov O, Altman A (2004) Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9:244–252. doi:10.1016/j.tplants.2004.03.006
Yordanov I (1995) Responses of photosynthesis to stress and plant growth regulators. Bulg J Plant Physiol 21:51–70
Acknowledgments
The research was financially supported by a Grant PE08050 from the Korea Polar Research Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, S., Lee, H., Kang, PS. et al. Complementary DNA library construction and expressed sequence tag analysis of an Arctic moss, Aulacomnium turgidum . Polar Biol 33, 617–626 (2010). https://doi.org/10.1007/s00300-009-0737-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-009-0737-8