Abstract
Key message
Induced mutations in the waxy locus in rice endosperm did not abolish GBSS activity completely. Compensatory mechanisms in endosperm and leaves caused a major reprogramming of the starch biosynthetic machinery.
Abstract
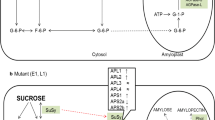
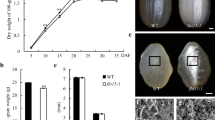
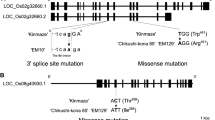
The mutation of genes in the starch biosynthesis pathway has a profound effect on starch quality and quantity and is an important target for plant breeders. Mutations in endosperm starch biosynthetic genes may impact starch metabolism in vegetative tissues such as leaves in unexpected ways due to the complex feedback mechanisms regulating the pathway. Surprisingly this aspect of global starch metabolism has received little attention. We used CRISPR/Cas9 to introduce mutations affecting the Waxy (Wx) locus encoding granule-bound starch synthase I (GBSSI) in rice endosperm. Our specific objective was to develop a mechanistic understanding of how the endogenous starch biosynthetic machinery might be affected at the transcriptional level following the targeted knock out of GBSSI in the endosperm. We found that the mutations reduced but did not abolish GBSS activity in seeds due to partial compensation caused by the upregulation of GBSSII. The GBSS activity in the mutants was 61–71% of wild-type levels, similarly to two irradiation mutants, but the amylose content declined to 8–12% in heterozygous seeds and to as low as 5% in homozygous seeds, accompanied by abnormal cellular organization in the aleurone layer and amorphous starch grain structures. Expression of many other starch biosynthetic genes was modulated in seeds and leaves. This modulation of gene expression resulted in changes in AGPase and sucrose synthase activity that explained the corresponding levels of starch and soluble sugars.








Similar content being viewed by others
References
Ball SG, Morell MK (2003) From bacterial glycogen to starch: understanding the biogenesis of the plant starch granule. Ann Rev Plant Biol 54:207–233
Baroja-Fernández E, Muñoz FJ, Li J, Bahaji A, Almagro G, Montero M, Etxeberria E, Hidalgo M, Sesma T, Pozueta-Romero J (2012) Sucrose synthase activity in the sus1/sus2/sus3/sus4 Arabidopsis mutant is sufficient to support normal cellulose and starch production. Proc Natl Acad Sci USA 109:321–326
Bassie L, Zhu C, Romagosa I, Christou P, Capell T (2008) Transgenic wheat plants expressing an oat arginine decarboxylase cDNA exhibit increases in polyamine content in vegetative tissue and seeds. Mol Breed 22:39–50
Baysal C, Bortesi L, Zhu C, Farré G, Schillberg S, Christou P (2016) CRISPR/Cas9 activity in the rice OsBEIIb gene does not induce off-target effects in the closely related paralog OsBEIIa. Mol Breed 36:108
Bortesi L, Zhu C, Zischewski J, Perez L, Bassié L, Nadi R, Forni G, Boyd-Lade S, Soto E, Jin X, Medina V, Villorbina G, Muñoz P, Farré G, Fischer R, Twyman R, Capell T, Christou P, Schillberg S (2016) Patterns of CRISPR/Cas9 activity in plants, animals and microbes. Plant Biotechnol J 14:2203–2216
Chari R, Mali P, Moosburner M, Church GM (2015) Unraveling CRISPR-Cas9 genome engineering parameters via a library-on-library approach. Nat Methods 12:823
Christou P, Ford TL, Kofron M (1991) Production of transgenic rice (Oryza sativa L.) plants from agronomically important indica and japonica varieties via electric discharge particle acceleration of exogenous DNA into immature zygotic embryos. Nat Biotechnol 9:957–962
CISC (Consejo Superior de Investigaciones Científicas) (2016) Método para la determinación “in situ” de actividades enzimáticas relacionadas con el metabolismo del carbono en hojas. ES Patent no 7:915.111.623.106, 2016-05-06
Crofts N, Abe N, Oitome NF, Matsushima R, Hayashi M, Tetlow IJ, Emes MJ, Nakamura Y, Fujita N (2015) Amylopectin biosynthetic enzymes from developing rice seed form enzymatically active protein complexes. J Exp Bot 66:4469–4482
Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Claverie JM, Gascuel O (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucl Acids Res 36:W465–W469
Farré G, Sudhakar D, Naqvi S, Sandmann G, Christou P, Capell T, Zhu C (2012) Transgenic rice grains expressing a heterologous ρ-hydroxyphenylpyruvate dioxygenase shift tocopherol synthesis from the γ to the α isoform without increasing absolute tocopherol levels. Transgenic Res 21:1093–1097
Fujita N, Yoshida M, Asakura N, Ohdan T, Miyao A, Hirochika H, Nakamura Y (2006) Function and characterization of starch synthase I using mutants in rice. Plant Physiol 140:1070–1084
Heigwer F, Kerr G, Boutros M (2014) E-CRISP: fast CRISPR target site identification. Nat Methods 11:122
Hirano HY (1993) Genetic variation and gene regulation at the wx locus in rice. Gamma Field Symp 24:63–79
Hirano HY, Sano Y (1998) Enhancement of Wx gene expression and the accumulation of amylose in response to cool temperatures during seed development in rice. Plant Cell Physiol 39:807–812
Itoh K, Nakajima M, Shimamoto K (1997) Silencing of waxy genes in rice containing Wx transgenes. Mol Gen Genet 255:351–358
Itoh K, Ozaki H, Okada K, Hori H, Takeda Y, Mitsui T (2003) Introduction of Wx transgene into rice wx mutants leads to both high-and low-amylose rice. Plant Cell Physiol 44:473–480
Jeon JS, Ryoo N, Hahn TR, Walia H, Nakamura Y (2010) Starch biosynthesis in cereal endosperm. Plant Physiol Biochem 48:383–392
Jiang D, Cao WX, Dai TB, Jing Q (2004) Diurnal changes in activities of related enzymes to starch synthesis in grains of winter wheat. Acta Bot Sin 46:51–57
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucl Acids Res 41:e188–e188
Jobling S (2004) Improving starch for food and industrial applications. Curr Opin Plant Biol 7:210–218
Juliano BO (1971) A simplified assay for milled-rice amylose. Cereal Sci Today 16:334–336
Kang TJ, Yang MS (2004) Rapid and reliable extraction of genomic DNA from various wild-type and transgenic plants. BMC Biotechnol 4:20
Kawagoe Y, Kubo A, Satoh H, Takaiwa F, Nakamura Y (2005) Roles of isoamylase and ADP-glucose pyrophosphorylase in starch granule synthesis in rice endosperm. Plant J 42:164–174
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protov 10:845
Li J, Baroja-Fernández E, Bahaji A, Muñoz FJ, Ovecka M, Montero M (2013) Enhancing sucrose synthase activity results in an increased levels of starch and ADP-Glucose in maize (Zea mays L.) seed endosperms. Plant Cell Physiol 54:282–294
Li C, Powell PO, Gilbert RG (2017) Recent progress toward understanding the role of starch biosynthetic enzymes in the cereal endosperm. Amylase 1:59–74
Liu L, Ma X, Liu S, Zhu C, Jiang L, Wang Y, Shen Y, Ren Y, Dong H, Chen L, Liu X, Zhao Z, Zhai H, Wan J (2009) Identification and characterization of a novel Waxy allele from a Yunnan rice landrace. Plant Mol Biol 71:609–626
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, Xie Y, Shen R, Chen S, Wang Z, Chen Y, Guo J, Chen L, Zhao X, Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8:1274–1284
Maddelein ML, Libessart N, Bellanger F, Delrue B, D’Hulst C, Van den Koornhuyse N, Fontaine T, Wieruszeski JM, Decq A, Ball S (1994) Toward an understanding of the biogenesis of the starch granule. Determination of granule-bound and soluble starch synthase functions in amylopectin synthesis. J Biol Chem 269:25150–25157
Martin C, Smith AM (1995) Starch biosynthesis. Plant Cell 7:971–985
Momma M, Fujimoto Z (2012) Interdomain disulfide bridge in the rice granule bound starch synthase I catalytic domain as elucidated by X-ray structure analysis. Biosci Biotechnol Biochem 76:1591–1595
Nakamura Y (2002) Towards a better understanding of the metabolic system for amylopectin biosynthesis in plants: rice endosperm as a model tissue. Plant Cell Physiol 43:718–725
Nakamura Y, Yuki K, Park SY, Ohya T (1989) Carbohydrate metabolism in the developing endosperm of rice grains. Plant Cell Physiol 30:833–839
Ohdan T, Francisco Jr PB, Sawada T, Hirose T, Terao T, Satoh H, Nakamura Y (2005) Expression profiling of genes involved in starch synthesis in sink and source organs of rice. J Exp Bot 56:3229–3244
Pérez L, Soto E, Villorbina G, Bassie L, Medina V, Muñoz P, Capell T, Zhu C, Christou P, Farré G (2018) CRISPR/Cas9-induced monoallelic mutations in the cytosolic AGPase large subunit gene APL2 induce the ectopic expression of APL2 and the corresponding small subunit gene APS2b in rice leaves. Transgenic Res 27:423–439
Preiss J (1982) Regulation of the biosynthesis and degradation of starch. Annu Rev Plant Physiol 33:431–454
Rösti S, Fahy B, Denyer K (2007) A mutant of rice lacking the leaf large subunit of ADP-glucose pyrophosphorylase has drastically reduced leaf starch content but grows normally. Funct Plant Biol 34:480–489
Ryoo N, Yu C, Park CS, Baik MY, Park IM, Cho MH, Bhoo SH, An G, Hahn TR, Jeon JS (2007) Knockout of a starch synthase gene OsSSIIIa/Flo5 causes white-core floury endosperm in rice (Oryza sativa L.). Plant Cell Rep 26:1083–1095
Sano Y (1984) Differential regulation of waxy gene expression in rice endosperm. Theor Appl Genet 68:467–473
Satoh H, Shibahara K, Tokunaga T, Nishi A, Tasaki M, Hwang SK, Okita TW, Kaneko N, Fujita N, Yoshida M, Hosaka Y, Sato A, Utsumi Y, Ohdan T, Nakamura Y (2008) Mutation of the plastidial α-glucan phosphorylase gene in rice affects the synthesis and structure of starch in the endosperm. Plant Cell 20:1833–1849
Seung D, Soyk S, Coiro M, Maier BA, Eicke S, Zeeman SC (2015) PROTEIN TARGETING TO STARCH is required for localising GRANULE-BOUND STARCH SYNTHASE to starch granules and for normal amylose synthesis in Arabidopsis. PLOS Biol 13:e1002080
Shan Q, Wang Y, Li J, Gao C (2014) Genome editing in rice and wheat using the CRISPR/Cas system. Nature Protoc 9:2395
Sudhakar D, Bong BB, Tinjuangjun P, Maqbool SB, Valdez M, Jefferson R, Christou P (1998) An efficient rice transformation system utilizing mature seed-derived explants and a portable, inexpensive particle bombardment device. Transgenic Res 7:289–294
Sun Y, Jiao G, Liu Z, Zhang X, Li J, Guo X, Du J, Francis F, Zhao Y, Xia L (2017) Generation of high-amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes. Front Plant Sci 8:298
Tang XJ, Peng C, Zhang J, Cai Y, You XM, Kong F, Yan HG, Wang GX, Wang L, Jin J, Chen WW, Chen XG, Ma J, Wang P, Jiang L, Zhang WW, Wan JM (2016) ADP-glucose pyrophosphorylase large subunit 2 is essential for storage substance accumulation and subunit interactions in rice endosperm. Plant Sci 249:70–83
Terada R, Nakajima M, Isshiki M, Okagaki RJ, Wessler SR, Shimamoto K (2000) Antisense waxy genes with highly active promoters effectively suppress waxy gene expression in transgenic rice. Plant Cell Physiol 41:881–888
Tetlow IJ (2011) Starch biosynthesis in developing seeds. Seed Sci Res 21:5–32
Tetlow IJ, Morell MK, Emes MJ (2004) Recent developments in understanding the regulation of starch metabolism in higher plants. J Exp Bot 55:2131–2145
Thitisaksakul M, Jiménez RC, Arias MC, Beckles DM (2012) Effects of environmental factors on cereal starch biosynthesis and composition. J Cereal Sci 56:67–80
Tran NA, Daygon VD, Resurreccion AP, Cuevas RP, Corpuz HM, Fitzgerald MA (2011) A single nucleotide polymorphism in the Waxy gene explains a significant component of gel consistency. Theor Appl Genet 123:519–525
Umemoto T, Terashima K (2002) Activity of granule-bound starch synthase is an important determinant of amylose content in rice endosperm. Funct Plant Biol 29:1121–1124
Valdez M, Cabrera-Ponce JL, Sudhakar D, Herrera-Estrella L, Christou P (1998) Transgenic Central American, West African and Asian elite rice varieties resulting from particle bombardment of foreign DNA into mature seed-derived explants utilizing three different bombardment devices. Ann Bot 82:795–801
Wang JC, Xu H, Zhu Y, Liu QQ, Cai XL (2013) OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm. J Exp Bot 64:3453–3466
Wong KS, Kubo A, Jane JL, Harada K, Satoh H, Nakamura Y (2003) Structures and properties of amylopectin and phytoglycogen in the endosperm of sugary-1 mutants of rice. J Cereal Sci 37:139–149
Yatou O, Amano E (1991) DNA structure of mutant genes in the waxy locus in rice. In: Plant mutation breeding for crop improvement, vol 2. IAEA, Vienna/FAO, Rome, pp 385–389
Yoshida S, Forno DA, Cock JH, Gomez KA (1976) Determination of sugar and starch in plant tissue. In: Laboratory manual for physiological studies of rice, 3rd edn. IRRI, Los Baños, Philippines, pp 46–49
Zhang G, Cheng Z, Zhang X, Guo X, Su N, Jiang L, Mao L, Wan J (2011) Double repression of soluble starch synthase genes SSIIa and SSIIIa in rice (Oryza sativa L.) uncovers interactive effects on the physicochemical properties of starch. Genome 54:448–459
Zhang MZ, Fang JH, Yan X, Liu J, Bao JS, Fransson G, Andersson R, Jansson C, Aman P, Sun C (2012) Molecular insights into how a deficiency of amylose affects carbon allocation–carbohydrate and oil analyses and gene expression profiling in the seeds of a rice waxy mutant. BMC Plant Biol 12:230
Zhang J, Zhang H, Botella JR, Zhu JK (2018) Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J Int Plant Biol 60:369–375
Zhu C, Bortesi L, Baysal C, Twyman RM, Fischer R, Capell T, Schillberg S, Christou P (2017) Characteristics of genome editing mutations in cereal crops. Trends Plant Sci 22:38–52
Acknowledgements
We thank Dr. Caixia Gao (Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing, China) for providing pJIT163-2NLSCas9 containing the rice codon-optimized cas9, and the empty pU3-gRNA vector. We thank Professor Kimiko Itoh, Graduate School of Science and Technology (Niigata University, Japan) for providing the KUR and Musa Wx lines. This work was supported by funding from the Spanish Ministry of Economy and Competitiveness (MINECO) (BIO2014-54426), and a Juan de la Cierva fellowship to GF (IJCI-2014-19528). LP is the recipient of MINECO fellowship. ES is the recipient of a PhD fellowship from the University of Lleida (BIO2014-54441-P).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared that no conflicts of interest exist.
Additional information
Communicated by Fabien Nogué.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pérez, L., Soto, E., Farré, G. et al. CRISPR/Cas9 mutations in the rice Waxy/GBSSI gene induce allele-specific and zygosity-dependent feedback effects on endosperm starch biosynthesis. Plant Cell Rep 38, 417–433 (2019). https://doi.org/10.1007/s00299-019-02388-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-019-02388-z